Translate this page into:
An electrochemical CRISPR/Cas12a biosensing platform utilizing Fe3O4/α-Fe2O3 nanosheets for highly sensitive detection of EGFR-19Del
* Corresponding authors: E-mail addresses: luckystar_lrj@ujs.edu.cn (R. Liu), aolinhe@163.com (A. He)
-
Received: ,
Accepted: ,
Abstract
A sensitive electrochemical biosensing platform CRISPR based was developed for the detection of EGFR-19Del. In this work, the trans-cleavage activity of target-induced CRISPR/Cas12a was investigated by monitoring the electrical signal changes of methylene blue (MB) on the electrode surface. The construction parameters of the biosensing platform were optimized, resulting in a linear correlation between ΔI% and the logarithm of target double-stranded DNA concentration within the range of 10 pM to 100 nM. The equation was expressed as ΔI% = 19.35 lg C (nM) + 52.99 (R2 = 0.997), with a limit of detection (LOD) of 0.8 pM and a limit of quantitation (LOQ) of 2.7 pM. The biosensing platform exhibited good selectivity by successful detection of interfering sequences target sequences. Five independent electrodes were employed for detecting tDNA at 100 pM and 10 nM concentrations, yielding relative standard deviation (RSD) values of only 3.05% and 4.61%, respectively, thus confirming excellent reproducibility for this biosensing platform. After 10 days of storage, the current response value remained at 97.3% of the initial, indicating the favorable stability of the biosensing platform. The recovery rates of the human serum samples were in the range of 94.73% to 114.50%, indicating that the biosensing platform had the potential to detect clinical samples. Therefore, this biosensor platform based on clustered regularly interspaced short palindromic repeats-associated protein 12a (CRISPR/Cas12a) could provide possibilities for other genetic tests.
Keywords
Detection
EGFR-19Del
Electrochemical
Fe3O4/α-Fe2O3
Nanosheet
1. Introduction
Lung cancer remains the leading malignant tumor with the highest mortality rate worldwide, posing a severe threat to public health and well-being [1]. Non-small cell lung cancer (NSCLC) is the main histological subtype, making up about 85% of all cases [2]. The epidermal growth factor receptor (egfr) mutation serves as a crucial driver gene in NSCLC and plays an indispensable role in targeted personalized therapy for patients [3]. The most prevalent egfr gene mutations are observed in 18-21 exons of the kinase domain [4], with 19 exon deletion mutations exhibiting the highest incidence, accounting for over 50% of all egfr gene mutations. Currently, many detection methods, such as polymerase chain reaction (PCR) [5,6], sanger sequencing [7,8], and amplification refractory mutation system (ARMS) [9,10], are commonly employed. However, these approaches exhibit limited sensitivity and high costs. In contrast, electrochemical detection methods have emerged as a prominent research area due to their cost-effectiveness, enhanced sensitivity, and exceptional specificity [11-13].
The Cas12a-based biosensors were introduced to enhance the detection sensitivity of the biosensors [14,15]. The clustered regularly interspaced short palindromic repeats-associated protein 12a (CRISPR/Cas12a) protein, functioning as an endonuclease, can be programmed with gRNA to specifically bind to a target sequence. Upon binding to the tDNA, the CRISPR/Cas12a protein induces cleavage in both strands of the tDNA, resulting in the cleavage of the double-stranded DNA (cis-cleavage) [16]. Furthermore, upon activation of its cleavage activity, Cas12a protein also cleaves surrounding non-specific single-stranded DNA (trans-cleavage) [17,18]. Consequently, the Cas12a/crRNA binary complex provides sequence-specific immunity against invading nucleic acids. The target can be quantitatively detected by monitoring the electrical signal changes of the signal probe immobilized on the electrode surface by an electrochemical workstation.
The effective immobilization of the signal probe on the electrode surface is a key step in determining the performance of the electrochemical biosensor. The integration of signal probes with magnetic nanocomposites (NCs) can enhance their stability and sensitivity in biological environments [19-21]. Iron oxide nanomaterials have garnered significant attention in research due to their excellent biocompatibility [22-24]. We fabricated magnetically controllable Fe3O4/α-Fe2O3 nanosheets with a large surface area and high surface energy [25]. Fe3O4/α-Fe2O3 nanosheets exhibit an appropriate saturation magnetization, and magnetically induced self-assembly takes place on the surface of the magnetic glassy carbon electrode (MGCE). This enables the signal probe to be immobilized onto the MGCE surface. Furthermore, due to their sheet-like nanostructure with a large specific surface area, more gold nanoparticles (AuNPs) can be anchored, facilitating immobilization of the signal probes onto the Fe3O4/α-Fe2O3@Au via the Au-S bond. Additionally, the Fe3O4/α-Fe2O3@Au NCs demonstrate excellent catalytic activity that enhances the reversibility of redox reactions on the electrode surface, thereby conferring significant advantages in electrochemical sensors and other applications [26,27]. Meanwhile, the combination of magnetic nanomaterials with other noble metal nanoparticles can further improve the sensitivity of the biosensors [28-30].
In this study, we presented the principle of an electrochemical biosensing platform based on CRISPR/Cas12a for ultrasensitive and specific determination of the EGFR-19Del (Figure 1). The Fe3O4/α-Fe2O3@Au NCs were prepared using the sodium borohydride reduction method [31]. Subsequently, sulfhydryl-modified hairpin DNA (hpS2) was covalently attached to the Fe3O4/α-Fe2O3@Au through Au-S bond formation, resulting in the formation of methylene blue-labeled Fe3O4/α-Fe2O3@Au-hpS2. Cas12a (Cpf1) harbored a RuvC endonuclease domain, which remained inactive without activation. Upon binding to a specific crRNA, it underwent a conformational change to form the Cas12a-crRNA binary complex [32,33]. With addition of the target dsDNA, the Cas12a-crRNA-target DNA ternary complex was formed, demonstrating robust non-specific ssDNA cleavage activity (trans cleavage). One end of the nonspecific ssDNA reporter gene (hpS2) was modified with methylene blue (MB) for signal transduction [34-36], while the other end was functionalized with a mercaptan group to enable binding to the surface of Fe3O4/α-Fe2O3@Au NCs for electrochemical detection. The existence of target dsDNA induced the activation of ssDNase activity, resulting in the cleavage of the MB-hpS2 probe into shorter fragments. Consequently, there was a significant reduction in electrochemical signals due to the dissociation of MB from the Fe3O4/α-Fe2O3@Au NCs. Depending on the Cas12a system, the electrochemical biosensing platform demonstrated a significant change in electrochemical current of the signaling probe (hpS2) in the presence or absence of the target analyte. In addition, different target sequences could be quantitatively detected by changing the sequence of crRNA, indicating that the biosensing platform has a potential application prospect.
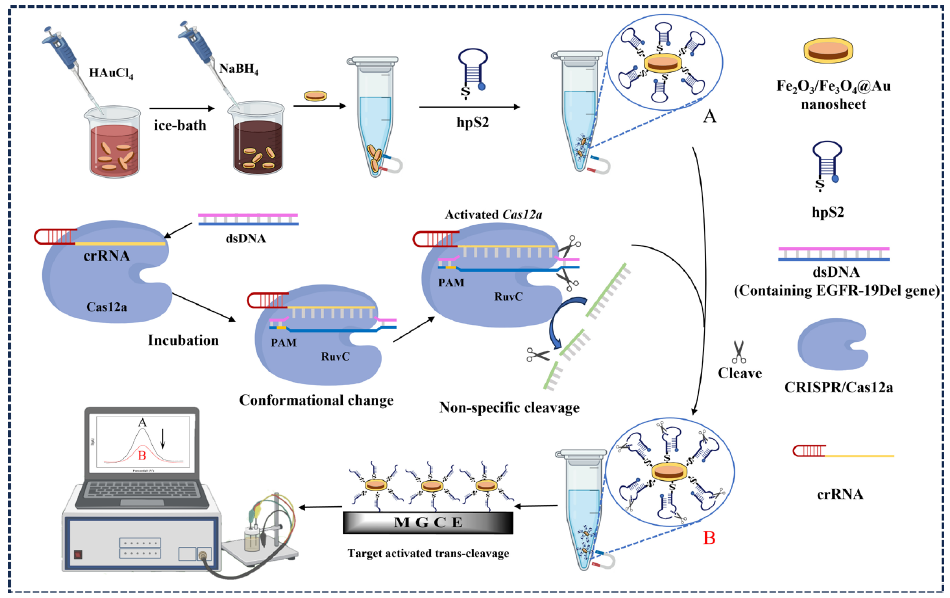
- The scheme of the electrochemical biosensor based on CRISPR/Cas12a. CRISPR: Clustered regularly interspaced short palindromic repeats, Cas12a: CRISPR-associated protein 12a, hps2: Human parainfluenza virus 2, mgce: Monodispersed graphene-cobalt-oxide electrode, dsDNA: Double-stranded DNA, crRNA: CRISPR RNA, ruC: Ruthenium complex.
2. Materials and Methods
2.1. Preparation of Fe3O4/α-Fe2O3@Au NCs
Firstly, FeCl3·6H2O (0.541 g) and NH4H2PO4 (16.6 mg) were dissolved in deionized water (100 mL). The solution of 80 mL was then put into a hydrothermal reactor and kept at 220°C for 24 hrs to obtain α-Fe2O3 nanosheets, and then, the precursor of α-Fe2O3 was partially reduced to Fe3O4 via high-temperature calcination (600°C, 4 hrs) with glucose as a reducing agent to prepare Fe3O4/α-Fe2O3 nanosheets. Subsequently, to improve the electrocatalytic properties of the Fe3O4/α-Fe2O3 nanosheets, sodium borohydride was used to reduce chloroauric acid to prepare Fe3O4/α-Fe2O3@Au NCs [37].
2.2. Construction of the biosensing platform
Construction of Fe3O4/α-Fe2O3@Au-hpS2 modified electrode: The hpS2 was prepared in TE buffer to a final concentration of 100 μM. Subsequently, hpS2 and TCEP (10 mM) were added in a molar ratio of 1:100 and allowed to react at room temperature for 1 hr to facilitate the reduction of disulfide bonds. The reaction mixture was then diluted with Phosphate-Buffered Saline (PBS) buffer (10 mM). The hpS2 (3 μM) solution (40 μL) was incubated with Fe3O4/α-Fe2O3@Au nanosheets (12 mg/mL) at 37°C for 3 hrs. After thorough cleaning with ultra-pure water, the excess water was removed to fabricate the Fe3O4/α-Fe2O3@Au-hpS2 modified electrode. The electrode modified with Fe2O3/α-Fe3O4@Au-hpS2/MCH was constructed by effectively blocking non-specific adsorption sites using MCH.
CRISPR/Cas12a cleavage activity: The Cas12a/crRNA double-stranded complex was formed by assembling 10 μL of a 200 nM Cas12a solution and 5 μL of a 400 nM crRNA in 5 μL of 10×NEBuffer 2.1 buffer, followed by incubation at 37°C for 30 mins. Subsequently, the cleavage activity was initiated by adding 10 μL of target double-stranded DNA (100 nM) and incubating the mixture with an additional 10 μL of Tris-HCl buffer at 37°C for another 30 mins. At this point, the Cas12a/crRNA complex reached a concentration of 50 nM. The Fe3O4/α-Fe2O3@Au-hpS2/MCH was then supplemented with a 40 μL solution of Cas12a/crRNA complex and tDNA, whereafter the execution of CRISPR/Cas12A-mediated cleavage reaction at 37°C for 1 hr. Subsequently, the obtained mixture was subjected to thermal inactivation of Cas12a enzyme at 65°C for 10 mins. After purification with ultra-pure water, dispersion was achieved by adding 40 μL ultra-pure water, and a volume of 9 μL solution was extracted for electrochemical detection. Electrochemical impedance spectroscopy (EIS) and cyclic voltammetry (CV) were utilized to investigate the construction process of the biosensing platform.
2.3. Polyacrylamide gel electrophoresis
Preparation of 12% polyacrylamide gels: Gels for electrophoresis experiments were made by mixing 2.4 mL of 30% (29:1) acrylamide/bis-acrylamide solution, 1.6 mL of 5×TBE buffer, 4 mL of deionized water, 56 μL of ammonium persulfate (APS) and 8 μL of TEMED solution. Subsequently, 10 μL of each sample and 2 μL of bromophenol blue up-sampling buffer were loaded into a 12% polyacrylamide gel. The polyacrylamide gel electrophoresis (PAGE) was placed in 1×TBE run at a constant voltage of 120 V for 45 mins, and then the lane results were observed by UV analyzer.
3. Results and Discussion
3.1. Characterization of modified materials on the electrode surface
Figure 2(a) showed the TEM image of the prepared Fe3O4/α-Fe2O3 nanosheets with an average diameter of 151 nm and thickness of 54 nm, respectively. Also, the size distribution has been shown in Figure S1A, B. The XRD pattern of Fe3O4/α-Fe2O3 nanosheets in Figure 2(b) was consistent with the Powder Diffraction File of Fe3O4 (JCPDS No. 19-0629) and α-Fe2O3 (JCPDS No. 33-0664), indicating the existent of α-Fe2O3 and Fe3O4. The characteristic peaks at 2θ of 24.1°, 33.2°, 35.6°, 40.8°, 49.4°, 54.0°, 62.4°, and 63.9° corresponded to the (012), (104), (110), (113), (024), (116), (214), and (300) crystal faces of α-Fe2O3. While the characteristic peaks at 2θ of 30.1°, 35.4°, 43.1°, 56.9°, and 73.9° corresponded to the (220), (311), (400), (511), and (533) crystal faces of Fe3O4. Additionally, the peak intensity ratio of Fe3O4/α-Fe2O3 nanosheets at 35.4° and 33.2° was higher than that of pure α-Fe2O3 because the presence of Fe3O4 was conducive to its characteristic peak at 35.4°. The average grain size of Fe3O4/α-Fe2O3 nanosheets was about 37 nm. The results confirmed the successful fabrication of Fe3O4/α-Fe2O3 nanosheets. Figure 2(c) showed the TEM image of Fe3O4/α-Fe2O3@Au NCs, AuNPs distributed on the surface of Fe3O4/α-Fe2O3 nanosheets, and the average diameter of AuNPs was about 11.3 nm. The size distributions of Fe3O4/α-Fe2O3@Au NCs and AuNPs have been plotted in Figure S1C, D, and E. The presence of Fe, O, and Au elements in the EDS spectra (Figure 2d) confirmed the successful preparation of the Fe3O4/α-Fe2O3@Au NCs, while the presence of other elements indicated that the DNA probes were successfully bound to the surface of the NCs. In addition, the elemental distribution mappings of the nanoprobes have been exhibited in Figure S2.
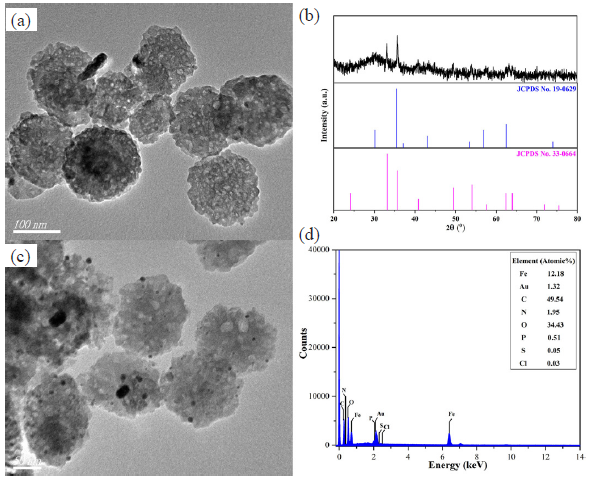
- (a) The TEM image and (b) XRD pattern of Fe3O4/α-Fe2O3 nanosheets, (c) the TEM image of Fe3O4/α-Fe2O3@Au NCs, and (d) the EDS spectral analysis of Fe3O4/α-Fe2O3@Au-hpS2 nanoprobes. TEM: Transmission electron microscopy, XRD: X-ray diffraction, EDS: Energy dispersive spectroscopy.
Furthermore, the chemical composition and electronic states of the elements in the NC were analyzed utilizing XPS, thus validating its successful synthesis (Figure 3). The full spectrum scanning of the NC has been displayed in Figure 3(a), with prominent peaks for Fe, C, Au, and O. The high-resolution spectrum of Fe 2p in Figure 3(b) exhibits two peaks at the binding energy (BE) of 710.70 and 724.40 eV, which correspond to Fe 2p3/2 and Fe 2p1/2, respectively. After peak differentiation imitating, the splitting peaks obtained at BE of 710.29 and 723.89 eV belonged to Fe(II), while the two at 712.14 and 725.64 eV were attributed to Fe(III), along with two satellite peaks at 718.01 and 732.54 eV. As depicted in Figure 3(c), after deconvolution, the C 1s spectrum had three peaks at the BE of 284.80, 285.54, and 288.46 eV, which could be ascribed to C–C/C–H, C–Oc, and O–C=O. Figure 3(d) illustrated two main peaks of the Au 4f region located at 83.84 and 87.52 eV, corresponding to Au(I) 4f7/2 and Au(I) 4f5/2 respectively. Additionally, it can be seen that the O 1s spectrum (Figure 3e) was composed of Fe–O, C–O/C–O=C, and H2O observed at 529.89, 530.88, and 531.91 eV, respectively. In summary, Fe3O4/α-Fe2O3@Au NCs were successfully prepared.
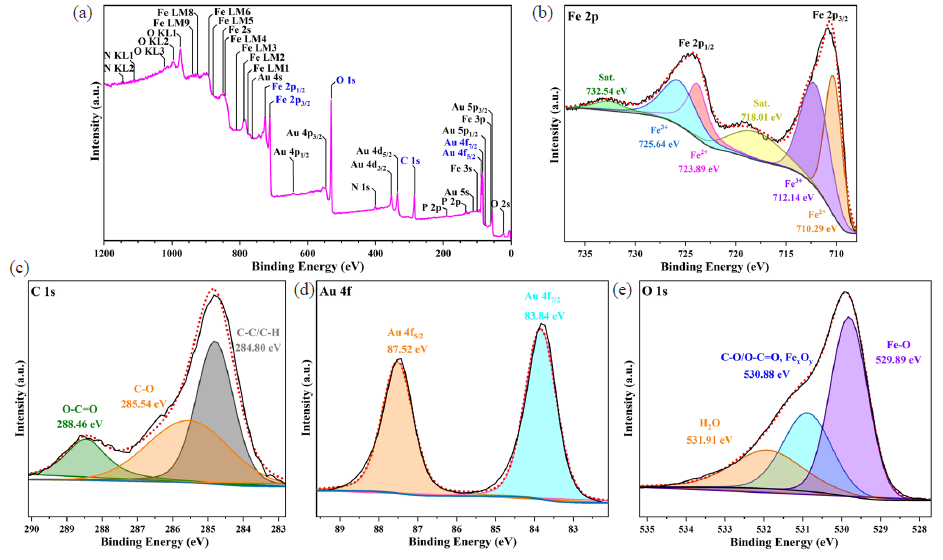
- XPS spectra of Fe3O4/α-Fe2O3@Au NCs nanocomposites: (a) Overview spectrum, (b) Fe 2p, (c) C 1s, (d) Au 4f, and (e) O 1s.
3.2. Electrochemical characterization of the biosensor
Interface properties of the biosensor construction process were investigated using cyclic voltammetry (CV), electrochemical impedance spectroscopy (EIS), and polyacrylamide gel electrophoresis (PAGE) techniques (Figure 4). In Figure 4(a), the curve a represents an unmodified electrode, while curve b corresponded to the deposition of Fe3O4/α-Fe2O3@Au on the electrode surface. The incorporation of AuNPs significantly enhanced the electrical conductivity, leading to an increase in redox current and a corresponding decrease in impedance. These results demonstrated the successful construction of a Fe3O4/α-Fe2O3@Au modified electrode. Subsequently, sulfhydryl-modified hpS2 was connected to Fe3O4/α-Fe2O3@Au by Au-S bonds, and the non-specific site was blocked with MCH to construct the Fe3O4/α-Fe2O3@Au/DNA/MCH modified electrode (curve c). Since the DNA phosphate backbone was negatively charged, there was electrostatic repulsion between it and Fe(CN)6]3-/4- in the solution, which made it more difficult for Fe(CN)6]3-/4- to reach the electrode surface, significantly reducing the redox current and increasing the charge transfer resistance. Curve d showed that the activated Cas12a protein was added to cut the hpS2. Because the nucleotide chain was cut into short fragments, the steric hindrance on the electrode surface decreased, and Fe(CN)6]3-/4- in the solution could reach the electrode surface more easily. Hence, the redox current increased, and the charge transfer resistance decreased.
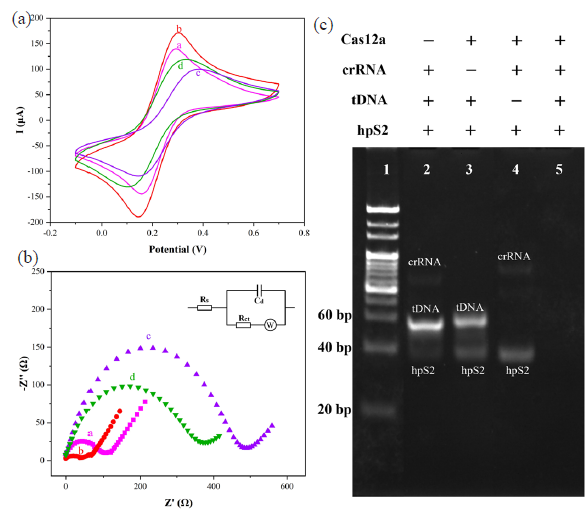
- (a) CV, (b) EIS, and (c) electrophoretic analysis of construction for CRISPR/Cas12a-based electrochemical biosensors. + stands for the factor exists, - means that it does not exist, CV: Cyclic voltammetry, EIS: Electrochemical impedance spectroscopy.
The Nyquist diagram in Figure 4(b) corresponded to the CV diagram, and the Randles equivalent circuit model in the inset of Figure 4(b) was employed for the fitting of the EIS. The examined ligand density of the hpS2 probes was 1.7 (± 0.3) × 1012 molecules per cm2 via the chronocoulometric method [38,39]. The cleavage activity has been confirmed in Figure 4(c). The hpS2 was designed as the substrate for Cas12a/crRNA trans-cleavage. In the absence of Cas12a (lane 2), crRNA (lane 3), or tDNA (lane 4), the silencing of Cas12a trans-cleavage activity was observed. However, specific tDNA binding facilitated the cleavage activity mediated by crRNA-directed Cas12a (lane 5). The results above verified the successful construction of an electrochemical biosensing platform based on CRISPR/Cas12a.
3.3. Optimization of the detection conditions
To achieve optimal sensing performance, the key influencing factors in the experimental process were optimized [40,41]. The concentration of target dsDNA was set at 100 nM, and each experiment group was repeated three times to observe signal changes of differential pulse voltammetry (DPV). Initially, the DPV method was employed to investigate the effect of Fe3O4/α-Fe2O3 @Au NCs’ concentration in the electrolyte solution containing 5 mM Fe(CN)6]3-/4-. As illustrated in Figure S3, as the concentration of NCs increased from 4 to 16 mg/mL, the DPV current exhibited an upward trend due to the good conductivity of AuNPs. However, excessive deposition of NCs on the electrode surface would create significant spatial resistance that hindered electron transfer. Therefore, it was determined that the optimal concentration for Fe3O4/α-Fe2O3@Au NCs was 16 mg/mL.
Moreover, the concentration of hpS2 emerged as a crucial parameter that exerted an impact on both the sensitivity and stability of the biosensing platform (Figure 5a). The peak current of MB served as an indicator for quantifying its concentration. The high concentration of hpS2 could result in electrode surface congestion, thereby impeding CRISPR/Cas12a cleavage activity. Thus, this experiment quantified the variations in current exhibited by hpS2 across a concentration range spanning from 1 μM to 3 μM. The DPV signal of MB was observed to be enhanced as the concentration of hpS2 increased from 1 μM to 2 μM, as illustrated in Figure 5(a). However, the current signal exhibited a subsequent decrease with an increase in the concentration. Consequently, 2 μM was determined to be the optimal concentration. The concentrations of Cas12a and crRNA were crucial factors directly impacting trans-cleavage efficiency. The maximum ΔI% was observed at the concentration of 40 nM as the Cas12a concentration increased. Henceforth, it was selected as the optimal Cas12a concentration (Figure 5b). In Figure 5(c), trans-cleavage activity increased with higher crRNA concentrations. Nevertheless, when exceeding 40 nM, ΔI% slightly decreased. Thus, 40 nM was selected as the optimal crRNA concentration. Furthermore, the effect of cleavage time was also investigated. It achieved stability after 60 min. Hence, 60 min was determined as the optimum condition (Figure 5d). It’s worth noting that ΔI% = (I - I0)/I × 100%, where I was the peak MB current obtained after optimization of hpS2 concentration, and I0 was the current value expressed by the sensor when each condition was optimized.
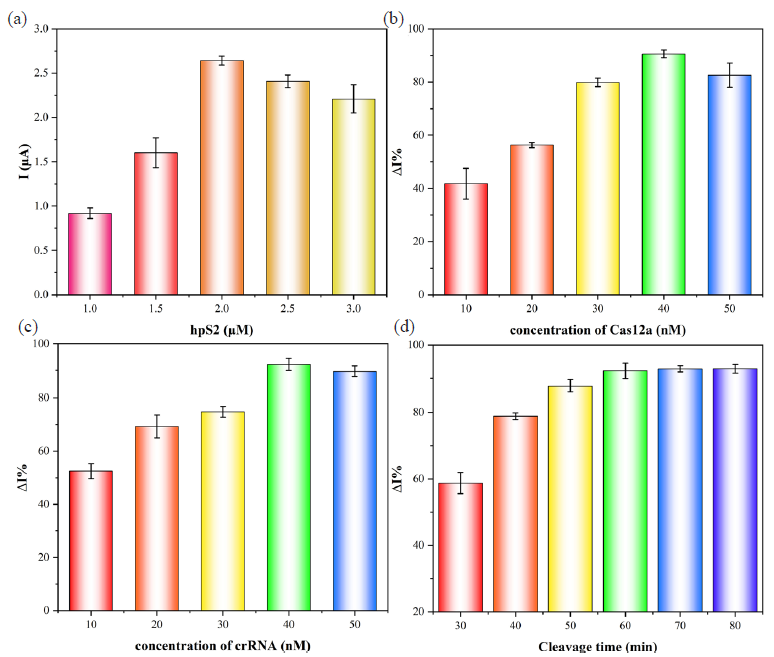
- Optimization: (a) Concentration of hpS2, (b) Cas12a, (c) crRNA, and (d) cleavage time.
3.4. Analytical performance of constructed biosensor towards EGFR-19Del
Under optimized experimental conditions, the performance of the biosensing platform was evaluated utilizing the DPV method, as depicted in Figure 6. The alteration in tDNA concentration could influence the trans-cleavage activity of CRISPR/Cas12a and subsequently resulted in variations in electrical signals on the electrode surface. Within the range of 10 pM to 100 nM, ΔI% was linearly correlated with the logarithm of the tDNA concentration (described by the equation ΔI% = 19.35 lg C (nM) + 52.99 with an R2 value of 0.997). The limit of detection (LOD) was determined as 0.8 pM (LOD=3σ/slope), while the limit of quantification (LOQ) was found to be 2.7 pM (LOQ=10σ/slope). In comparison with previously reported methods (Table 1 and Table S2), this CRISPR-based electrochemical biosensor platform exhibited superior sensitivity with a wider linear range as well as lower LOD, thereby serving as a valuable reference for detecting other nucleic acid sequences using CRISPR-based biosensors.
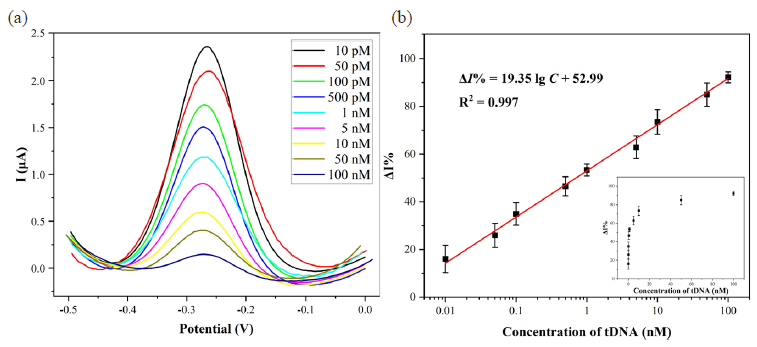
- (a) DPV curves of the biosensing platform for the detection of different concentrations of tDNA and (b) the linear relationship between ΔI% and the logarithm of tDNA concentration. DPV: Differential pulse voltammetry. tDNA: target DNA.
3.5. Selectivity, stability, and reproducibility of the biosensing platform
The selectivity of the biosensing platform was investigated by examining its ability to discriminate between tDNA and non-target sequence, with a concentration of 1 nM for tDNA and 100 nM for the non-target sequences. The current change (ΔI%) of the tDNA was significantly higher than that of the non-complementary sequences, indicating its good selectivity (Figure 7a). The detection of tDNA of 100 pM and 10 nM was performed using a set of five electrodes, yielding RSD values of 3.05% and 4.61%, respectively. These results demonstrated excellent reproducibility (Figure 7b). Moreover, the stability was investigated by recording the DPV current responses for consecutive 10 days. The current fluctuated slightly without significant difference, and the current response value remained still 97.3% of the initial value after 10 days, which indicated the biosensing platform had excellent stability (Figure 7c).
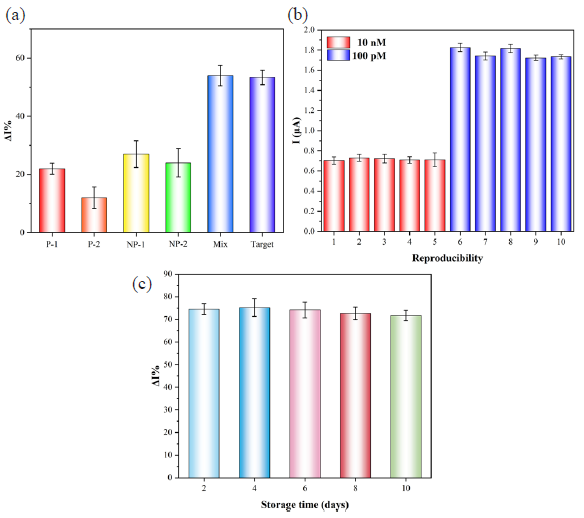
- (a) The selectivity, (b) reproductivity, and (c) stability of the CRISPR/Cas12a-based biosensing platform.
3.6. Analysis of EGFR-19Del in human serum samples
To assess the capability of the biosensing platform for biological sample analysis [42], we conducted tests on serum-labeled samples with various concentrations (Table 2). The serum was diluted 20-fold using a diluent of 20 mM PBS (pH 7.4). The experimental findings revealed that the RSD values ranged from 1.23% to 6.55%, while the recovery rate fell within the range of 94.73% to 114.50%. These results strongly suggested that the biosensor platform had great potential for clinical sample detection.
| Samples | Spiked | Detected | RSD (%) | Recovery (%) |
|---|---|---|---|---|
| 1 | 100 pM | 114.5 pM | 1.54 | 114.50 |
| 1 nM | 1.048 nM | 1.23 | 104.78 | |
| 10 nM | 11.35 nM | 3.77 | 113.54 | |
| 2 | 100 pM | 94.73 pM | 4.15 | 94.73 |
| 1 nM | 1.060 nM | 1.79 | 106.02 | |
| 10 nM | 10.96 nM | 4.37 | 109.61 | |
| 3 |
100 pM 1 nM 10 nM |
99.31 pM 1.138 nM 10.90 nM |
4.20 3.70 6.65 |
99.31 113.77 108.95 |
EGFR-19Del: Epidermal growth factor receptor - 19 base pair deletion, RSD: Relative standard deviation.
4. Conclusions
As demonstrated, an electrochemical biosensing platform based on CRISPR/Cas12a was proposed to detect the EGFR-19Del. Compared with other signal amplification strategies, the proposed electrochemical biosensor platform not only took advantage of the excellent electrical conductivity of Fe3O4/α-Fe2O3@Au NCs to amplify the electrochemical signal but also utilized the target recognition activity of Cas12a to boost the specificity of the biosensor platform. After optimization, the analytical performance was explored. The linear range of the biosensor platform was from 10 pM to 100 nM with a LOD of 0.8 pM. This biosensor platform also had the potential to detect clinical samples with recovery rates in the range of 94.73% to 114.50% in real human serum analysis. More importantly, the CRISPR based biosensor platform could be applied in other gene detectors by rationally designing the sequence of crRNA, thus promoting the development of electrochemical biosensors.
Acknowledgment
Postgraduate Research & Practice Innovation Program of Jiangsu Province (Grant No. KYCX24_4043).
CRediT authorship contribution statement
Zhihao Xu: Experimental studies, Data acquisition, Data analysis, Manuscript preparation. Min Liu: Definition of intellectual content, Literature search, Concepts. Yun Ni: Design, Data acquisition, Experimental studies. Jie Wang: Data acquisition, Data analysis. Hexiao Zhang: Experimental studies. Aolin He: Concepts, Definition of intellectual content, Manuscript editing and review. Ruijiang Liu: Concepts, Design, Definition of intellectual content, Data analysis, Manuscript editing and review.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Declaration of Generative AI and AI-assisted technologies in the writing process
The authors confirm that there was no use of artificial intelligence (AI)-assisted technology for assisting in the writing or editing of the manuscript and no images were manipulated using AI.
Supplementary data
Supplementary material to this article can be found online at https://dx.doi.org/10.25259/AJC_111_2024
References
- Ultrasensitive immunosensor for multiplex detection of cancer biomarkers carcinoembryonic antigen (CEA) and yamaguchi sarcoma viral oncogene homolog 1 (YES1) based on eco-friendly synthesized gold nanoparticles. Talanta. 2024;266:124934. https://doi.org/10.1016/j.talanta.2023.124934
- [CrossRef] [PubMed] [Google Scholar]
- Electrochemical peptide nucleic acid functionalized α-Fe2O3/Fe3O4 nanosheets for detection of CYP2C19*2 gene. Mikrochimica Acta. 2023;190:189. https://doi.org/10.1007/s00604-023-05781-4
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Design, synthesis, and biological evaluation of 4-(2-fluorophenoxy)-7-methoxyquinazoline derivatives as dual EGFR/c-met inhibitors for the treatment of NSCLC. European Journal of Medicinal Chemistry. 2024;263:115939. https://doi.org/10.1016/j.ejmech.2023.115939
- [CrossRef] [PubMed] [Google Scholar]
- Overview on therapeutic options in uncommon EGFR mutant non-small cell lung cancer (NSCLC): New lights for an unmet medical need. International Journal of Molecular Sciences. 2023;24:8878. https://doi.org/10.3390/ijms24108878
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Evaluation of colorimetric RT-LAMP for screening of SARS-coV-2 in untreated wastewater. The Science of the Total Environment. 2024;907:167964. https://doi.org/10.1016/j.scitotenv.2023.167964
- [CrossRef] [PubMed] [Google Scholar]
- Development and validation of a multiplex quantitative polymerase chain reaction assay for detecting and genotyping red sea bream iridoviral disease. Aquaculture. 2024;580:740358. https://doi.org/10.1016/j.aquaculture.2023.740358
- [CrossRef] [Google Scholar]
- Infection with human cytomegalovirus, epstein-barr virus, and high-risk types 16 and 18 of human papillomavirus in EGFR-mutated lung adenocarcinoma. Croatian Medical Journal. 2023;64:84-92. https://doi.org/10.3325/cmj.2023.64.84
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Sequence variation of the epstein-barr virus nuclear antigen 1 (EBNA1) gene in chronic lymphocytic leukemia and healthy volunteer subjects. Archives of Virology. 2023;169:1. https://doi.org/10.1007/s00705-023-05933-0
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Intelligent genetic decoding system based on nucleic acid isothermal amplification for non-small cell lung cancer diagnosis. Micromachines. 2023;14:647. https://doi.org/10.3390/mi14030647
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Highly selective, single-tube colorimetric assay for detection of multiple mutations in the epidermal growth factor receptor gene. The Journal of Molecular Diagnostics : JMD. 2023;25:313-9. https://doi.org/10.1016/j.jmoldx.2023.02.006
- [CrossRef] [PubMed] [Google Scholar]
- Molecularly imprinted polymers-aptamer electrochemical sensor based on dual recognition strategy for high sensitivity detection of chloramphenicol. Food Chemistry. 2024;437:137933. https://doi.org/10.1016/j.foodchem.2023.137933
- [CrossRef] [PubMed] [Google Scholar]
- Nitrite determination in food using electrochemical sensor based on self-assembled MWCNTs/AuNPs/poly-melamine nanocomposite. Food Chemistry. 2023;437:137773. https://doi.org/10.1016/j.foodchem.2023.137773
- [CrossRef] [PubMed] [Google Scholar]
- Enzymatic construction au NPs-rGO based MIP electrochemical sensor for adulteration detection of bovine-derived allergen in camel milk. Food Chemistry. 2024;436:137638. https://doi.org/10.1016/j.foodchem.2023.137638
- [CrossRef] [PubMed] [Google Scholar]
- “OR” logic gate multiplexed photoelectrochemical sensor for high-risk human papillomaviruses: “One pot” recombinase polymerase amplification and logic discrimination. Talanta. 2024;266:125090. https://doi.org/10.1016/j.talanta.2023.125090
- [CrossRef] [PubMed] [Google Scholar]
- Near-infrared light activated and hybridization chain reaction cascaded CRISPR/Cas12a system under the enhancement of Mn2+ for intracellular biosensing. Sensors and Actuators B: Chemical. 2024;398:134777. https://doi.org/10.1016/j.snb.2023.134777
- [Google Scholar]
- Engineering of DNA structures attached to magnetic particles for effective trans- and cis-cleavage in Cas12-based biosensors. International Journal of Molecular Sciences. 2023;24:4484. https://doi.org/10.3390/ijms24054484
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- CRISPR/Cas systems for the detection of nucleic acid and non-nucleic acid targets. Nano Research 2023:1-14. https://doi.org/10.1007/s12274-023-5567-4
- [CrossRef] [Google Scholar]
- CRISPR applications in cancer diagnosis and treatment. Cellular & Molecular Biology Letters. 2023;28:73. https://doi.org/10.1186/s11658-023-00483-4
- [CrossRef] [PubMed] [Google Scholar]
- A tandem DNA nanomachines-supported electrochemiluminescence assay for attomolar detection of miRNA at ambient-temperature. Chemical Engineering Journal. 2024;480:148161. https://doi.org/10.1016/j.cej.2023.148161
- [CrossRef] [Google Scholar]
- Ultrasensitive detection of genetic variation based on dual signal amplification assisted by isothermal amplification and cobalt oxyhydroxide nanosheets/quantum dots. Mikrochimica Acta. 2023;191:12. https://doi.org/10.1007/s00604-023-06097-z
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Rapid, ultrasensitive and non-enzyme electrochemiluminescence detection of hydrogen peroxide in food based on the ssDNA/g-C3N4 nanosheets hybrid. Food Chemistry. 2021;357:129753. https://doi.org/10.1016/j.foodchem.2021.129753
- [CrossRef] [PubMed] [Google Scholar]
- Iron oxide nanoparticles: A review on the province of its compounds, properties and biological applications. Materials (Basel, Switzerland). 2022;16:59. https://doi.org/10.3390/ma16010059
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Design and development of multi-functional graphitic carbon nitride heterostructures embedded with copper and iron oxide nanoparticles as versatile sensing platforms for environmental and agricultural applications. Ceramics International. 2023;49:20688-20698. https://doi.org/10.1016/j.ceramint.2023.03.200
- [CrossRef] [Google Scholar]
- Nanomaterial-based electrochemical sensing platform for the determination of olaparib. Electrochimica Acta. 2023;449:142198. https://doi.org/10.1016/j.electacta.2023.142198
- [CrossRef] [Google Scholar]
- A hydrothermal-calcination process with ammonium dihydrogen phosphate as restricted growth agent for the fabrication of magnetic Fe3O4/α-Fe2O3 heterogeneous nanosheets. Journal of Inorganic and Organometallic Polymers and Materials. 2024;34:1015-1027. https://doi.org/10.1007/s10904-023-02879-3
- [Google Scholar]
- Encapsulating ultrastable metal nanoparticles within reticular schiff base nanospaces for enhanced catalytic performance. Cell Reports Physical Science. 2021;2:100289. https://doi.org/10.1016/j.xcrp.2020.100289
- [CrossRef] [Google Scholar]
- Solid-solution alloy nanoclusters of the immiscible gold-rhodium system achieved by a solid ligand-assisted approach for highly efficient catalysis. Nano Research. 2020;13:105-111. https://doi.org/10.1007/s12274-019-2579-1
- [CrossRef] [Google Scholar]
- Ag decorated cuGaO2 nanosheets for enhanced ethylene glycol detection. Journal of Alloys and Compounds. 2024;970:172563. https://doi.org/10.1016/j.jallcom.2023.172563
- [CrossRef] [Google Scholar]
- DNAzyme-driven bipedal DNA walker and catalytic hairpin assembly multistage signal amplified electrochemical biosensor based on porous auNPs@Zr-MOF for detection of Pb2. Food Chemistry. 2024;435:137503. https://doi.org/10.1016/j.foodchem.2023.137503
- [CrossRef] [PubMed] [Google Scholar]
- Construction of nanoflowers-like au@ZnO/ZnS/SnO2 heterostructure for conductometric cyclohexanone detection. Sensors and Actuators B: Chemical. 2024;399:134754. https://doi.org/10.1016/j.snb.2023.134754
- [CrossRef] [Google Scholar]
- Magnetically induced self-assembly electrochemical biosensor with ultra-low detection limit and extended measuring range for sensitive detection of HER2 protein. Bioelectrochemistry (Amsterdam, Netherlands). 2024;155:108592. https://doi.org/10.1016/j.bioelechem.2023.108592
- [CrossRef] [PubMed] [Google Scholar]
- Sensitive and selective DNA detecting electrochemical sensor via double cleaving CRISPR Cas12a and dual polymerization on hyperbranched rolling circle amplification. Biosensors & Bioelectronics. 2023;224:115078. https://doi.org/10.1016/j.bios.2023.115078
- [CrossRef] [PubMed] [Google Scholar]
- Electrochemical detection of the p53 gene using exponential amplification reaction (EXPAR) and CRISPR/Cas12a reactions. Mikrochimica Acta. 2023;190:113. https://doi.org/10.1007/s00604-023-05642-0
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Aptamer-based electrochemical biosensor with S protein binding affinity for COVID-19 detection: Integrating computational design with experimental validation of S protein binding affinity. Sensors and Actuators B: Chemical. 2024;399:134775. https://doi.org/10.1016/j.snb.2023.134775
- [CrossRef] [Google Scholar]
- G-quadruplex functionalized nano mesoporous silica for assay of the DNA methyltransferase activity. Analytica Chimica Acta. 2015;879:34-40. https://doi.org/10.1016/j.aca.2015.04.023
- [CrossRef] [PubMed] [Google Scholar]
- An ultra-sensitive dual-signal ratiometric electrochemical aptasensor based on functionalized MOFs for detection of HER2. Bioelectrochemistry (Amsterdam, Netherlands). 2022;148:108272. https://doi.org/10.1016/j.bioelechem.2022.108272
- [CrossRef] [PubMed] [Google Scholar]
- Triptonodiol, a diterpenoid extracted from tripterygium wilfordii, inhibits the migration and invasion of non-small-cell lung cancer. Molecules (Basel, Switzerland). 2023;28:4708. https://doi.org/10.3390/molecules28124708
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Amplified electrochemical genotyping of single-nucleotide polymorphisms using a graphene–gold nanoparticles modified glassy carbon platform. RSC Advances. 2015;5:29285-29293. https://doi.org/10.1039/c5ra03794h
- [Google Scholar]
- Dual amplification of single nucleotide polymorphism detection using graphene oxide and nanoporous gold electrode platform. The Analyst. 2014;139:5192-99. https://doi.org/10.1039/c4an01171f
- [CrossRef] [PubMed] [Google Scholar]
- An electrochemical sensing method based on CRISPR/Cas12a system and hairpin DNA probe for rapid and sensitive detection of salmonella typhimurium. Sensors and Actuators B: Chemical. 2022;369:132301. https://doi.org/10.1016/j.snb.2022.132301
- [CrossRef] [Google Scholar]
- CRISPR/Cas12a-mediated interfacial cleaving of hairpin DNA reporter for electrochemical nucleic acid sensing. ACS Sensors. 2020;5:557-562. https://doi.org/10.1021/acssensors.9b02461
- [CrossRef] [PubMed] [Google Scholar]
- Magnetic self-assembled label-free electrochemical biosensor based on Fe3O4/α-Fe2O3 heterogeneous nanosheets for the detection of tau proteins. Bioelectrochemistry (Amsterdam, Netherlands). 2024;157:108678. https://doi.org/10.1016/j.bioelechem.2024.108678
- [CrossRef] [PubMed] [Google Scholar]
- Exploring the trans-cleavage activity of CRISPR-Cas12a (cpf1) for the development of a universal electrochemical biosensor. Angewandte Chemie (International ed. in English). 2019;58:17399-17405. https://doi.org/10.1002/anie.201910772
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- A CRISPR-Cas12a powered electrochemical sensor based on gold nanoparticles and MXene composite for enhanced nucleic acid detection. Sensors and Actuators B: Chemical. 2023;380:133342. https://doi.org/10.1016/j.snb.2023.133342
- [CrossRef] [Google Scholar]
- Enhanced chemiluminescence imaging sensor for ultrasensitive detection of nucleic acids based on HCR-CRISPR/Cas12a. Biosensors & Bioelectronics. 2022;212:114428. https://doi.org/10.1016/j.bios.2022.114428
- [CrossRef] [PubMed] [Google Scholar]







