Translate this page into:
Baicalin suppresses colorectal cancer cell proliferation, potentially via ARRDC4: Bioinformatics and experimental analysis
⁎Corresponding authors. wzhang@must.edu.mo (Wei Zhang), yueyinzi@126.com (Yinzi Yue)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
Colorectal cancer (CRC) is a malignant digestive tumor mostly prevalent among adolescents and has great proliferation potential. Baicalin is a flavonoid derived from the dried root of Scutellaria baicalensis, it has tumor-suppressing effects on various kinds of tumors. Although baicalin has been shown to regulate genes associated with CRC, its role in cancer proliferation remains poorly illustrated. In this work, we evaluated the effect of baicalin on the proliferative ability of three CRC cell lines using a cell counting kit-8 assay. Furthermore, we screened for differentially expressed genes in HCT-116 cells using baicalin-treated samples and control samples on a gene expression profile microarray. We identified 19 genes were identified significantly downregulated by baicalin, which may potentially play an indispensable role in the proliferation of HCT-116 cells. High-content screening evaluated whether silencing the 19 genes affected HCT-116 cell growth. The results showed that knocking down ARRDC4 exhibited the most potent inhibition of cell proliferation. ARRDC4 expression was knocked down by transfecting CRC cells with lentiviral shRNA. MTT assay, Caspase-Glo 3/7 assay and immunohistochemistry analysis were performed to explore the effects of baicalin on proliferation and apoptosis of CRC cells. Besides, RT-PCR showed that the expression levels of ARRDC4 mRNA decreased significantly in HCT-116 cells after baicalin treatment. In conclusion, these findings indicated that baicalin may attenuated the expression of ARRDC4 via regulating the levels of LCN2, Claudin-2 and GSTA4 to inhibit HCT-116 cell proliferation. The theoretical underpinnings for the investigation of the pharmacological effects of baicalin in the treatment of human CRC were also established in this study.
Keywords
Baicalin
Colorectal cancer
RNA-sequencing
Proliferation
Apoptosis
Arrestin domain-containing protein 4 (ARRDC4)
1 Introduction
Colorectal cancer (CRC), a heterogeneous disease of intestinal epithelial cells, is characterized by the accumulation of mutations and an immune response disorder. It is one of the three most common malignant tumors, making this fatal disease a serious public health problem (Li et al., 2021a, 2021b, 2021c). According to the GLOBO CAN 2020 cancer incidence and mortality assessment report released by the International Agency for Research on Cancer, it is found that among all types of cancer, there were >1.93 million new CRC cases, and 940,000 relevant deaths. It ranked third by new incidence rate (10%) and second by mortality rate (9.4 %) (Sung et al., 2021). Similarly, the incidence of CRC in China has also shown a significant upward trend, with around 555.4 million new cases and 286.1 million deaths in 2020 (Sun et al., 2020). Besides, CRC ranked third and fifth in incidence and mortality for tumors (Cao et al., 2021). The burden of CRC in China is expected to increase in 2022, leading the cancer situation in China to converge with that in the America (Xia et al., 2022). Although the incidence of CRC in China is lower than the world average, due to the relatively large population and gradually westernized lifestyle, China ranks first in the number of new CRC cases and related deaths (Feng et al., 2019). Thus, CRC is a public health problem that endangers the health of Chinese residents.
The development of CRC is a multi-factor, multi-stage and multi-gene process; age, heredity, intestinal homeostasis, diet structure, precancerous lesions, and other factors cause malignant lesions in the mucosal epithelium leading to CRC (Biller and Schrag, 2021). At present, the main clinical treatments for CRC are surgical resection, adjuvant radiotherapy, and chemotherapy (Dekker et al., 2019). Radical resection is the preferred treatment for early CRC. Surgery alone increases the five-year survival rate of patients by 50%, but there are still 30 % of patients with stage I and II colon cancer, who experience complications such as recurrence and postoperative metastasis. Meanwhile, patients with CRC who receive radical surgery at the metastatic stage have a very poor prognosis and survival rate (Roque-Castellano et al., 2020). National Comprehensive Cancer Network (NCCN) Guidelines recommend chemotherapy and the clinical application of targeted biological agents for stage III and IV CRC to improve outcomes (Messersmith, 2019). Fluoropyrimidine-based chemotherapy (e.g., 5-fluorouracil) has been applied to treat advanced CRC for more than half a century, and has become the first-line systemic chemotherapy for patients with CRC and postoperative patients (Vodenkova et al., 2020). However, most patients receiving chemotherapy eventually develop drug resistance, which is also a major cause of failure in CRC treatment (Piawah and Venook, 2019). Although chemotherapy and radiotherapy are important CRC treatment methods that can prolong the survival time and improve the quality of life of patients, they are limited by the slow treatment progress, lack of specificity, numerous side effects, and unsatisfactory treatment effect (Nozawa et al., 2020). Therefore, the identification of genes and cell processes involved in the pathogenesis of CRC and the search for drug targets related to CRC growth are crucial for developing effective targeted therapeutic drugs. Clinicians need a new, effective, and safe alternative or adjuvant therapy to prolong the survival and improve the quality of life of patients.
Traditional Chinese medicine is increasingly regarded as a new source of anti-tumor drugs because its multi-component and multi-target treatments usually cause few side effects. Huang Qin is the dried root of Scutellaria baicalensis (S. baicalensis, Huangqin in Chinese); it was first mentioned in Shen Nong Ben Cao Jing (Sheng Nong’s Herbal Classics). According to Chinese herbology, it has a variety of curative effects, such as clearing heat, drying dampness, purging fire, and removing toxins. Its root has been used for over 2000 years to treat jaundice, hemorrhage, hepatitis, hypertension, diarrhea, insomnia, and infections of the respiratory and gastrointestinal tracts (Zhao et al., 2016). So far, more than 2000 compounds have been identified in SB, and 132 of them have been isolated, including 100 flavonoids (Murch et al., 2004; Wang et al., 2018a, 2018b). The primary bioactive chemical elements of S. baicalensis (SB) are flavonoids and their glycosides, with baicalein, baicalin, wogonin, wogonoside, and oroxylin A constituting the majority of the root-specific 4′-deoxyflavones (also known as Scutellariae radix, S. radix) (Zhao et al., 2019). Baicalin, a flavone glycoside, is the most abundant and crucial bioactive ingredient extracted from the roots of S. baicalensis. Its pharmacological effects include clearing up free radicals, enhancing apoptosis, blocking calcium channels, and inhibiting aldose reductase (Wang et al., 2018a, 2018b). Baicalin is often used to treat pneumonia, infection and tumor in clinic because of its protective effect on the immune system, cardio-cerebrovascular system, digestive system and nervous system (Zhou et al., 2021; Srinivas, 2010).
Here, we focused on the anticancer effects of baicalin on CRC, investigated its main underlying molecular mechanisms, performed in vivo studies, and evaluated its potential clinical applications. To find relevant studies, we used international electronic databases, i.e., MEDLINE (through PubMed), SciFinder, Google Scholar, and AMED, and China Journals Full-Text Databases (i.e., CNKI, VIP, and Wanfang) from their inception to April 2022, without language restriction (but most relevant studies were published in English and Chinese) using the terms baicalin, colorectal carcinoma, colorectal cancer, and colon carcinoma. Several reports have proved the high activity of baicalin on a variety of CRC cells through various targets and mechanisms. Baicalin exerts an anti-tumor effect by inducing cell cycle arrest (Yang et al., 2013; Dou et al., 2018; Xu et al., 2017) and apoptosis (Chen et al., 2012) through different mechanisms, such as inhibiting the Wnt signaling pathway (Jia et al., 2019), downregulating the SP1 transcription factor (Ma et al., 2019), reducing the expression of c-Myc and oncomiRs (Tao et al., 2018), upregulating Baculoviral IAP repeat-containing protein 3 (BIRC3) expression (Phan et al., 2020), and downregulating circMYH9 and HDGF and upregulating miR-761 (Zhang et al., 2021). Additionally, baicalin blocks cell migration, invasion, and metastasis through various signaling pathways in CRC cells, such as the Ras/Raf/MEK/ERK, TLR4/NF-κB, and TGFβ/Smad pathways (Wang et al., 2018a, 2018b; Song et al., 2022; Yang et al., 2020). However, the definite effect and molecular mechanism of baicalin on CRC cell proliferation have not been elucidated.
High-content screening (HCS) is a fluorescence labeling technique allowing the quantification of intracellular indicators by microscopic imaging, and the analysis of intracellular biological activity through image analysis (Mattiazzi et al., 2016). At present, with the continuous development of biotechnology, genomics, epigenetics, transcriptomics and proteomics technology screening has been applied to daily basic research work. However, the functional verification of screened differentially expressed genes (DEGs) remains labor-intensive and time-consuming (Mattheakis, 2013). Thanks to its high throughput and high spatio-temporal consistency, HCS can greatly help researchers with the functional verification of DEGs (Fraietta and Gasparri, 2016). HCS can yield high-quality morphological structure information of cells through a microscopic imaging system, provide intuitive morphological information on cell proliferation, and automatically identify, as well as qualitatively and quantitatively analyze different cell characteristics through software analysis.
The present study documents the effect of baicalin on the proliferation of CRC cells in vitro and in vivo. To explore the underlying mechanism and the biological activities exerted by baicalin, we performed microarray analysis, high-content short hairpin RNA (shRNA) screening, and other assays. Our findings shed light on the molecular processes of CRC and suggest a unique CRC treatment approach. Our results indicate that baicalin is promising CRC treatment candidates.
2 Materials and methods
2.1 Chemicals and reagents
Baicalin (MW:446.37, Fig. A.1, ≥98% purity) was purchased from Shanghai yuanye Bio-Technology Co., Ltd (catalog no.:N15GB167969). Caspase-Glo® 3/7 Assay was obtained from Promega (Madison, WI, USA, #G8091). Antibodies against ABP1 (#16338–1-AP), DAPP1 (#14722–1-AP), NLRP3 (#19771–1-AP), GSTA4 (17271–1-AP) and LCN2 (26991–1-AP) were purchased from Proteintech (Chicago, USA). The antibody against GGT5 (#ab283267) was purchased from Abcam (USA). The antibody against Claudin-2 (#48120) was purchased from Cell Signaling Technology (Boston, MA, USA). ANKRD63 polyclonal antibody (#PA5-49152) and ARRDC4 polyclonal antibody (# PA5-36518) were purchased from Invitrogen (Carlsbad, USA). Cell counting kit-8 kit (#CK04, Dojindo, Kumamoto, Japan), apoptosis kit (#88–8007, eBioscience, California, USA), BCA protein assay kit (Beyotime, Shanghai, China, #P0010S), radioimmunoprecipitation assay (RIPA) buffer (Beyotime, #P0013B), endoFree midi plasmid kit (#DP118-2, TIANGEN, Beijing, China) and enhanced chemiluminescent (ECL) plus reagents (#M3121/1859022, Thermo Fisher Scientific, Waltham, MA, USA) were used in the study.
2.2 Cell culture and baicalin treament
The two human CRC cell lines HCT-116 and HT-29 were obtained from the Cell Bank of the Chinese Academy of Science Type Culture Collection (Shanghai, China). RKO cells were obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA). The cell lines were all cultured in RPMI 1640 media (Thermo Fisher Scientific,Waltham, MA, USA), McCoy’s 5A and Eagle’s Minimum Essential Medium (Sigma Aldrich, Germany), respectively. All mediums were supplemented with 10% fetal bovine serum (#VS500T, Ausbian, Australia), 100 μg/mL streptomycin (Gibco, CA, USA), and 100 units/mL penicillin (Gibco, CA, USA). Cells were all incubated in a humidified incubator under a 5% CO2 atmosphere at 37 °C. The indicated concentrations of baicalin were achieved by adding appropriate amounts of stock baicalin solution to the culture medium. The cells then incubated for a certain time. The dimethyl sulfoxide solution that did not contain baicalin was used the blank reagent.
2.3 Cell viability and drug sensitivity assays
Cells were seeded into 96 wells (4 × 103 cells/well) and incubated for 24 h at 37 °C for adhesion, then treated with baicalin at different concentrations or physiological saline (control) for 48 h. Then, the CCK-8 assay was performed to evaluate cell viability (%). A non-linear regression model was used to calculate half maximal (50%) inhibitory concentration (IC50) values with a sigmoidal dose–response curve in GraphPad Prism 8. Three independent experiments were performed.
2.4 RNA sequencing and quantitative real-time PCR (qRT-PCR)
Total RNA was extracted using Trizol reagent from baicalin-treated cells and was reversely transcribed into cDNA with a Fast King RT kit. The primers are listed in Table 2. RNA purity was confirmed using a NanoPhotometer® spectrophotometer (IMPLEN, CA, USA). Next, we screened the RNA to identify DEGs with an Agilent 2100 bioanalyzer (Agilent, Santa Clara, CA, USA). Three ARRDC4 knockdown samples and three negative control samples were used for microarray analysis using the human GeneChip PrimeView array. The cDNA library construction and purification as well as transcriptome sequencing were conducted based on the Shanghai Genechem Company’s instructions. The gene or exon expression levels were normalized as the number of reads per kilobase per million mapped reads (RPKM) (Mortazavi et al., 2008). DEGs were defined as having a P value below 0.05 and an expression fold change above 2. The GoTaq® qPCR Master Mix was applied for qRT-PCR. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as an internal control.
2.5 Construction of ARRDC4 gene recombinant lentiviral vectors and transfection
The cell lines with stably knocked-down genes were obtained as previously described (Dave et al., 2014). The knockdown efficiency was confirmed by real-time PCR and western blot, and the shRNA sequences were shown in Table 1. The interference sequence-containing single-strand DNA oligonucleotide was then created, and annealing was used to create the double-strand DNA. After digestion by an enzyme, the two ends of the oligonucleotide were then immediately joined to the lentiviral vector. The prepared Escherichia coli cells received the ligated products. The positive recombinants were then located by PCR, and their sequences were used to verify them and remove their plasmids. Lentiviral vector DNA and packaging vectors were transfected in 293 T cells (Chinese Academy of Science Type Culture Collection, Shanghai, China). We purchased the lentivirus with the ARRDC4 gene overexpression vector (Shanghai Genechem Co., Ltd., Shanghai, China). The lentiviral vector system was composed of GV493, a pHelper 1.0 vector and a pHelper 2.0 vector prior to packaging, and the full-length ARRDC4 gene was encoded into the GV493 vector after being labeled with the enhanced green fluorescence protein (eGFP). The empty GV493 lentiviral vector was used as the shRNA control (shCtrl). After 48 h of culture, supernatants containing the lentiviruses, including shARRDC4 and shCtrl, were harvested and purified. Three days later, GFP expression was quantified by fluorescence microscopy (BX51; Olympus, Tokyo, Japan). Cells with stable eGFP expression were harvested for the following analysis.
Primer name
shRNA sequences
ARRDC4-RNAi(97573-1)
GCATTATCAGAGTGGACTATT
ARRDC4-RNAi(97574-1)
CGCTTTCAACTTCCATCTGAA
ARRDC4-RNAi(97575-1)
CGGTATTATGAAACCAAGAAA
COL24A1-RNAi(97582-1)
GCTGCTATTCAAGCCTTGATT
COL24A1-RNAi(97583-1)
CGCCCAAACTATTTGCTGAAA
COL24A1-RNAi(97584-1)
GCAGACAAATACCAACCTGAA
GASK1B-RNAi(97585-1)
CTTGATAAAGTGTATTGGGAA
GASK1B-RNAi(97586-1)
GACAGGAGTGAAGATAACTTA
GASK1B-RNAi(97587-1)
CCAAGATGGCACTCTTTGATT
CIBAR2-RNAi(97567-1)
CATTATGACACTCGGCTGCTT
CIBAR2-RNAi(97568-1)
GTGACTTTGTAACTATTGAGA
CIBAR2-RNAi(97569-1)
CAGCTCATCGACTTTGCCAAC
EMC9-RNAi(97579-1)
GACTTTGACTGCCACCTTGAT
EMC9-RNAi(97580-1)
CCCTAAGGATAAGAACTTAGT
EMC9-RNAi(97581-1)
GCTGGTTACTACCATGCCAAT
ANKRD63-RNAi(97564-1)
TGGCCCGGAGTTAGAGGCCAA
ANKRD63-RNAi(97565-1)
CGGCTCTGGCCGCCTGGGTTT
ANKRD63-RNAi(97566-1)
TTGGCCCGCTTCGTGTTGGAT
C2orf78-RNAi(97570-1)
GCTCGACCTGATTCTACTAAC
C2orf78-RNAi(97571-1)
GATAGGAAGCCAGGTCTATTA
C2orf78-RNAi(97572-1)
GTCTATGACAGCCCAGTATTA
FOXL2NB-RNAi(97576-1)
GAAGCTTCACTGTGTCTATTA
FOXL2NB-RNAi(97577-2)
AGCGGGAGAGAATAGAGCTTG
FOXL2NB-RNAi(97578-1)
TTCACATGGCTGTCCGGCATT
METTL7A-RNAi(97588-2)
ATAGTGTGAGCTGGCAGTTAA
METTL7A-RNAi(97589-1)
GTGAGGTTCACTGTGATATAC
METTL7A-RNAi(97590-1)
CACCTGCAGTTTGAGCGCTTT
PCDHGB2-RNAi(97714-1)
CGACAAAGGATGATTTGGATT
PCDHGB2-RNAi(97715-1)
CCACCTTAATGACAACGAGTA
PCDHGB2-RNAi(97716-1)
GCAGTAATTGTGCAGGATATA
PRR19-RNAi(97711-1)
CAACAGCGTTTGACTTGTTAA
PRR19-RNAi(97712-1)
GAAGCAACAAGGGACAAAGGA
PRR19-RNAi(97713-1)
CTTCTGGATTAATAGCCCTGA
RBBP8NL-RNAi(97669-1)
GCAGTACAGAAGATGAAGACA
RBBP8NL-RNAi(97670-1)
GCAGGACTGTGCCCTAGACAA
RBBP8NL-RNAi(97671-1)
GAAGGAAGAGAACGAGACCTT
RCBTB2-RNAi(97666-1)
CACAAACTGCTGTGGCTGTTT
RCBTB2-RNAi(97667-1)
GACCATAGCATGTGGGCAGAT
RCBTB2-RNAi(97668-1)
AGTGGCAGCTTTGCAAGGCAT
DDX25-RNAi(97702-1)
GCCTAGCTCCTACTTATGAAT
DDX25-RNAi(97703-1)
GTGAACTTTGATCTCCCTGTA
DDX25-RNAi(97704-1)
CTGAACAACATCCGGCAATAT
PRR16-RNAi(97699-1)
CGAGAACGAGTTCGGTTTAAT
PRR16-RNAi(97700-1)
TGACTGTGATACCCGGTATAA
PRR16-RNAi(97701-1)
CTTCTACGAAATGGAGGCTTA
ZNF780A-RNAi(97696-1)
CGGTATCCAGATTTGGAGTTA
ZNF780A-RNAi(97697-1)
GCCCTGGATGGTTGTAAGGAA
ZNF780A-RNAi(97698-1)
ACGGTATCCAGATTTGGAGTT
PRXL2B-RNAi(97708-2)
GGATGAGAGCAAGCAGCTTTA
PRXL2B-RNAi(97709-1)
AAGCAGCTTTACAAGGAGCTA
PRXL2B-RNAi(97710-1)
GCTCTACCTGGATGAGAGCAA
SPOCK3-RNAi(97660-1)
TCAGGCATGTGTCTTAGGAAA
SPOCK3-RNAi(97661-11)
AAGCATTTACCTTGATAAGAA
SPOCK3-RNAi(97662-1)
ACTCACTTGGCTGGATGTTTA
SMIM32-RNAi(97663-1)
CACCTACCTGCTGCTCTTCTT
SMIM32-RNAi(97664-1)
GGTCAAGGCAGAAGGCGCTTT
SMIM32-RNAi(97665-11)
CGGCGACATATTCAACGCCAC
Genes
Forward (5′-3′)
Reverse (5′-3′)
COL24A1
TGCTGTGGATCTCACAAACCA
TTTGCCTCACTGATGCGATGA
NUDT4B
TTCTCATTCACAGCATTAG
ATAGCCATATCAGGTAGC
PRR19
CACCCAGTTCCCCAGAGTT
TTGTCCCTTGTTGCTTCCT
ARRDC4
GCAGACATATTTGGCTAGTGG
GATGGAGTAACAGGTGGGATT
METTL7A
ACAGATGGCAAGCAAGAAGCG
TTGGGGTTGGGGTCAATACAG
PRXL2B
AGGCTTCAAGCGGTACAACA
GGCCTTGGCAGCCACATC
SPOCK3
GACTCACTTGGCTGGATGT
CTTTACCCCTTGCCGCTTC
GASK1B
AGGACCCTGCCGTCTGTGA
GCTTGGGTACTCGGCCATT
ANKRD63
TTTGCGCCGACGCTCCACA
GCCTCTAACTCCGGGCCACTCT
C2orf78
CTGTGGCTTACCCTGCTCG
AACTGGGTTGGGTTGCTTT
CIBAR2
CAAAGCGGTGGAGGTGTATT
AACTCCTTGCATCTTGGCTC
EMC9
CAGTTGGCTCCTCACTCCTCT
CACCATCTGCCGTGACTCTT
FOXL2NB
CAGCCCTGGTGAAGAAGAG
CCGGACAGCCATGTGAAGG
PCDHGB2
GTTGCCCTTCCTGTTGTCT
CGCATATCTGTTCTCGGTCT
RBBP8NL
AGCGGCAGCAGGAGTTCGA
TCAGCCCGTTCATCTCGTTG
RCBTB2
GAACCTCGGAGACTGGATT
TATGACAGGGCACTAAACC
SMIM32
CCACGGTCAAGGCAGAAGG
AGCAGGTAGGTGGGCAGGTC
DDX25
GCGAATCATCCCTGACCCT
TCCCATCCCGAAACCTCTG
PRR16
AAACGGACACGCTGAATAG
TACAGGATTGGCAGTTGGA
ZNF780A
TCAGGAGGAGTGGGAGTGC
TCTGGATACCGTCTGCTTG
GAPDH
TGACTTCAACAGCGACACCCA
CACCCTGTTGCTGTAGCCAAA
2.6 High-content screening and cell growth curve analysis
Using HCS, we discovered the effects of 19 putative target genes on the proliferation of HCT-116 cells were discovered (Supplementary Material). In brief, HCT-116 cells were seeded at 2000 cells/well in 96-well plates and transfected with shRNA lentivirus targeting candidate target genes or negative control lentivirus. The expression of GFP was quantified by fluorescence microscopy. Cells were collected for further experiments when they reached 80% confluence. The system achieves high throughput screening through an automatic Celigo cell imaging analyzer (Nexcelom, USA). In the HCS proliferation assay, the detection target was cells expressing GFP after infection with the virus. The Celigo cell imaging analyzer recognizes cells with green fluorescence and records pictures. Then, the images were analyzed and processed by software to calculate the number of cells in various groups in the orifice plate. After 5 days of continuous reading, the cell growth curves were drawn. A fold change of the proliferation ratio of two or more suggested slowing-down cell proliferation, which was enough to examine the effect of RNAi lentivirus infection on cell proliferation (Mandavilli et al., 2018).
2.7 Western blot
After harvesting baicalin-treated cells, cell pellets were washed with ice-cold phosphate-buffered saline (PBS) and lyzed with RIPA buffer. Total proteins were quantified using a bicinchoninic acid (BCA) assay kit. After the total lysates were treated with a sodium dodecyl sulfate (SDS)-loading buffer at 100 °C, sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) was performed. The proteins were then transferred to polyvinylidene difluoride (PVDF) membranes, which were blocked with 5% dried skimmed milk, followed by overnight incubation with the specific primary antibodies. Finally, the HRP-conjugated secondary antibody were added and protein expression was quantified using the ECL plus reagents.
2.8 Apoptosis assay
Before the apoptosis assay, cells were seeded in a six-well plate and treated with baicalin for 48 h. The cells (attached and floating) were collected by centrifugation. To perform the apoptosis assay, these cells were resuspended in PBS and adjusted to 1 × 106 cells/mL. Next, 10 μL of Annexin V-APC staining solution (eBioscience, California, USA) was added to the 300 μL cell suspension for staining and the samples were incubated for 10–15 min at room temperature in the dark. Finally, the cells were counted and analyzed using a FAC Sort Flow Cytometer (BD, CA, USA).
2.9 Caspase-Glo 3/7 assay
The caspase activity was detected using a Caspase-Glo® 3/7 Assay Kit (Promega, WI, USA) (Li et al., 2021a, 2021b, 2021c). The cells were seeded in 96-well plates, incubated overnight and then for 24 h with the compounds at various concentrations according to the manufacturer’s instructions. The luminescent signal resulting from caspase 3/7 activity was detected with an EnVision® Multilabel Reader (PerkinElmer, Thermo Fisher Scientific, Shanghai, China).
2.10 In vivo tumor growth assay
Female BALB/c nude mice (4 weeks old, 25–30 g) from Shanghai Lingchang Biotechnology Co., Ltd (SCXK(hu)2018–0003), were maintained under a specific pathogen -free environment according to the Regulations for the Care and Use of Laboratory Animals. All experiments were in accordance with the guidelines of China for animal care, which confirm to the internationally accepted principles in the care and use of experimental animals. This study obtained approval from the Ethics Committee of Animal Experiments of Suzhou TCM Hospital Affiliated to Nanjing University of Chinese Medicine. A xenograft model was established by subcutaneously injecting HCT-116 cells and ARRDC4 knockdown HCT-116 cells under logarithmic growth into the right flank of nude mice (1 × 107/mL cells/mouse). Next, we monitored tumor growth. The treatment was initiated when the volume of the tumor reached approximately 100 mm3 (day 1). Mice were randomly divided into four groups of six mice each. The vehicle and control groups were treated with baicalin (50 or 100 mg/kg/day) for 14 consecutive days. The tumors were measured twice a week using microcalipers and the tumor volume (V) was calculated using: V = (Length × Width × Width)/2. All mice were sacrificed at the end of the experiment or when the tumor volume reached 800 mm3 to avoid unnecessary animal suffering, and their tumors were weighed and photographed.
2.11 Hematoxylin and eosin (HE) staining of subcutaneous tumor tissue sections
Subcutaneous tumor tissues were fixed in 4% polyformaldehyde, then washed thoroughly with water, dehydrated with methanol at different concentrations, and transparentized with dimethylbenzene. They were then embedded in paraffin, sectioned, and subjected to HE staining. The pathological changes of subcutaneous tumor tissues were observed by optical microscopy and photographed (×200, ×400).
2.12 Immunohistochemistry (IHC) analysis
We performed IHC on the paraffin sections of tumor tissues using a kit according to the manufacturer’s instructions. Briefly, the tissue sections (4 μm) were dehydrated and subjected to peroxidase blocking. After blocking, the sections were incubated with the Ki67 antibody for overnight incubation at 4 °C. Next, we incubated the slides with the horseradish peroxidase-conjugated secondary antibody and stained them with 3,3′-diaminobenzidine (DAB). Ki67-positive cells appeared brown under a microscope, and images were recorded. Image J image analysis software was used for semi-quantitative analysis.
2.13 Statistical analysis
All quantitative data were expressed as mean ± standard deviation and analyzed using Student’s t-test or analysis of variance (ANOVA). P-values < 0.05 indicated statistically significant differences. All statistical analyses were conducted using SPSS (version 13.0) and visualized with GraphPad Prism 8 (GraphPad Software, La Jolla, CA, USA).
3 Results
3.1 Cytotoxicity of baicalin in CRC cells
To study the effect of baicalin on CRC cells, we treated three cell lines with various baicalin doses for 48 h. The calculated IC50 values ranged from 0 to 1000 μM (Fig. 1B) and indicated that HCT116 cells were more sensitive to baicalin than the other lines. Baicalin markedly inhibited HCT-116 cell proliferation and reduced cell growth by more than 30% after 48 h (Fig. 1C and D). Cells treated with 90 µM baicalin were more significantly inhibited than untreated cells. Based on the pre-experiment, baicalin dose-dependently inhibited HCT-116 cell proliferation (1–40 μM). Therefore, we used baicalin concentrations of 80 µM for RNA sequencing and 10 µM for the HCS experiment.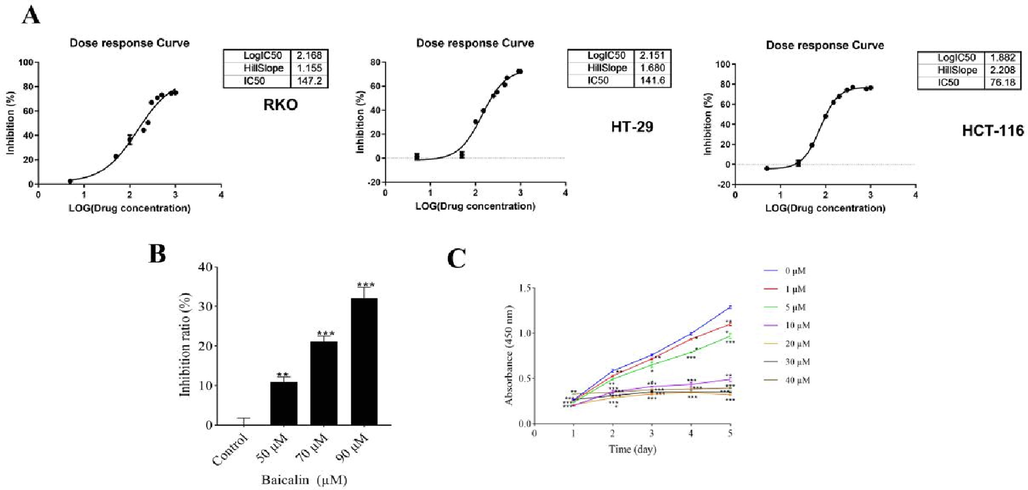
The inhibitory effect of baicalin was investigated based on the selected cell line. (A-C) Using an MTT assay, the effects of baicalin on the viability of human RKO, HT-29 and HCT-116 cells treated with baicalin (0–1000 μM) for 48 h are illustrated.
3.2 Identification of ARRDC4 as a critical gene promoting CRC proliferation
In order to study the underlying mechanisms of the anti-proliferation effect of baicalin on HCT-116 cells, we conducted a transcriptomic analysis. We treated HCT-116 cells with baicalin (80 μM) or negative control for 48 h, performed RNA sequencing. and identified 9601 DEGs were identified (4817 upregulated and 4784 downregulated) (Fig. 2A and 2B; Tables A1-A3). Among them, 114 had a fold change larger than 2 (Table A4). We then conducted a GO enrichment analysis on these genes to further characterize the mechanism of baicalin’s effect on CRC cell proliferation (Fig. 2C). To identify novel oncogenes, we focused on 19 downregulated genes that have not been extensively investigated for potential association with CRC.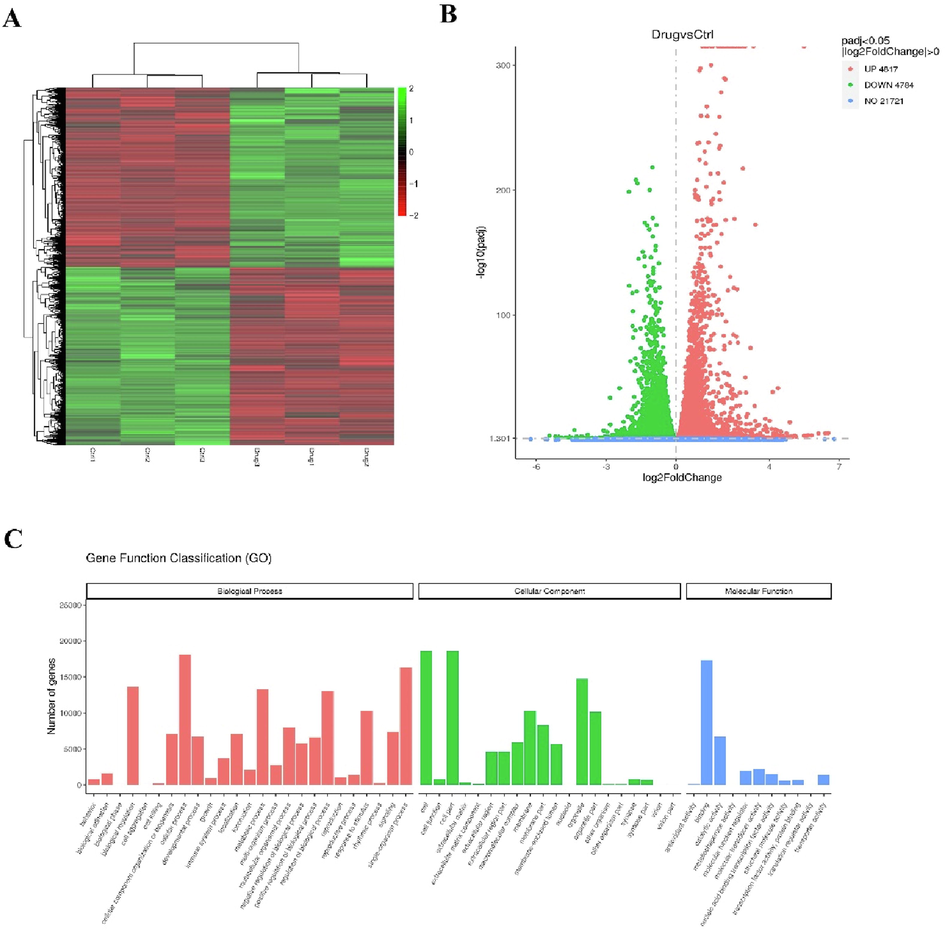
Agilent mRNA microarray identified expression profiling of baicalin-treated HCT-116 cells for 48 h by using RNA sequencing technology. (A) Heat map showing gene expression profiles. Each row represents a gene and each column represents a sample. Red indicates high expression, whereas green indicates low expression. (B) Volcano plot representing differentially expressed genes between baicalin-treated and baicalin-untreated HCT-116 cells. (C) GO enrichment (BP, CC and MF) of host genes of significantly differentially expressed circRNAs. X-axis represents the enriched GO term ordered by BP, CC and MF. Y -axis indicates the number (Left) of the genes in the corresponding terms.
We confirmed these DEGs by qPCR in accordance with the RNA-seq sample preparation guidelines. Except for NUDT4B, 19 genes were compatible with the sequencing data (Fig. 3).Next, we performed HCS on cells infected with a mix of three targets of lentivirus carrying 19 genes to identify genes involved in cell proliferation. Silencing each of the 19 candidate genes in HCT-116 cells revealed that silencing ANKRD63, SMIM32, and ARRDC4 strongly inhibited cancer cell growth (Fig. 4A, B and C). These results suggested that ARRDC4 might play a role in CRC development (Fig. 4D). ARRDC4 has been recently extensively studied in prostate cancer (Choueiri et al., 2015), but few studies have documented its role in CRC. Therefore, we assessed its anti-proliferation properties and the underlying mechanisms.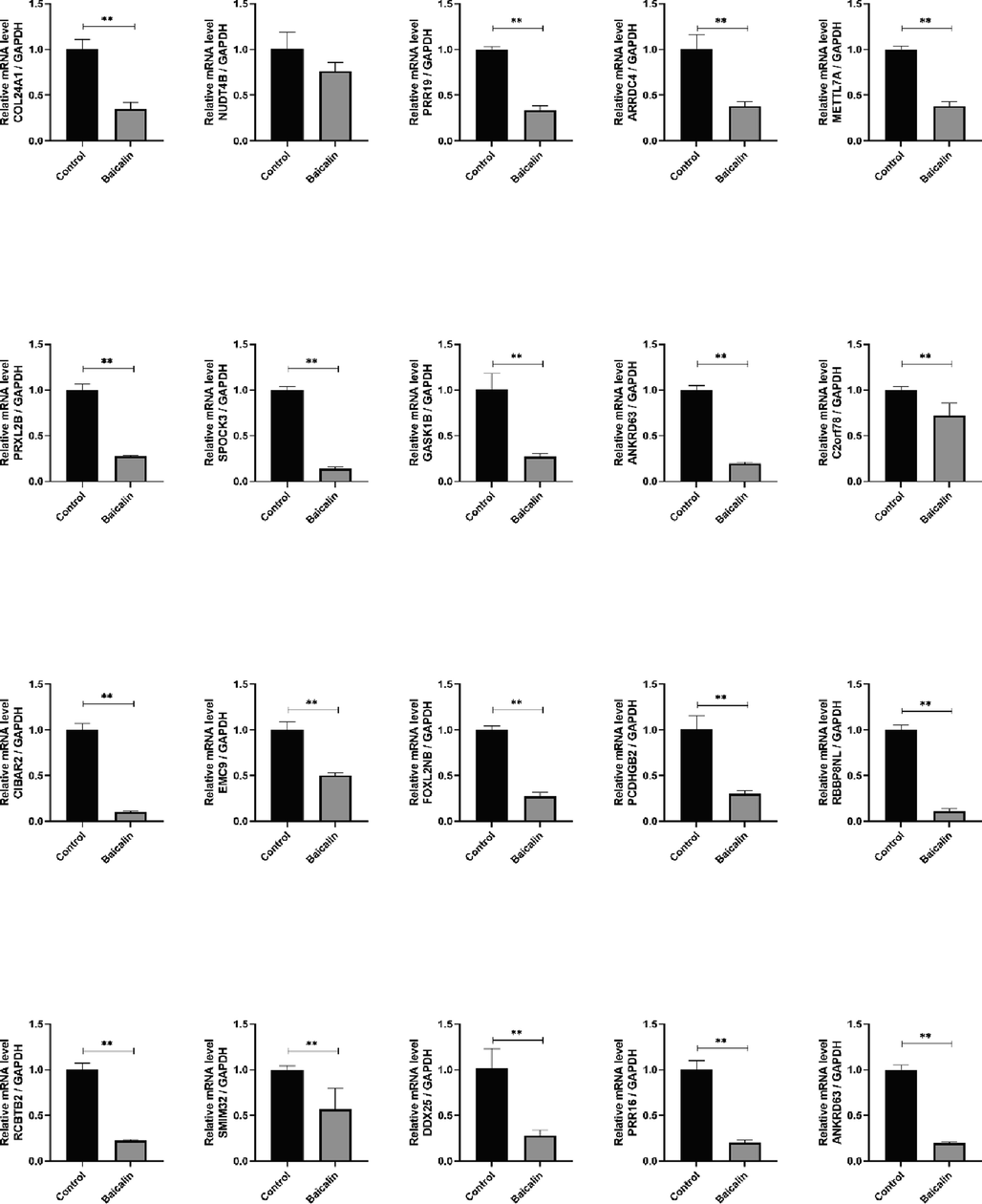
RNA-seq transcriptome profiling identifies 19 genes (except NUDT4B) as potential targets for colorectal cancer proliferation. Values were expressed as mean ± SD. Compared with the control group, ##P < 0.01.
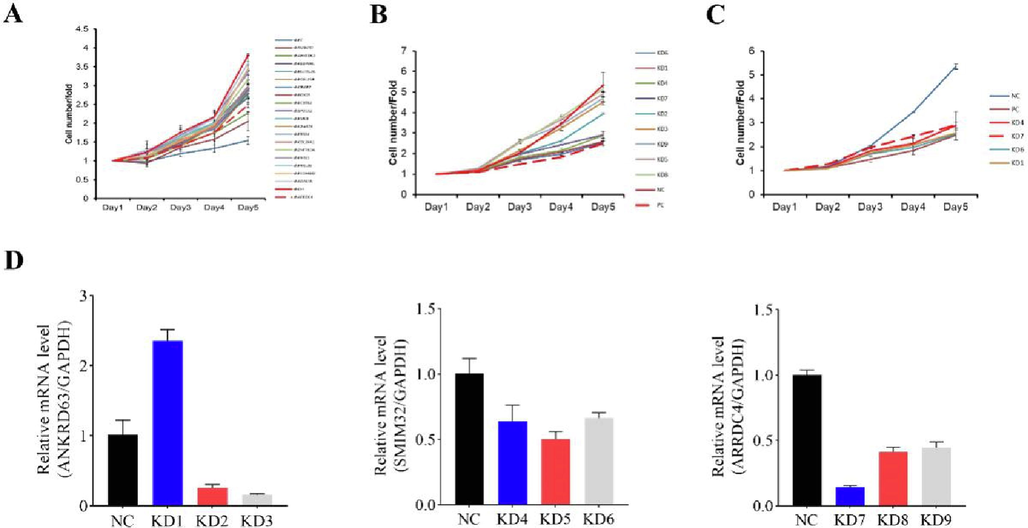
High-content screening identified ARRDC4 as a critical gene in promoting colorectal cancer proliferation. (A) A total of 19 genes were selected for validation by high-content screening. (B-C) Representative cell proliferation count of high-content screening for ANKRD63, SMIM32 and ARRDC4. (D) Knockdown efficiency of shRNA targeting ANKRD63, SMIM32 and ARRDC4. Cells were infected with lentivirus containing shRNA, RNA was collected and knockdown efficiency was determined by Real-time PCR.
3.3 Knocking down ARRDC4 strengthened the baicalin-mediated inhibition of HCT-116 cell proliferation
To determine the role of ARRDC4 in CRC cell proliferation, we established a stable ARRDC4 knockdown HCT-116 cell line using a lentiviral delivery system (Fig. 5A), confirmed the downregulation of ARRDC4 protein and mRNA levels in this cell line (Fig. 5B and C). The role of ARRDC4 in baicalin-mediated activities was further clarified, to down-regulation of ARRDC4 and then investigate baicalin’s activity. HCT-116 cells were knocked down ARRDC4 for 72 h, then the cells were treated with 10 μM baicalin, and cell viability was assessed by the CCK-8 assay (Fig. 5D). The inhibition of CRC cell proliferation by baicalin was dramatically strengthened in ARRDC4-low expression cells, as evaluated by Celigo and cell growth curve analysis utilizing a fluorescent imaging equipment.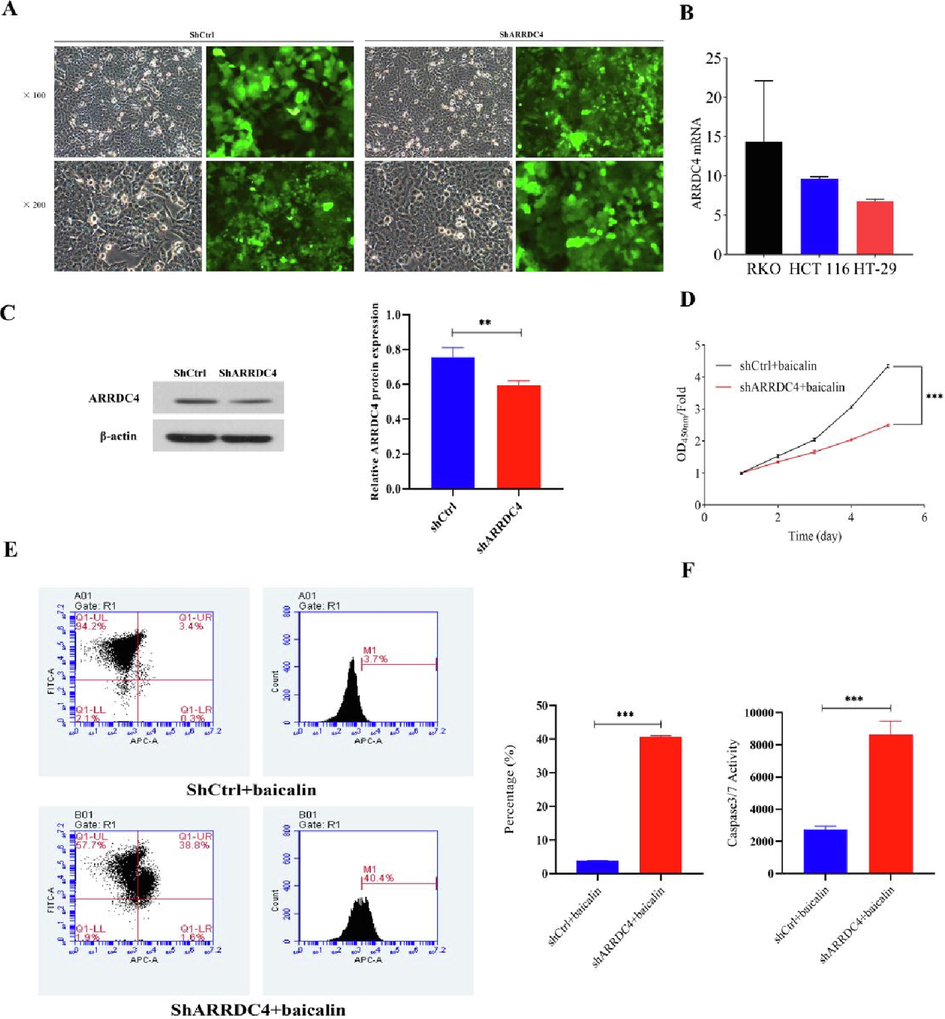
Knockdown of ARRDC4 inhibits HCT-116 cell proliferation in vitro. (A) Infection efficiency was determined at 48 h after infection of lentivirus shARRDC4 or shCtrl in HCT-116 cells. Original magnification × 100, ×200. (B) ARRDC4 mRNA in CRC cells was measured by q-PCR, and normalized to GAPDH. (C) ARRDC4 protein expression was analyzed by western blot analysis in HCT-116 infected with shARRDC4 or shCtrl. **P < 0.001 shARRDC4 vs shCtrl. (D) Cell growth curve analysis comparing ARRDC4 knockdown (shARRDC4 + baicalin) with negative control (shCtrl + baicalin) HCT-116 cells. (E) Knocked down of ARRDC4 accelerated the effect of baicalin on apoptosis. Cell apoptosis was analyzed was analyzed by flow cytometry. (F) Activity of cleaved caspase-3/7 was determined in HCT-116 cells transfected with shCtrl + baicalin or shARRDC4 + baicalin (***P < 0.001).
3.4 Knocking down ARRDC4 accelerated the baicalin-induced apoptosis of HCT-116 cells
To determine whether ARRDC4 regulated apoptosis in the baicalin-mediated CRC cell proliferation inhibition, we performed Annexin V-APC labeling and flow cytometry analysis. We observed 5% of apoptosis in HCT-116 cells treated with 10 μM baicalin, but the ARRDC4-knockdown cells had a significantly higher apoptotic ratio (Fig. 5E). Morever, compared with the shCtrl + baicalin group, the cleaved caspase-3/7 activity was statistically significantly increased in the shARRDC4 + baicalin group (Fig. 5F). These findings indicated that the baicalin-mediated induction of apoptosis in HCT-116 cells was strongly connected to its effect on ARRDC4.
3.5 Baicalin restrained CRC xenograft growth in vivo
After the in vitro exploration of ARRDC4′s role, we assessed whether knocking down ARRDC4 affected the in vivo formation of tumors. First, we established an in vivo tumor xenograft model by subcutaneously injecting sh ARRDC4-HCT-116 (KD groups) or HCT-116 cells (NC groups) into BALB/c nude mice (Fig. 6A). As shown in Fig. 6B-D, the KD + baicalin groups had significantly smaller and lighter tumors than the NC + baicalin (50 mg/kg) group. At the end of the experiment, the tumors of the vehicle group reached around 500 mm3, while those of the KD + baicalin group averaged 300 mm3. Overall, baicalin (50 and 100 mg/kg) inhibited tumor growth on both NC and KD groups, but the effect was weaker on NC groups. HE staining (Fig. 7A) revealed that the tumors in the NC + baicalin groups presented different degrees of degenerative lesions, and some areas showed obvious cell necrosis. These lesions were more obvious in the KD + baicalin groups, and vacuolar degeneration and necrosis could be seen in some cells. Unlike the NC + baicalin groups, the KD + baicalin groups showed large necrotic areas and cells with more nuclear shrinkage, fragmentation, and dissolution. Subsequently, we investigated effects of baicalin on cell proliferation markers in vivo by IHC analysis. The KD + baicalin groups had substantially lower levels of Ki-67, a well-known cell proliferation marker, than the NC + baicalin groups (Fig. 7B and C). Furthermore, the Western blot analysis revealed that knocking down ARRDC4 in groups treated with baicalin dramatically reduced the protein expression of p21 (Fig. 7D). Contrarily, ARRDC4 knockdown with baicalin treatment might upregulate Cyclin D1 protein expression (Fig. 7D). The results demonstrated that knocking down ARRDC4 in groups treated with baicalin might eventually suppress tumor growth.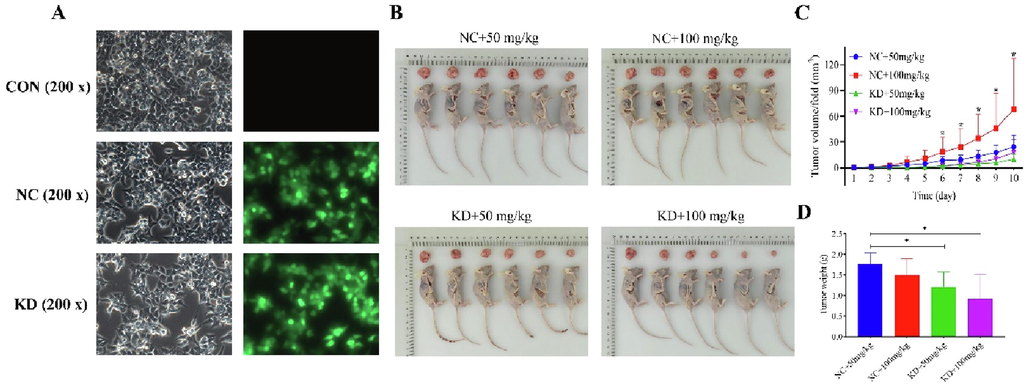
Knockdown of ARRDC4 with baicalin inhibited HCT-116 xenograft growth in vivo. (A) Morphology of HCT-116 cells after silencing ARRDC4. (B) HCT116 cells expressing sh Ctrl or sh ARRDC4 were injected into the right flank of nude mice for xenograft model, then after 2 weeks, the tumors of nude mice from the NC + baicalin (50 mg/kg), NC + baicalin (100 mg/kg), KD + baicalin (50 mg/kg) and KD + baicalin (100 mg/kg) groups were observed and tumor samples were separately collected. (C) Tumor volume growth curves and (D) tumor weight change for subcutaneous xenografts.
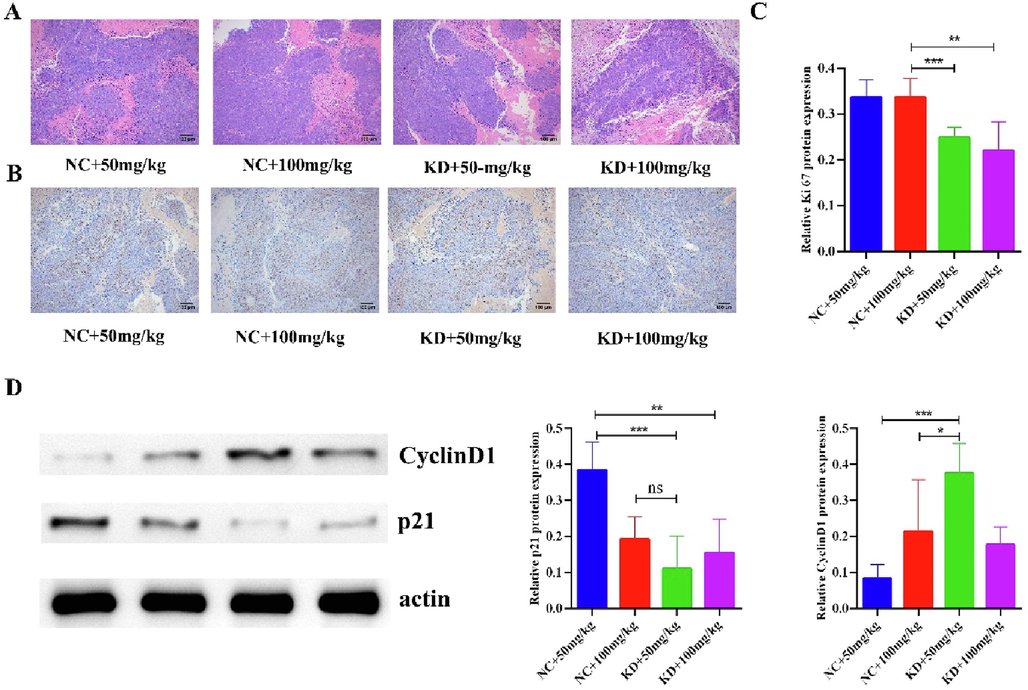
Knockdown of ARRDC4 with baicalin exerts antitumor effect in vivo. (A) Effect of knockdown of ARRDC4 with baicalin on histological changes was visualized using HE staining (×100). (B) Representative images of IHC staining for Ki67 cell in the tumors from the mice in each group (×100). (C) Quantification of immunohistochemistry intensity of Ki67 area in the tumor tissues of mice. The IHC score was calculated and analyzed. (D) Protein expression levels of CyclinD1 and p21 in tumors of nude mice from the NC + baicalin (50 mg/kg), NC + baicalin (100 mg/kg), KD + baicalin (50 mg/kg) and KD + baicalin (100 mg/kg) groups. All data are presented as mean ± SD. *P < 0.05, compared with NC + baicalin (100 mg/kg) group; **P < 0.01, compared with KD + baicalin (100 mg/kg) group; ***P < 0.01, compared with KD + baicalin (50 mg/kg) group.
3.6 Baicalin inhibited cell proliferation by regulating LCN2, Claudin-2 and GSTA4
To explore the target of ARRDC4, we performed RNA sequencing with shARRDC4-HCT-116 and HCT-116 cells, and we screened 25 genes with consistent trends with the previous sequencing results (Fig. 8). Knocking down ARRDC4 significantly increased LCN2 levels and reduced Claudin-2 and GSTA4 levels (Fig. 9). These findings suggest that ARRDC4 mediates CRC cell proliferation through LCN2, Claudin-2, and GSTA4.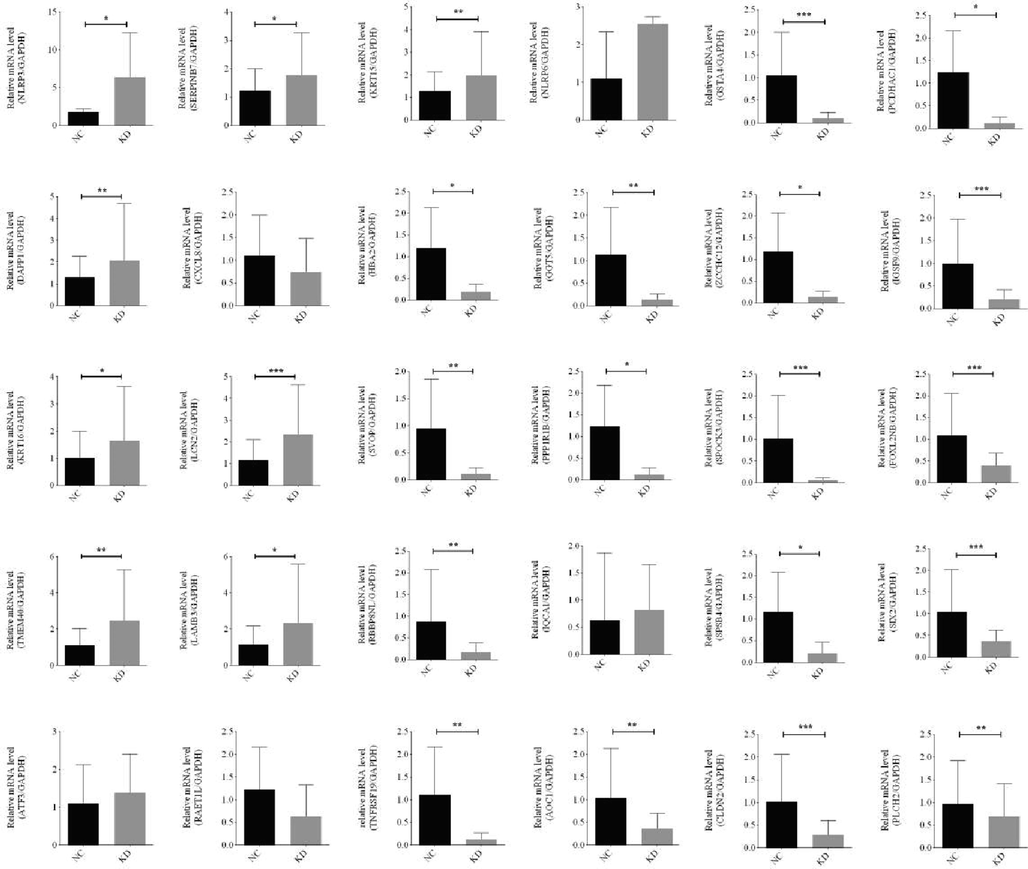
RNA-seq sequencing was performed with sh ARRDC4-HCT-116 or HCT-116 cells, and we screened 30 genes with consistent trends with the previous sequencing results.
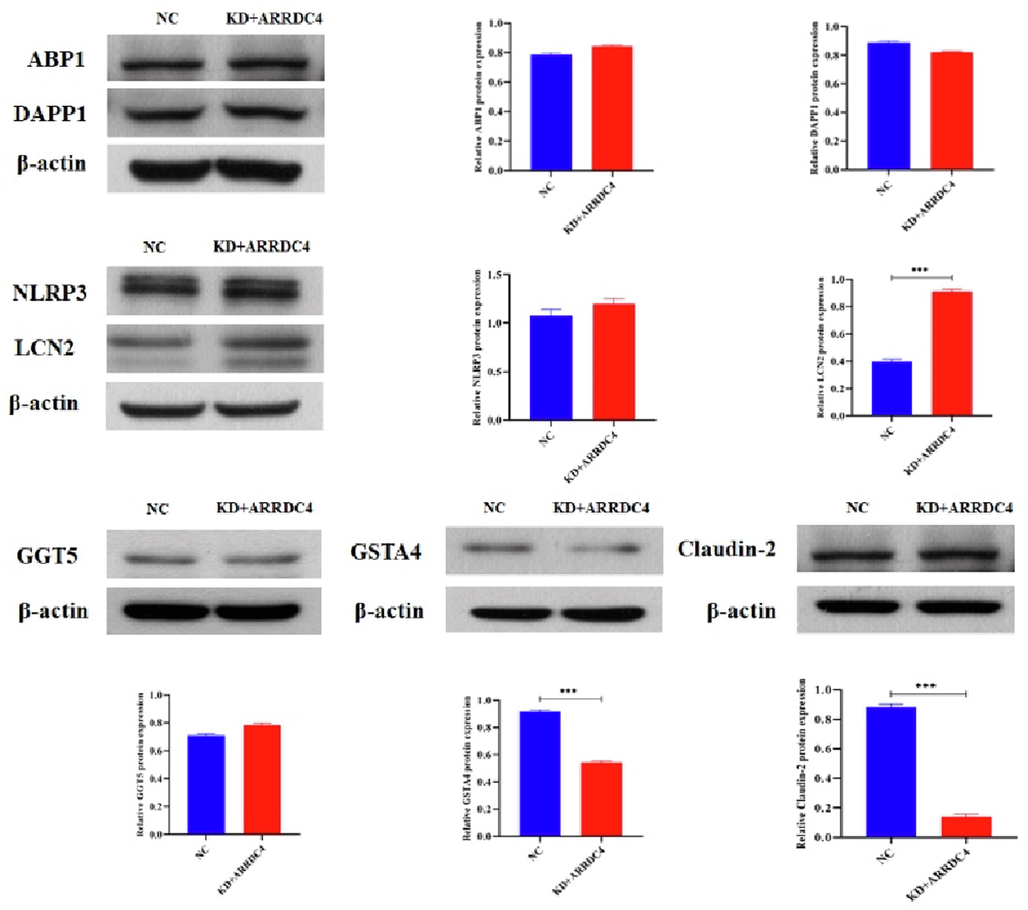
Western blot assay was performed to test ABP1, DAPP1, NLRP3, LCN2, GGT5, GSTA4 and Claudin-2 expression in the HCT-116 cells with or without ARRDC4. β-actin was performed as an internal control. Data are presented as the mean ± SD and are analyzed by one-way ANOVA. ***P < 0.001.
4 Discussion
Traditional Chinese medicine (TCM) has long been an important part of cancer treatment in China, which exerts great effects in increasing efficiency and reducing toxicity, improving life quality and prolonging survival time (Xiang et al., 2019). The majority of researchers favor Traditional Chinese medicine remedies with a single active ingredient, as their efficacy is clearer and they are usually less toxic and have fewer side effects on normal cells (Zhu et al., 2021; Li et al., 2021a, 2021b, 2021c). For example, berberine reduces the risk of developing colon cancer in clinical practice (Chen et al., 2020) and ApcMin/+ mice treated it exhibit fewer and smaller polyps in the intestine, along with lower cyclin D1 and c-Myc expression levels (Zhang et al., 2013). Studies have shown that flavonoids in Chinese herbal extracts have anti-tumor, anti-bacterial, immunosuppressive and anti-inflammatory pharmacological activities (Jucá et al., 2020). Nowadays, targeted therapy has become an important approach for CRC with good curative effect (Zhao et al., 2022). For instance, the molecular targeted drug panitumumab has been applied in clinical practice and demonstrated a strong therapeutic effect (Peeters et al., 2018). Nevertheless, finding new therapeutic targets remains an urgent task. Baicalin inhibits the development of CRC, but its mechanisms need to be explored. In our study, we used microarray technology to screen the DEGs between the control HCT-116 cells and the baicalin-treated HCT-116 cells; we thus identified potential targets of baicalin associated with the inhibition of CRC cell proliferation. We found 19 oncogenes and, based on the HCS outcomes, we chose to confirm the biological role of ARRDC4 in HCT-116 cells. We confirmed that transfection with the lentiviral shRNA targeting ARRDC4 decreased the mRNA and protein expression of ARRDC4 in the CRC cell line HCT-116. The MTT assay and Celigo analysis results indicated that knocking down ARRDC4 notably inhibited the proliferation of CRC cells, in accordance with reports on other tumors. Besides, ARRDC4 plays a pivotal role for baicalin to exert its activity on CRC cells. We also confirmed that baicalin inhibited mRNA and protein expressions of ARRDC4, in line with the RNA sequencing results. ARRDC4 is closely homologous to TXNIP (thioredoxin interacting protein), another alpha-arrestin, and plays a vital role in glucose metabolism and G-protein-coupled receptor-related physiological and pathological processes (Patwari and Lee, 2012; Patwair et al., 2009). Note-worthily, ARRDC4 expression is closely correlated with predicted pathway activities of lactic acidosis and is correlated with favorable clinical outcomes in human cancers (Patwari and Lee, 2012; Soung et al., 2014). ARRDC4 was identified as a downregulated gene by interrogating the Gene Expression Omnibus (GEO) for microarray expression data in metformin-treated and metformin-untreated groups (Soung et al., 2014). The expression of ARRDC4 is much higher in normal tissues than in tumor tissues of patients with CRC, according to the GEPIA data platform (Huang, et al., 2023). Additionally, high ARRDC4 expression is positively correlated with favorable survival outcomes in cancer patients (Chen et al., 2010). However, very little research has addressed the regulatory ability of ARRDC4 on CRC. Our results showed that ARRDC4 knockdown significantly suppressed CRC proliferation, an effect similar to baicalin intervention. Moreover, the combination of ARRDC4 knockdown and baicalin treatment enhanced the baicalin-induced inhibition of CRC progression.
Knocking down ARRDC4 significantly increased LCN2 levels and decreased GSTA4 and Claudin-2 levels (Fig. 9). LCN2, a member of the secretory lipoprotein family, is involved in innate immunity and apoptosis (Wu et al., 2021). What’s interesting, some researches reported a reciprocal modulation as TME regulated LCN2, SLC22A17, and MMP9 expressions which in turn modified TME dynamism (Candido et al., 2022). According to the present study, GSTA4 is a detoxifying enzyme for 4-HNE, 13, which is greatly expressed in macrophages and epithelial cells of microbiome-driven murine colitis and biopsies of human colon adenomas and carcinomas (Yang et al., 2016). GSTA4 is activated at the initial stage of colorectal carcinogenesis, making it a potential CRC biomarker (Zhang et al., 2022).GSTA4 overexpression promotes cell proliferation, tumorigenesis, and chemoresistance in CRC; thus, inhibiting GSTA4 is a potential CRC therapy approach. As a component of cellular tight junctions, claudin-2 is involved in the progression of various cancers where its expression is elevated (Wei et al., 2021). In addition, claudin-2 expression is upregulated in CRC samples and is correlated with poor survival chances.
There are several limitations in the present study. We did not perform IHC staining of selected markers. In addition, if baicalin inhibits CRC proliferation by down-regulating ARRDC4, what happens when only the gene is downregulated in vivo? If ADRRC4 inhibits cell proliferation by regulating the expression of LCN2, Claudin-2, and GSTA4, functional recovery verification is possible. Since LCN2 is involved in innate immunity, we will test the possibility of baicalin being an active cytotoxic compound activating the immune system, for example by doing co-culture assays of HCT-116 cell lines and macrophages. After co-culture, the activation of macrophages can be monitored by detecting inflammatory factors and determining the flow cytometry index of macrophage differentiation. To overcome these limitations, we plan to study the ubiquitination or protein interactions from the perspective of protein stability using co-immunoprecipitation.
5 Conclusion
We found that baicalin is demonstrated to have profound anti-tumor activity against CRC as inhibiting tumor cell proliferation. To sum up, ARDDC4 may be a critical gene that regulates CRC cell proliferation, which may serve as a potential target of baicalin in treating patients with CRC. Further verification of these findings is needed.
Funding
This work was funded by Macau Young Scholars Program (Project code: AM2020020), the fifth batch of Gusu health personnel training project in Suzhou (Project code: GSWS2020085), Natural Science Foundation of Nanjing University of Chinese Medicine (Project code: XZR2020038), Science and Technology Innovation Project of Suzhou Medical and Health Care (Project code: SKJY2021136).
CRediT authorship contribution statement
Shuai Yan: Investigation, Methodology, Project administration, Writing – review & editing. Yahui Wang: Data curation, Visualization. Yunhui Gu: Data curation, Visualization. Mingyue Zhou: . Lianlin Su: Data curation, Visualization. Tianpeng Yin: . Wei Zhang: Conceptualization, Funding acquisition. Yinzi Yue: Data curation, Visualization.
Acknowledgments
We would like to thank all the people who participated in this study.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Diagnosis and treatment of metastatic colorectal cancer: a review. J. Am. Med. Assoc.. 2021;325:669-685.
- [Google Scholar]
- Bioinformatic analysis of the LCN2-SLC22A17-MMP9 network in cancer: the role of DNA methylation in the modulation of tumor microenvironment. Front. Cell Dev. Biol.. 2022;10:945586
- [Google Scholar]
- Changing profiles of cancer burden worldwide and in China: a secondary analysis of the global cancer statistics 2020. Chin Med. J. (Engl).. 2021;134:783-791.
- [Google Scholar]
- Berberine versus placebo for the prevention of recurrence of colorectal adenoma: a multicentre, double-blinded, randomised controlled study. Lancet Gastroenterol. Hepatol.. 2020;5:267-275.
- [Google Scholar]
- Baicalin induces apoptosis in SW620 human colorectal carcinoma cells in vitro and suppresses tumor growth in vivo. Molecules. 2012;17:3844-3857.
- [Google Scholar]
- Lactic acidosis triggers starvation response with paradoxical induction of TXNIP through MondoA. PLoS Genet.. 2010;6:e1001093.
- [Google Scholar]
- Cabozantinib versus Everolimus in advanced renal-cell carcinoma. N. Engl. J. Med.. 2015;373:1814-1823.
- [Google Scholar]
- Targeting RPL39 and MLF2 reduces tumor initiation and metastasis in breast cancer by inhibiting nitric oxide synthase signaling. PNAS.. 2014;111:8838-8843.
- [Google Scholar]
- Baicalein and baicalin inhibit colon cancer using two distinct fashions of apoptosis and senescence. Oncotarget.. 2018;9:20089-20102.
- [Google Scholar]
- Current cancer situation in China: good or bad news from the 2018 Global Cancer Statistics? Cancer Commun. (Lond).. 2019;39:22.
- [Google Scholar]
- The development of high-content screening (HCS) technology and its importance to drug discovery. Expert Opin. Drug Discov.. 2016;11:501-514.
- [Google Scholar]
- Brusatol suppresses the tumor growth and metastasis of colorectal cancer via upregulating ARRDC4 expression through modulating PI3K/YAP1/TAZ Pathway. Phytomedicine. 2023;109:154567
- [Google Scholar]
- Baicalin induced colon cancer cells apoptosis through miR-217/DKK1-mediated inhibition of Wnt signaling pathway. Mol. Biol. Rep.. 2019;46:1693-1700.
- [Google Scholar]
- Flavonoids: biological activities and therapeutic potential. Nat. Prod. Res... 2020;34:692-705.
- [Google Scholar]
- Genetic and biological hallmarks of colorectal cancer. Genes Dev.. 2021;35:787-820.
- [Google Scholar]
- Discovery of MTR-106 as a highly potent G-quadruplex stabilizer for treating BRCA-deficient cancers. Invest. New Drugs. 2021;39:1213-1221.
- [Google Scholar]
- SANT, a novel Chinese herbal monomer combination, decreasing tumor growth and angiogenesis via modulating autophagy in heparanase overexpressed triple-negative breast cancer. J. Ethnopharmacol.. 2021;266:113430
- [Google Scholar]
- Baicalin induces apoptosis in SW480 cells through downregulation of the SP1 transcription factor. Anticancer Drugs.. 2019;30:153-158.
- [Google Scholar]
- Cell-Based High Content Analysis of Cell Proliferation and Apoptosis. Methods Mol. Biol.. 2018;1683:47-57.
- [Google Scholar]
- High-content screening: imaging, analysis, and applications. J. Biomol. Screen.. 2013;18:845-847.
- [Google Scholar]
- High-Content screening for quantitative cell biology. Trends Cell Biol.. 2016;26:598-611.
- [Google Scholar]
- NCCN guidelines updates: management of metastatic colorectal cancer. J. Natl. Compr. Canc. Netw... 2019;17:599-601.
- [Google Scholar]
- Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods. 2008;5:621-628.
- [Google Scholar]
- A metabolomic analysis of medicinal diversity in Huang-qin (Scutellaria baicalensis Georgi) genotypes: discovery of novel compounds. Plant Cell Rep.. 2004;23(6):419-425.
- [Google Scholar]
- Postoperative chemotherapy is associated with prognosis of stage IV colorectal cancer treated with preoperative chemotherapy/chemoradiotherapy and curative resection. Int. J. Colorectal Dis.. 2020;35:177-180.
- [Google Scholar]
- Thioredoxin-independent regulation of metabolism by the alpha-arrestin proteins. J. Biol. Chem.. 2009;284:24996-25003.
- [Google Scholar]
- An expanded family of arrestins regulate metabolism. Trends Endocrinol. Metab.. 2012;23:216-222.
- [Google Scholar]
- Relationships between tumour response and primary tumour location, and predictors of long-term survival, in patients with RAS wild-type metastatic colorectal cancer receiving first-line panitumumab therapy: retrospective analyses of the PRIME and PEAK clinical trials. Br. J. Cancer. 2018;119:303-312.
- [Google Scholar]
- Inhibition of autophagy amplifies baicalein-induced apoptosis in human colorectal cancer. Mol. Ther. Oncolytics.. 2020;19:1-7.
- [Google Scholar]
- Targeted therapy for colorectal cancer metastases: a review of current methods of molecularly targeted therapy and the use of tumor biomarkers in the treatment of metastatic colorectal cancer. Cancer. 2019;125:4139-4147.
- [Google Scholar]
- Colorectal cancer surgery in selected nonagenarians is relatively safe and it is associated with a good long-term survival: an observational study. World J. Surg. Oncol... 2020;18:120.
- [Google Scholar]
- Baicalin triggers apoptosis, inhibits migration, and enhances anti-tumor immunity in colorectal cancer via TLR4/NF-κB signaling pathway. J. Food Biochem... 2022;46:e13703.
- [Google Scholar]
- Epigenetic silencing of ARRDC3 expression in basal-like breast cancer cells. Sci. Rep.. 2014;4:3846.
- [Google Scholar]
- Baicalin, an emerging multi-therapeutic agent: pharmacodynamics, pharmacokinetics, and considerations from drug development perspectives. Xenobiotica. 2010;40:357-367.
- [Google Scholar]
- Cancer burden and trends in China: A review and comparison with Japan and South Korea. Chin. J. Cancer Res.. 2020;32:129-139.
- [Google Scholar]
- Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin.. 2021;71:209-249.
- [Google Scholar]
- Baicalin, the major component of traditional Chinese medicine Scutellaria baicalensis induces colon cancer cell apoptosis through inhibition of oncomiRNAs. Sci. Rep... 2018;8:14477.
- [Google Scholar]
- 5-fluorouracil and other fluoropyrimidines in colorectal cancer: Past, present and future. Pharmacol. Ther.. 2020;206:107447
- [Google Scholar]
- Baicalin induces cellular senescence in human colon cancer cells via upregulation of DEPP and the activation of Ras/Raf/MEK/ERK signaling. Cell Death Dis... 2018;9:217.
- [Google Scholar]
- A comprehensive review on phytochemistry, pharmacology, and flavonoid biosynthesis of Scutellaria baicalensis. Pharm. Biol.. 2018;56:465-484.
- [Google Scholar]
- Claudin-2 promotes colorectal cancer growth and metastasis by suppressing NDRG1 transcription. Clin. Transl. Med.. 2021;11:e667.
- [Google Scholar]
- The effect of lipocalin-2 (LCN2) on apoptosis: a proteomics analysis study in an LCN2 deficient mouse model. BMC Genomics.. 2021;22:892.
- [Google Scholar]
- Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin Med J (Engl).. 2022;135:584-590.
- [Google Scholar]
- Traditional Chinese medicine as a cancer treatment: modern perspectives of ancient but advanced science. Cancer Med.. 2019;8:1958-1975.
- [Google Scholar]
- Baicalin promotes cell cycle G2 /M arrest to inhibit growth of orthotopic xenografts with colorectal cancer cells that are deficient in a mismatch repair gene in nude mice. Pharmocol Clin. Chin Mater. Med.. 2017;33:24-28.
- [Google Scholar]
- Inhibiting EMT, stemness and cell cycle involved in baicalin-induced growth inhibition and apoptosis in colorectal cancer cells. J. Cancer. 2020;11:2303-2317.
- [Google Scholar]
- Inhibitory effects of baicalin on orthotopic xenografts of colorectal cancer cells that are deficient in a mismatch repair gene in nude mice. Int. J. Colorectal Dis.. 2013;28:547-553.
- [Google Scholar]
- Glutathione S-transferase alpha 4 induction by activator protein 1 in colorectal cancer. Oncogene.. 2016;35:5795-5806.
- [Google Scholar]
- Berberine potently attenuates intestinal polyps growth in ApcMin mice and familial adenomatous polyposis patients through inhibition of Wnt signalling. J. Cell Mol. Med.. 2013;17(11):1484-1493.
- [Google Scholar]
- Use Chou's 5-steps rule to study how Baicalin suppresses the malignant phenotypes and induces the apoptosis of colorectal cancer cells. Arch. Biochem. Biophys.. 2021;705:108919
- [Google Scholar]
- Glutathione S-Transferase Alpha 4 promotes proliferation and chemoresistance in colorectal cancer cells. Front. Oncol.. 2022;12:887127
- [Google Scholar]
- Scutellaria baicalensis, the golden herb from the garden of Chinese medicinal plants. Sci. Bull. (Beijing). 2016;61(18):1391-1398.
- [Google Scholar]
- Colorectal cancer immunotherapy-Recent progress and future directions. Cancer Lett.. 2022;545:215816
- [Google Scholar]
- The reference genome sequence of scutellaria baicalensis provides insights into the evolution of wogonin biosynthesis. Mol. Plant. 2019;12:935-950.
- [Google Scholar]
- Drug-herb interactions between Scutellaria baicalensis and pharmaceutical drugs: Insights from experimental studies, mechanistic actions to clinical applications. Biomed. Pharmacother.. 2021;138:111445
- [Google Scholar]
- Tripterine: a potential anti-allergic compound. Curr. Pharm. Biotechnol.. 2021;22:159-167.
- [Google Scholar]
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105141.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1
Supplementary data 2
Supplementary data 2







