Translate this page into:
Computational insights into the antioxidant and antidiabetic mechanisms of cannabidiol: An in vitro and in silico study
⁎Corresponding author. matsabisamg@ufs.ac.za (Motlalepula G. Matsabisa)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
The present study was aimed at investigating the molecular mechanism involved the antioxidant and antidiabetic activities of cannabidiol using in vitro and in silico models. The antioxidant activities of cannabidiol were determined by assaying its ability to scavenge 2,2-diphenyl-1-picryl-hydrazyl-hydrate (DPPH) radical and reduce ferric ion (Fe3+). The antidiabetic activity of cannabidiol was determined by its ability to inhibit α-glucosidase activity. The DPPH scavenging activity of cannabidiol was further investigated using the Density Functional Theory (DFT) analysis. The molecular interactions of cannabidiol and α-glucosidase activity was determined via molecular docking and molecular dynamics simulation. Cannabidiol significantly scavenged DPPH radical and reduced Fe3+, with IC50 values of 211.93 and 101.72 μg/mL, respectively. It also inhibited the activity of α-glucosidase, with an IC50 value of 311.26 μg/mL. DFT analysis revealed sequential proton loss followed by electron transfer (SPLET) as the preferred mechanism by which cannabidiol scavenge DPPH, followed by hydrogen atom transfer (HAT) and single electron transfer followed by proton transfer (SET-PT). The highest occupied molecular orbital (HOMO) orbitals were delocalised on the benzene ring and O—H groups, while the lowest energy orbital (LUMO) orbitals were distributed on both the parent and substituent molecules. Cannabidiol bounded favourably to α-glucosidase enzyme, with G, root mean square deviation (RMSD), root mean square fluctuation (RMSF) values of −28.49 kcal/mol, 1.36 Å and 0.835 Å as well as higher correlated motions. Therefore, the antioxidant activity of cannabidiol involves the abstraction of its H1 by the radical and delocalization of the radical’s unpaired electron. Its antidiabetic activity can be attributed to the potent inhibitory effect on α-glucosidase, as well as its high affinity with the enzyme.
Keywords
Antioxidant
Cannabis sativa
Cannabidiol
Computations
Density functional theory
Diabetes
1 Introduction
There has been increasing search for drugs from natural products to treat type 2 diabetes (T2D) and its complications. The search can be attributed to the high cost associated with the treatment of T2D (Kähm et al., 2018; Kang et al., 2018), as well as the side effects associated with commercial antidiabetic drugs (Palanisamy et al., 2018). Type 2 diabetes is the highest contributor to mortality morbidity associated with diabetes mellitus (DM) as it accounts for over 90% of all diabetes types (Erukainure et al., 2017; Forouhi and Wareham, 2019). The therapeutic aim of antidiabetic drugs is to suppress hyperglycemia and improve glucose homeostasis. Hyperglycemia arising from insulin resistance and pancreatic β-cell dysfunction is the hallmark of T2D. Hyperglycemia exacerbates the production of free radicals, with concomitant suppression of the body’s endogenous antioxidant system and thereby inducing oxidative stress (Yaribeygi et al., 2019). Oxidative stress has been implicated in the pathophysiology of diabetic complications such as neuropathology, retinopathy and nephropathy (Bhatti et al., 2022).
Inhibition of glucose metabolizing enzymes especially α-glucosidase, is a major antihyperglycemic mechanism of some antidiabetic drugs termed α-glucosidase inhibitors. Common α-glucosidase inhibitors are acarbose, miglitol, and voglibose, with acarbose being the most common (Joshi et al., 2015). This mechanism has also been reported for most antidiabetic medicinal plants and their isolated compounds (Dirir et al., 2021; Papoutsis et al., 2021). Amongst these compounds is cannabidiol (Ghasemi-Gojani et al., 2022) (Fig. 1).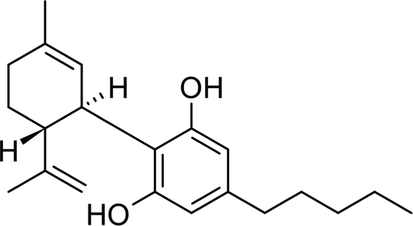
Chemical structure of cannabidiol.
Cannabidiol is among the non-psychoactive phytocannabinoids found in Cannabis sativa, with a chemical formula of C21H30O2. Its chemical structure consists of a phenolic ring, cyclohexene ring, pentyl side chain, 21 carbon atoms, hydroxyl groups at the C-10 and C-50 positions, methyl group at the C-1 position of the cyclohexene ring, and a pentyl chain at the C-30 of the phenolic ring (Atalay et al., 2020b). Cannabidiol has been reported for its antioxidant and antidiabetic properties (Atalay et al., 2020b; Ghasemi-Gojani et al., 2022; Zorzenon et al., 2019). Its antioxidant properties have been attributed to its ability to scavenge free radicals, chelate transition metals, reduce ferric ion to ferrous ion, activate antioxidant enzymes, and reduce malondialdehyde (MDA) level (Atalay et al., 2020a; Atalay et al., 2020b; Khaksar et al., 2022). The antidiabetic properties of cannabidiol have been attributed to its ability to modulate glucose homeostasis, improve insulin secretion, decrease AGEs levels, and inhibit α-glucosidase activity (Suttithumsatid et al., 2022; Zorzenon et al., 2019). Using bio-computational tools to support the experimental evidence reported earlier, this study seeks to unravel the probable mechanism of simultaneous inhibition of α-glucosidase by Cannabidiol during 100 ns molecular dynamic simulation (Bian et al., 2019). Computational tools including molecular dynamic simulations, Docking and post-analyses profiling predictions were utilized to gain more knowledge on the structural and molecular mechanism of Cannabidiol in the inhibition of α-glucosidase (Aljoundi et al., 2019, 2020; Mohamed et al., 2022).
Despite the reported antioxidant and antidiabetic properties of cannabidiol, there is still a dearth on the molecular mechanism involved in the activation of these activities by the compound. The present study is aimed at demystifying the molecular mechanism by which cannabidiol activates its antioxidant and antidiabetic activities using in vitro and in silico models.
2 Materials and methods
2.1 Chemical and reagents
2,2-diphenyl-1-picryl-hydrazyl-hydrate, sodium phosphate buffer, potassium ferricyanide, trichloroacetic acid, ferric chloride, ascorbic acid, α-glucosidase enzyme, p-nitrophenyl-α-D-glucopyranoside and acarbose were obtained from Sigma-Aldrich, Johannesburg, South Africa.
2.2 Cannabidiol
Cannabidiol was obtained from Sigma-Aldrich, Johannesburg, South Africa.
2.3 Permit approval
The present study was undertaken under the permit approval (Permit No. POS 014/2021/2022) from the South African Health Products Regulatory Authority to import, conduct, collect, possess, transport and store cannabis plant, plant parts and products for research purposes.
2.4 Biological activities
2.4.1 Antioxidant activities
2.4.1.1 2,2-diphenyl-1-picryl-hydrazyl-hydrate (DPPH) radical scavenging activity
The free radical scavenging activity of cannabidiol was determined using previously published method with slight modifications (Oyebode et al., 2019). Briefly, 200 μL of cannabidiol at various concentrations (30–250 µg/mL) was incubated with 100 µL of 0.3 mM DPPH solution at room temperature for 30 min in the dark. Absorbance was read at 517 nm against blank without sample. The scavenging activity was calculated using the formula:
Ascorbic acid was used as the standard antioxidant drug (positive control).
2.4.1.2 Ferric (Fe3+) reducing antioxidant power (FRAP) activity
The reducing ability of cannabinoid was determined using a previously published method with slight modifications (Oyebode et al., 2019). Briefly, 100 μL of cannabinoid at different concentration (30–250 µg/mL) was incubated with 100 μL sodium phosphate buffer (0.2 M, pH 6.6) and 100 mL of 1% potassium ferricyanide at 50 °C for 30 min. 100 μL of 10% trichloroacetic acid was used to acidify the reaction mixture. The reaction mixture was further mixed with 100 μL of distilled water and 200 µL of 0.1% ferric chloride (FeCl3). Absorbance was read at 700 nm using a microplate reader. The results were calculated using the formula below and expressed as a percentage of the absorbance of the samples to the absorbance of gallic acid. Ascorbic acid was used as the standard antioxidant drug (positive control).
2.4.2 α-Glucosidase inhibitory activity
The α-glucosidase inhibitory activity of cannabinoid was carried out according to a previously published method (Oboh et al., 2017) with slight modifications. Briefly, 100 µL of 1.0 U mL−1 of α-glucosidase enzyme dissolved in phosphate buffer (100 mmol/L, pH 6.8) was incubated with 50 µL of different concentrations of cannabinoid (30–240 μg/mL) for 20 min at 37 °C. Fifty (50) µL of p-nitrophenyl-α-D-glucopyranoside (pNPG) solution (5 mmol/ L; prepared in the same phosphate buffer) was added to the reaction mixture and further incubated for another 30 min at 37 °C. Absorbance was read at 405 nm using a microplate reader. The inhibitory activity was calculated using the formula:
Blank = reaction mixture without sample. Acarbose was used as the standard antidiabetic drug (positive control).
2.5 Computational studies
2.5.1 Density functional theory (DFT) analysis
All quantum chemical calculations were performed in Gaussian 16 program (Frisch et al., 2016). Geometry optimization of the reaction species were performed using the B3LYP/6-311++G(3df,2p) level of theory (Epifanovsky et al., 2021). Frequency calculations were performed at the same level of theory as optimization to confirm if the optimised structures are ground state structures on the potential energy surface. Enthalpy parameters bond dissociation enthalpy (BDE), ionization potential (IP), proton dissociation enthalpy (PDE), proton affinity (PA), and electron transfer enthalpy (ETE) values were calculated using equation explained below:
2.5.1.1 Theoretical background
The most common reaction mechanisms used to study the interactions between antioxidant (RXH) molecule and radical (R•) species include hydrogen atom transfer (HAT), single electron transfer followed by proton transfer (SET-PT), and sequential proton loss followed by electron transfer (SPLET). These reaction mechanisms are utilized to obtain valuable information on the reaction species in order to evaluate the chosen RXH's radical-scavenging abilities (Dehkordi et al., 2022; Zhou et al., 2019). The reaction mechanisms are provided as equations (1), (2), and (3) for the HAT, SET-PT, and SPLET processes, respectively.
The numerical parameters known as the BDE, IP, PDE, PA, and ETE which can be calculated by determining the change in enthalpy of the reaction products and reactants species, are responsible for controlling the mechanisms.
Since this study was performed experimentally in the presence of ethanol solvent, implicit solvation model was used to model the reactions in the presence of the ethanol solvent through solvation model density (Michalík et al., 2019). Gaussview 06 program was used to draw and visualise the reaction species (Dennington et al., 2016).
2.5.2 Molecular docking and molecular dynamics
2.5.2.1 System preparation
X-ray crystal structures of α-glucosidase enzyme (PDB code:3CTT) (Cardona et al., 2009) was attained from RSCB Protein Data Bank (PDB). To prepare these structures for molecular dynamics simulation, Molegro Molecular Viewer (MMV) (Kusumaningrum et al., 2014), and UCSF Chimera software package (Pettersen et al., 2004), were employed. All missing residues from α-glucosidase enzyme were added by a structural refinement tool known as Modeller (Eswar et al., 2007).
2.5.2.2 Molecular docking and molecular dynamics simulation
Preceding docking, UCSF Chimera was used to eliminate all co-factors, non-standard residues such as Cl−, Na+, etc. as well as water molecules surrounding the protein (Pettersen et al., 2004). Afterwards, cannabidiol and acarbose were charged and hydrogenated. AutoDock Tools GUI was used to estimate the ligand docking and adding of a partial Gasteiger charge (Aljoundi et al., 2020). The dimensions and co-ordinates of the grid box was defined as follows, size x = 13.22, y = 20.96, z = 10 with center x = 23.27, y = 9.56, z= −1.24. Overall, system consisting of the α-glucosidase bound to cannabidiol and acarbose underwent a 100 ns MD simulation using the AMBER18 CPU and GPU packages (Case et al., 2022), PMEMD engine. An atomic partial charge was generated for cannabidiol inhibitor by means of the AMBER FF14SB of the ANTECHAMBER program (Wang et al., 2004). The Leap module of Amber 18 enabled the addition of hydrogen atoms, sodium (Na+), or chloride (Cl−) counter ions to the systems for neutralization. The systems were initially minimized for 2500 steps with 500 kcal/mol Å2 restraint potential, and a whole minimization step of 5000 steps was further run without restraint using the conjugate algorithm. Thereafter, the system underwent gradual heating from 0 to 300 K for 50 ps by means of a Langevin thermostat in a canonical assemblage (NVT). Following heating, equilibration was undertaken at a temperature of 300 K, excluding all restraints, and a maintained atmospheric pressure of 1 bar by utilizing the Berendsen barostat until all systems had reached equilibration. The complex then underwent a 100 ns MD simulation with the SHAKE algorithm being employed to constrict all hydrogen atoms. Co-ordinates of the α-glucosidase- cannabidiol and acarbose complexes were saved after the 100 ns production period at every 1 ps intervals. The generated trajectories of each system were analyzed using the PTRAJ and CPPTRAJ modules (Roe and Cheatham III, 2013). The generated data and subsequent complexes were visualized using Microcal Origin analytical software (Seifert, 2014).
2.5.2.3 Post-molecular dynamics simulation analysis
Post analysis for the systems MD simulation included protein, stability (RMSD), flexibility (RMSF). Binding free energy is computed by employing Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) technique.
2.5.2.4 Thermodynamics (free binding energy) computations
To determine an estimate of the binding free energy α-glucosidase bound to cannabidiol and acarbose, a method known as the Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) approach (Homeyer and Gohlke, 2012), has been implemented. Binding free energy represented by ΔGbind is a reliable tool that is extensively used to measure the energies that contribute to the binding of the protein and ligand to form a complex, and is calculated using the following equation:
The ΔGbind is the sum of the gas and solvent energy minus entropy (TS).
The following equation is representative of the solvent energy calculation:
GGB is representative of the polar solvation impact and GSA is representative of the non-polar solvation impact which is calculated using the solvent accessible surface area (SASA). This is attained by using a water probe radius of 1.4 A°. A surface tension constant (c) was represented by a measure of 0.0072 kcal/mol and ‘b’ to 0 kcal/mol (Richard et al., 2016).
2.5.2.5 Dynamic Cross-Correlation (DCCM) analysis
We used dynamic cross-correlation analysis to investigate the fluctuations and movements in the backbone of the α carbon atoms. Cross-correlation components for i and j Cα atoms are shown in Equation (1):
2.5.2.6 Residual overall motions clustering and principal component analysis
Principle component analysis can convert the high-dimensional data of protein dynamics into the low dimensional space to obtain a series of eigenvectors and eigenvalues that replicate overall motions in the protein. The PCA analysis can be applied to any system and permits to study the effect of any varying parameters, by reducing the density of the collective motion, which is associated with the phase space behavior related to protein functions and stability. Consequently, it is often used to distinguish different conformational variances, which are involved in protein open-close mechanism and folding of ion channels, and conformational dynamics.
2.6 Statistical analysis
All biological analyses were carried out in triplicates. Data were analyzed with a statistical software package (SPSS for Windows, version 25, IBM Corporation, USA) using Tukey’s-HSD multiple range post-hoc test. Values were considered significantly different at p < 0.05, reported as mean ± standard deviation (SD).
3 Results and discussion
The role of antioxidant and α-glucosidase inhibitors in the treatment of T2D and its complications have been well documented (Hossain et al., 2020; Unuofin and Lebelo, 2020). Studies have shown phytochemicals to possess potent antioxidant and α-glucosidase inhibitory properties, and have over the years been used in the treatment of T2D (Ademiluyi et al., 2015; Papoutsis et al., 2021). The present study applied in vitro and in silico studies in elucidating the molecular antioxidant and α-glucosidase inhibitory mechanism of cannabidiol.
3.1 Antioxidant properties
The ability to scavenge free radicals is a major antioxidative mechanism for suppressing oxidative stress. As shown in Fig. 2A, cannabidiol displayed a significantly (p < 0.05) scavenging activity against DPPH, with an IC50 value of 211.93. The activity was dose-dependent, with the highest concentration (250 μg/mL) of cannabidiol having the best activity. This activity indicates the potential of cannabidiol to scavenge free radicals and thus, portraying an ability to alleviate oxidative stress. This corroborates previous reports on the free radical scavenging activity of cannabidiol (Russo et al., 2021; Tura et al., 2019), and its ability to mitigate oxidative stress (Atalay et al., 2019; Pereira et al., 2021).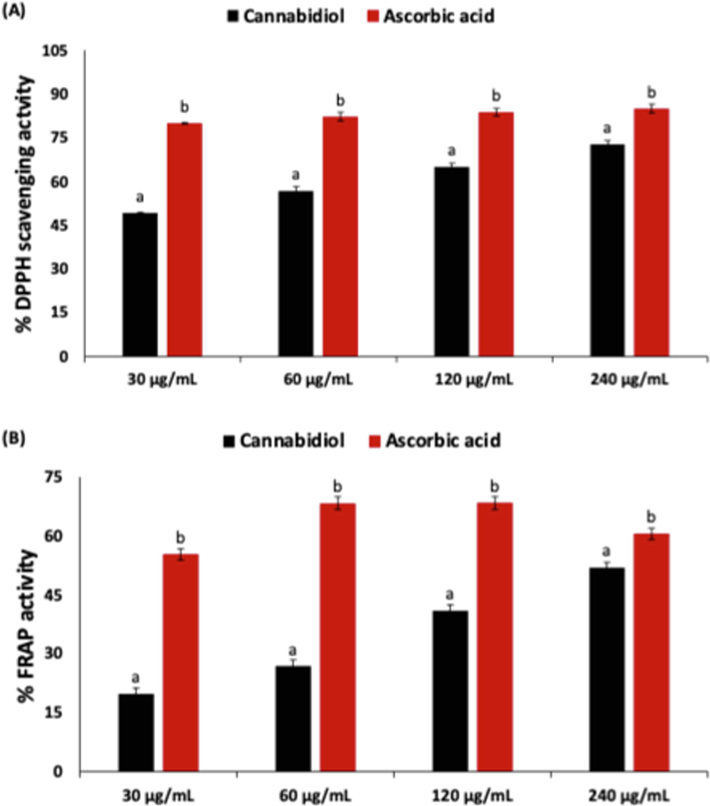
(A) DPPH scavenging and (B) reducing power activities of cannabidiol. Data = mean ± SD; n = 3. a,b Values with different letter above the bars are significantly (p < 0.05) different from each other.
The oxidation of Fe2+ to Fe3+ has been implicated in the generation of hydroxyl radicals (·OH) in the presence of hydrogen peroxide (H2O2) via Fenton’s reaction, thereby triggering oxidative stress (Latunde-Dada, 2017). The reduction of Fe3+ to Fe2+ has been shown to be an antioxidative mechanism as it suppresses the generation of ·OH (Salau et al., 2019). As shown in Fig. 2B, cannabidiol showed significant (p < 0.05) FRAP activity, with an IC50 value of 101.72. This activity was dose dependent, with the highest of concentration of cannabidiol having the best activity. This activity also corroborates previous reports on the ability of cannabidiol to reduce Fe3+ (Dawidowicz et al., 2021). Thus, further demonstrating its antioxidant properties.
3.2 DFT analysis
Understanding the geometrical characteristics of the cannabidiol conformers is crucial to comprehending their antioxidant action. Four structural confirmations of cannabidiol are shown in Fig. 3; the difference between them is the orientation of the O—H groups with regards to the pentyl group. The acronym CBD-1 denotes conformer with both O—H groups oriented towards the pentyl group, CBD-2 represents conformer with both O—H groups oriented away the pentyl group, CBD-3 and CBD-4 conformers are identical in that one OH group is oriented towards and the other OH group is oriented away the pentyl group. The difference between CBD-3 and CBD-4 is that in CBD-3 the OH group oriented towards pentyl group is located away from the alkene group while in CBD-4 it is relatively close to the alkene group. The abbreviations H1 and H2 represent H-atoms of O—H groups oriented towards C⚌C and CH3 groups (first ring), respectively. The reported relative energies in Fig. 3 suggest that the energy difference (relative to the CBD-2 energy) is less than 1 kcal/mol; therefore, all four conformations were further investigated for DPPH radical scavenging activity.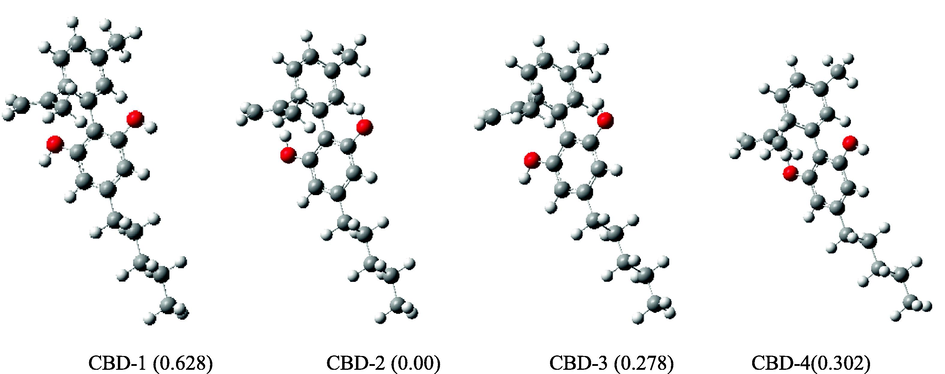
Optimized cannabidiol (CBD) conformers. Relative energies (kcal/mol) in parenthesis were calculated with respect to lowest energy conformation.
In this section, we report the findings and the analysis of the DFT investigation of the DPPH radical scavenging activity of cannabidiol through different mechanistic pathways in ethanol medium. The first part of the discussion is the analysis of the enthalpy values obtained for the HAT mechanism, followed by SET-PT and the SPLET mechanism. The second part of the discussion is the frontier molecular orbitals of the different conformations. After geometry optimisation, the BDE values for the HAT mechanism are shown in Table 1. The BDE values were positive, indicating that the reactions through HAT mechanisms are endothermic. Therefore, an amount of energy is needed to start these reactions. The results also suggests that the BDE value of compound CBD-4 is the highest and lowest compared to the other conformations when the H2 and H1 are abstracted, respectively. This indicates that H1 in CBD-4 can be easily abstracted by the DPPH radical.
Compound
HAT
SET-PT
SPLET
BDE
IP
PDE
PA
ETE
CBD-1-H1
5.164
24.196
–19.032
11.931
–6.767
CBD-1-H2
4.986
24.196
–19.211
11.616
–6.630
CBD-2-H1
4.797
23.234
–18.437
11.616
–6.819
CBD-2-H2
4.946
23.234
–18.288
11.921
–6.975
CBD-3-H1
4.763
23.717
–18.955
11.998
–7.235
CBD-3-H2
4.751
23.717
–18.966
11.723
–6.972
CBD-4-H1
5.490
23.001
–17.511
12.257
–6.767
CBD-4-H2
4.552
23.001
–18.449
11.372
–6.820
The values of IP and PDE for the SET-PT mechanism are also reported in Table 1. The IP values were positive, and the single electron transfer (SET) is likely preferred in CBD-4 followed by CBD-1, CBD-2 and CBD-3. Further analysis of IP values indicates that the lower the IP value of the compound, the stronger the antioxidant activity in comparison to other compounds. In this way, CBD-4 is considered to have the strongest antioxidant potential. The PDE values were negative, which indicate that it is a thermodynamically favoured process. CBD-4 has the lowest PDE value while CBD-1 has the highest PDE value, suggesting that CBD-4 is more likely to transfer a proton than CBD-1. Even though SET is considered to be a rate determining step of the SET-PT mechanism, it is clear that the HAT mechanism is more favoured than the SET-PT mechanism.
The SPL and ET steps are controlled by PA and ETE parameters, respectively and are also shown in Table 1. All PA values were positive, suggesting that the reactions are endothermic. The analysis of the results also suggests that the PA value of compound CBD-4 is the highest and lowest for conformations involving H2 and H1, meaning that it is easier to lose a proton in CBD-4-H2 than CBD-4-H1 active sites. This trend is similar to the one observed in the HAT mechanism analysis. The ETE values are negative, indicating that the ET step is exothermic. Since the ET step is the rate determining step for the SPLET mechanism, we can then conclude that the SPLET mechanism is the preferred mechanism followed by HAT and SET-PT. The analysis of the ETE values also suggest that the CBD-2 and CBD-3 conformers have stronger antioxidant ability than CBD-1 and CBD-4. The most preferred active sites are CBD-3-H1 followed by CBD-2-H2 and CBD-3-H2 while the least active sites are CBD-1-H1, CBD-1-H2 and CBD-4-H2.
According to recent studies, free radical scavenging activity and frontier molecular orbital energy are related (Wang et al., 2015). The electron density distribution of HOMO can be used to anticipate the areas where electron transfer and free radical attack are most likely to occur. The distribution pattern of the HOMO and LUMO molecular orbitals are shown in Fig. 4. It was observed that the HOMO orbitals were delocalised on the benzene ring and O—H groups (i.e., phenolic ring) while the LUMO orbitals were distributed on both the parent and substituent molecules. Therefore, the free radical reaction takes place primarily on the phenolic ring.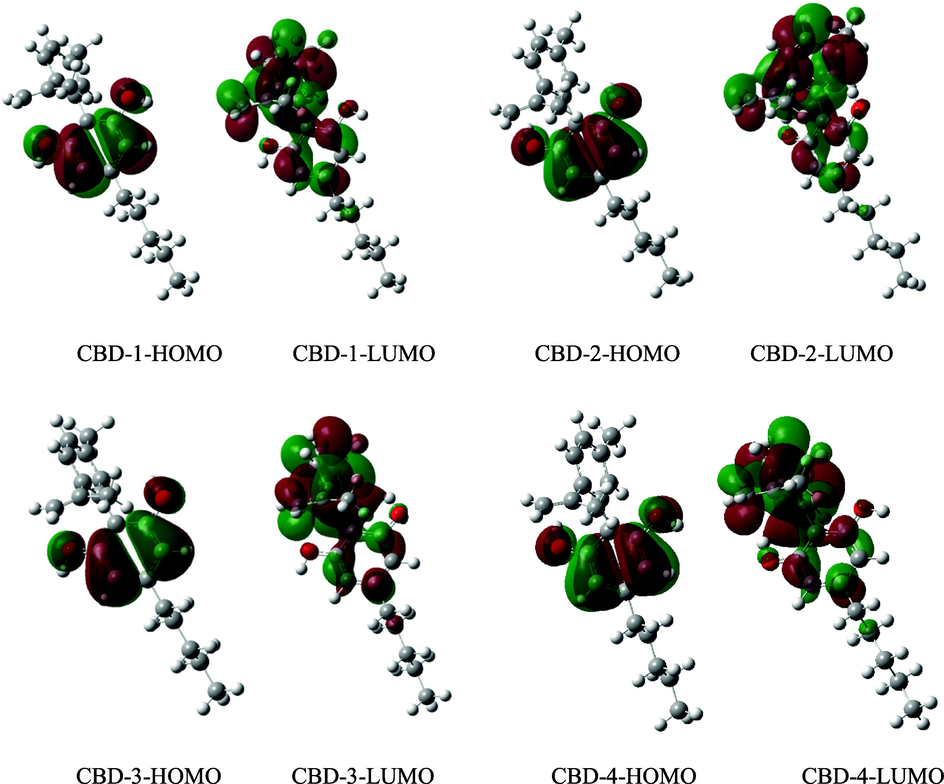
Representative diagram of the distribution pattern of the high occupied molecular orbital and lowest unoccupied molecular orbitals. Positive and negative orbital phases are indicated by the green and red patches, respectively.
The HOMO orbitals energy (EHOMO) and Lumo (ELUMO) and the energy difference (ΔEGap) of the studied compound are shown in Table 2. We can infer information about the chemical activity of the molecule from the energy difference (ΔEGap) between EHOMO and ELUMO. The lower ΔEGap is associated with greater antioxidant activity (Dehkordi et al., 2022; Wang et al., 2015). CBD-3 and CBD-4 are the same while CBD-1 is slightly the lowest; this is explained by the uniform HOMO densities on the compounds which are shown in Fig. 4. The fact that CBD-2 has the highest antioxidant activity whereas CBD-1 has the lowest antioxidant activity is indicated by the ΔEGap value, which is lowest for CBD-2 and highest for CBD-1. From these findings, it may also be inferred that the most stable conformers have higher antioxidant activity whereas the least stable conformers have lower antioxidant activity (Liang et al., 2012; Wang et al., 2015).
Compound
EHOMO
ELUMO
ΔEGap
CBD-1
–0.224
–0.023
0.201
CBD-2
–0.223
–0.031
0.191
CBD-3
–0.223
–0.025
0.197
CBD-4
–0.223
–0.029
0.195
The spin density is a characteristic that well captures the stability and reactivity of radical species. Spin density analysis is a good technique to evaluate the delocalization of an unpaired electron in the radical species. From literature, it has been reported that more delocalized spin density in the radical to simpler radical synthesis, lower BDE, therefore higher radical scavenging activity (Osorio et al., 2013). The spin density distribution on the radical species formed through the HAT mechanism is shown in Fig. 5. The unpaired electron of the radical was delocalised and extended over the benzene ring; this enables the high antioxidant activity.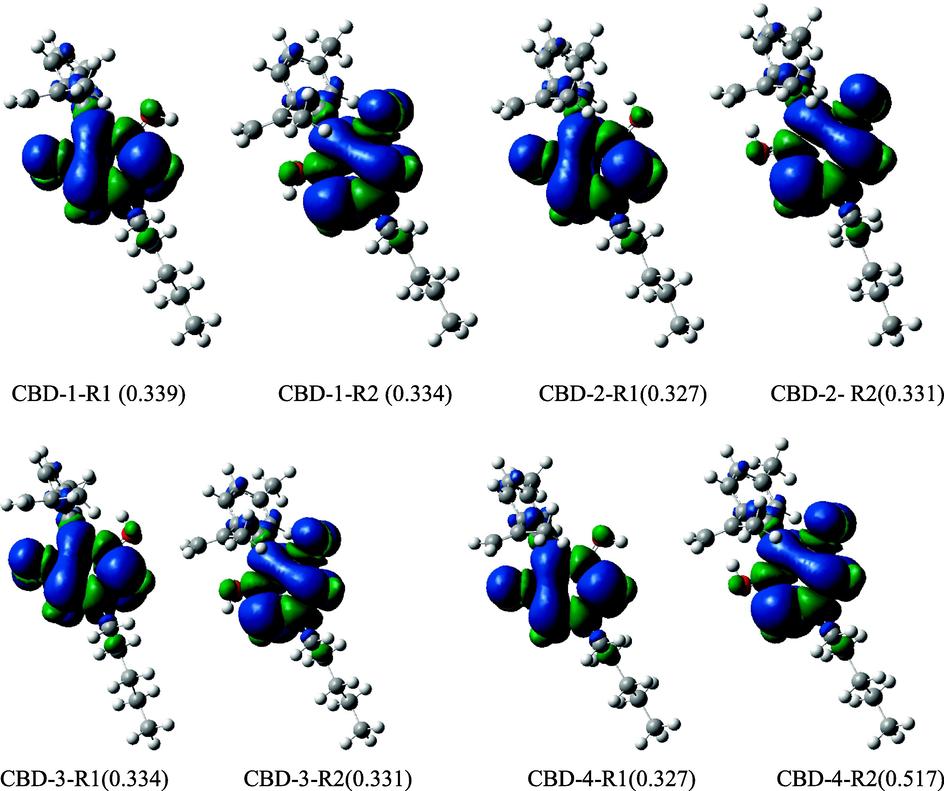
Spin density distribution of the CBD most stable radicals formed through the hydrogen atom transfer mechanism: the blue and green color codes denote the α spin density and the β spin density, respectively. The values in brackets represent the spin density of the oxygen atom from which the hydrogen atom was abstracted. The acronyms R1 and R2 denote the position of which the H-atom was removed.
3.3 α-Glucosidase activity
The inhibition of α-glucosidase activity has been shown to contribute to hypoglycemia as it suppresses the hydrolysis of dietary carbohydrate to glucose by the enzyme (Xiao et al., 2020). Thus, preventing postprandial rise in blood glucose level. As shown in Fig. 6, cannabidiol significantly (p < 0.05) inhibited the activities of α-glucosidase dose-dependently, with an IC50 value of 311.26 μg/mL. This indicates its potential to suppress postprandial rise of blood glucose, thereby contributing to hypoglycemia. This activity corroborates previous studies on the inhibitory effect of cannabidiol on α-glucosidase activity (Suttithumsatid et al., 2022; Zorzenon et al., 2019).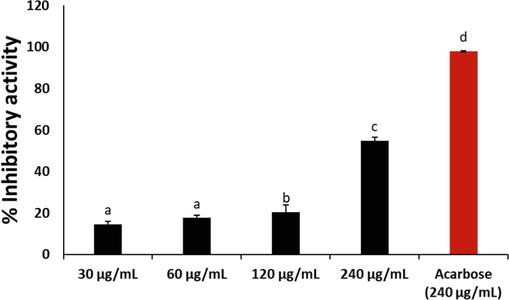
Inhibitory effect of cannabidiol on α-glucosidase activity. Data = mean ± SD; n = 3. a,b,c Values with different letter above the bars are significantly (p < 0.05) different from each other.
3.4 Molecular docking and molecular dynamics simulation
From the results obtained, it was found that cannabidiol and acarbose bounded to α-glucosidase favourably, with
G values measuring −28.49 and-32.14 kcal/mol, respectively (Table 3). Moreover, the promising
G values could indicate analogous modes of binding that possibly underlie the ability of cannabidiol to bind concurrently to the enzyme. All energies are in kcal/mol.
= electrostatic energy;
= van der Waals energy;
= total binding free energy;
= solvation free energy;
= gas phase free energy.
Systems
Energy components(kcal/mol)
Ligand
Cannabidiol
–32.19 ± 0.11
−15.43 ± 0.27
− 47.62 ± 0.27
19.13 ± 0.16
−28.49 ± 0.16
Acarbose
−42.35 ± 0.12
−12.27 ± 0.06
− 54.63 ± 0.13
22.48 ± 0.06
–32.14 ± 0.14
3.5 Exploration of stability and flexibility of the Inter-atomic motion and dynamics of cannabidiol and acarbose bound to α-glucosidase
The therapeutic impact of a small molecule depends on its stability in a target enzyme binding region. The type of interactions between the small molecule and the site residues is the primary factor that determines its stability. The root mean square deviation (RMSD) measures the difference between a protein’s backbones from its initial structural conformation to its final position. However, the residual conformational analysis is a measure of the nature of fluctuation exhibited by individual residue corresponding to the effect of ligand induction on the protein cumulatively yielding its therapeutic efficacy. The average RMSD value of the systems was estimated, the plot revealed the cannabidiol and acarbose had a mean RMSD value of 1.36 Å and 2.29 Å, respectively (Fig. 7A). While the average of the root mean square fluctuation (RMSF) values were 0.835 Å and 0.883 Å, respectively (Fig. 7B). An elevated RMSF value is associated with high fluctuations in the backbone structure, resulting in a more flexible structure (Kumar et al., 2014). Thus, the mean RMSD value indicates that the binding of cannabidiol to α-glucosidase induced stability in the backbone atoms of the enzyme. This suggests that acarbose is less stable in comparison.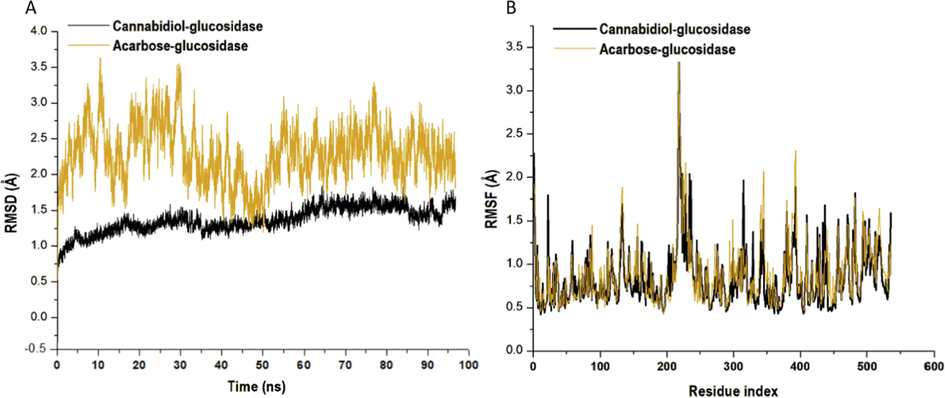
Structural representation of alterations on the binding of cannabidiol (black) and acarbose (orange) bound to α-glucosidase. (A) The conformational stability, C-α atoms RMSD for α-glucosidase enzyme to acarbose bound. (B) The time evolution RMSF of each residue of the enzyme C-α atom over 100 ns for cannabidiol (Black) and acarbose (orange) bound to α-glucosidase.
3.6 Domain cross-correlation matrix (DCCM)
Different enzyme dynamics between the three conditions were examined and plotted using a dynamic cross-correlation matrix for observing the correlated movements of all residues. DCCM plots are represented by different colors, highly positive regions range from green to red (strongly correlated), and highly negative range from light blue to black (strong anticorrelated) movement for specific residues, respectively. As evident in Fig. 8, Cannabidiol-α-glucosidase complex displayed higher correlated motions compared to the Acarbose-α-glucosidase complex. These results correlate with that of RMSF and RMSD which justified that the region of the binding site exhibits a highly correlated motion during the simulation per time. Taken together, these results suggest that the conformation flexibility and stability of the entire enzyme was improved by the binding of cannabidiol.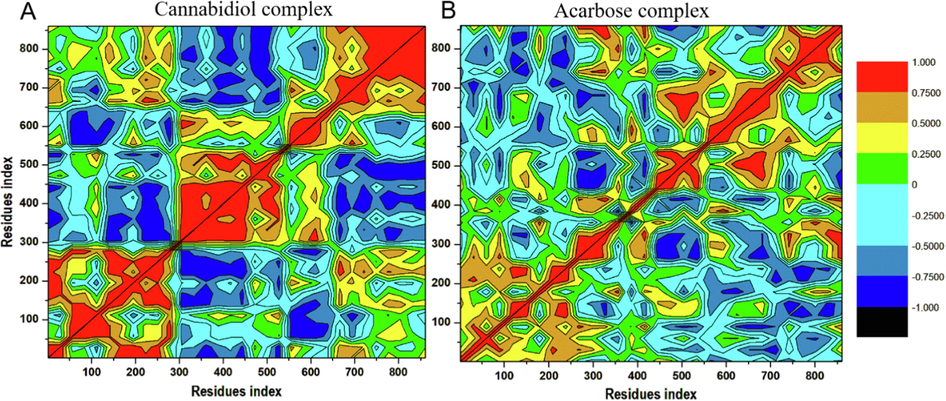
Dynamic cross-correlation matrix analyses for (A) Cannabidiol-α-glucosidase and (B) Acarbose-α-glucosidase. Numbers closer to 1 indicate high correlation, while those closer to −1 indicate anticorrelation between pairs of residues.
3.7 Residual overall motions clustering and principal component analysis
Principle component analysis and clustering have been carried out for cannabidiol and acarbose complexes to realize the overall concerted motion of α-glucosidase. The structural distribution of conformational changes and the proportion of variance of the captured eigenvectors are shown in (Fig. 9A and B). The first three principal components accounted for 51.22% and 30.77% of the total variance observed in the MD trajectories for cannabidiol and acarbose complexes. The clustering analysis also shows conformational distribution variance along with the first, second, and third principal components with each dot representing a single complex conformation.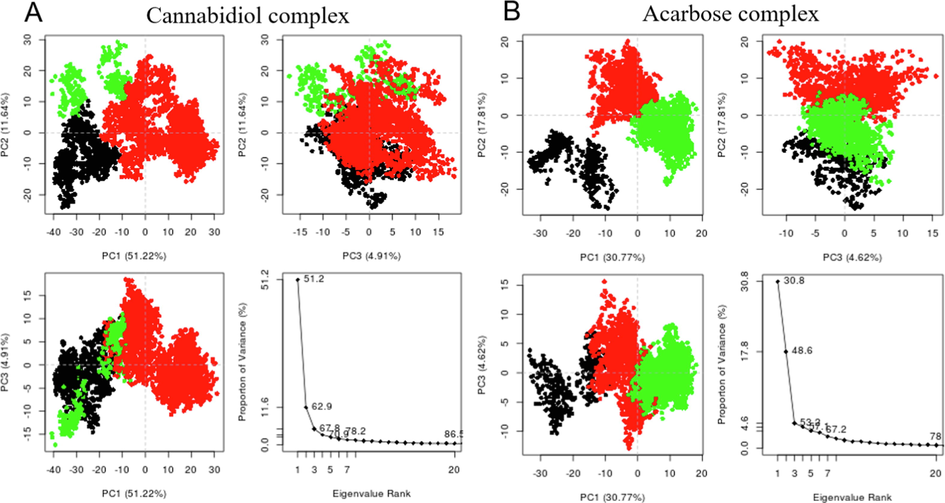
Clustering and principal component analysis on 100 ns of equidistance conformations using the Bio3D package in R. The plots show the first three eigenvectors for (A) Cannabidiol-α-glucosidase complex and (B) Acarbose-α-glucosidase complex. Conformers are colored according to the k-means clustering: cluster 1, black; 2, green; 3, red. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
4 Conclusion
Taken together, these results indicate the antioxidant and antidiabetic properties of cannabidiol. The antioxidant activity can be attributed to the abstraction of H1 in cannabidiol by the DPPH radical, and delocalisation of the radical’s unpaired electron on its (cannabidiol) benzene ring. Its antidiabetic activity can be attributed to the potent inhibitory effect on α-glucosidase, as well as its high affinity with the enzyme. Further studies in in vivo models are recommended to decipher these mechanisms of cannabidiol in type 2 diabetes.
Funding
Dr. Erukainure OL was supported by the University of the Free State, Bloemfontein, South Africa for Incentives for Rated Researchers (2019060769); and the National Research Foundation (NRF) for Scarce Skills Postdoctoral Research Grant (UID: 132822). Prof. Matsabisa MG was supported by the IKS Based Technology Innovation Unit of DSI South Africa, for financial support (Grant contracts: DST/CON 0162/201 and DST/CON 0206/2019/2020).
Acknowledgement
We appreciate the staff and students of the Indigenous Knowledge System (IKS) division, Department of Pharmacology, University of the Free State, Bloemfontein, South Africa for their support and assistance towards this study.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Antioxidant properties and in vitro α-amylase and α-glucosidase inhibitory properties of phenolics constituents from different varieties of Corchorus spp. J. Taibah Univ. Med. Sci.. 2015;10:278-287.
- [Google Scholar]
- Turning to computer-aided drug design in the treatment of diffuse large B-cell lymphoma: has it been helpful? Anticancer Agents Med. Chem.. 2019;19:1325-1339.
- [Google Scholar]
- Covalent versus non-covalent enzyme inhibition: which route should we take? A justification of the good and bad from molecular modelling perspective. Protein J.. 2020;39:97-105.
- [Google Scholar]
- Antioxidative and anti-inflammatory properties of cannabidiol. Antioxidants. 2019;9:21.
- [Google Scholar]
- Cannabidiol protects keratinocyte cell membranes following exposure to UVB and hydrogen peroxide. Redox Biol.. 2020;36:101613
- [Google Scholar]
- Antioxidative and anti-inflammatory properties of cannabidiol. Antioxidants. 2020;9:21.
- [Google Scholar]
- Oxidative stress in the pathophysiology of type 2 diabetes and related complications: Current therapeutics strategies and future perspectives. Free Radic. Biol. Med. 2022
- [Google Scholar]
- Computational systems pharmacology analysis of cannabidiol: a combination of chemogenomics-knowledgebase network analysis and integrated in silico modeling and simulation. Acta Pharmacol. Sin.. 2019;40:374-386.
- [Google Scholar]
- Total syntheses of casuarine and its 6-O-α-glucoside: complementary inhibition towards glycoside hydrolases of the GH31 and GH37 families. Chem.–A Eur. J.. 2009;15:1627-1636.
- [Google Scholar]
- Resilience of S309 and AZD7442 monoclonal antibody treatments against infection by SARS-CoV-2 Omicron lineage strains. Nat. Commun.. 2022;13:1-11.
- [Google Scholar]
- CBG, CBD, Δ9-THC, CBN, CBGA, CBDA and Δ9-THCA as antioxidant agents and their intervention abilities in antioxidant action. Fitoterapia. 2021;152:104915.
- [Google Scholar]
- A Comparative DFT Study on the Antioxidant Activity of Some Novel 3-Hydroxypyridine-4-One Derivatives. Chem. Biodivers.. 2022;19:e202100703.
- [Google Scholar]
- GaussView, version 6.0. 16. Semichem Inc Shawnee Mission KS; 2016.
- A review of alpha-glucosidase inhibitors from plants as potential candidates for the treatment of type-2 diabetes. Phytochem. Rev. 2021:1-31.
- [Google Scholar]
- Software for the frontiers of quantum chemistry: An overview of developments in the Q-Chem 5 package. J. Chem. Phys.. 2021;155:084801.
- [Google Scholar]
- Dacryodes edulis enhances antioxidant activities, suppresses DNA fragmentation in oxidative pancreatic and hepatic injuries; and inhibits carbohydrate digestive enzymes linked to type 2 diabetes. Biomed. Pharmacother.. 2017;96:37-47.
- [Google Scholar]
- Current Protocols in Protein Science, Chapter 2, Unit 2.9. New York: John Wiley & Sons; 2007. [Google Scholar]
- Frisch, M.E., Trucks, G., Schlegel, H., Scuseria, G., Robb, M., Cheeseman, J., Scalmani, G., Barone, V., Petersson, G., Nakatsuji, H., 2016. Gaussian 16. Gaussian, Inc. Wallingford, CT.
- Cannabinoids and terpenes for diabetes mellitus and its complications: from mechanisms to new therapies. Trends Endocrinol. Metab.. 2022;33:828-849.
- [Google Scholar]
- Free energy calculations by the molecular mechanics Poisson− Boltzmann surface area method. Mol. Inf.. 2012;31:114-122.
- [Google Scholar]
- An overview on the role of bioactive α-glucosidase inhibitors in ameliorating diabetic complications. Food Chem. Toxicol.. 2020;145:111738.
- [Google Scholar]
- Therapeutic potential of α-glucosidase inhibitors in type 2 diabetes mellitus: an evidence-based review. Expert Opin. Pharmacother.. 2015;16:1959-1981.
- [Google Scholar]
- Health care costs associated with incident complications in patients with type 2 diabetes in Germany. Diabetes Care. 2018;41:971-978.
- [Google Scholar]
- Cost-related medication non-adherence among US adults with diabetes. Diabetes Res. Clin. Pract.. 2018;143:24-33.
- [Google Scholar]
- Antioxidant and anti-apoptotic effects of cannabidiol in model of ischemic stroke in rats. Brain Res. Bull.. 2022;180:118-130.
- [Google Scholar]
- Computational analysis reveals the association of threonine 118 methionine mutation in PMP22 resulting in CMT-1A. Adv. Bioinforma. 2014
- [CrossRef] [Google Scholar]
- The molecular docking of 1, 4-naphthoquinone derivatives as inhibitors of Polo-like kinase 1 using Molegro Virtual Docker. J. Appl. Pharmaceut. Sci.. 2014;4:047-053.
- [Google Scholar]
- Latunde-Dada, G.O., 2017. Ferroptosis: role of lipid peroxidation, iron and ferritinophagy. Biochimica et Biophysica Acta (BBA)-General Subjects 1861, 1893–1900.
- Transport properties of puerarin and effect of Radix Angelicae Dahuricae extract on the transport of puerarin in Caco-2 cell model. J. Ethnopharmacol.. 2012;144:677-682.
- [Google Scholar]
- Thermodynamics of primary antioxidant action of flavonols in polar solvents. Acta Chim. Slovaca. 2019;12:108-118.
- [Google Scholar]
- The antioxidant and antidiabetic potentials of polyphenolic-rich extracts of Cyperus rotundus (Linn.) J. Biomol. Struct. Dyn.. 2022;40:12075-12087.
- [Google Scholar]
- Inhibition of enzymes linked to type-2 diabetes and hypertension by essential oils from peels of orange and lemon. Int. J. Food Prop.. 2017;20:S586-S594.
- [Google Scholar]
- Why is quercetin a better antioxidant than taxifolin? Theoretical study of mechanisms involving activated forms. J. Mol. Model.. 2013;19:2165-2172.
- [Google Scholar]
- Phytochemical constituents, antioxidant and antidiabetic activities of different extracts of the leaves, stem and root barks of Alstonia boonei: an in vitro and in silico study. Botany Lett.. 2019;166:444-456.
- [Google Scholar]
- Systematic review of efficacy and safety of newer antidiabetic drugs approved from 2013 to 2017 in controlling HbA1c in diabetes patients. Pharmacy. 2018;6:57.
- [Google Scholar]
- Fruit, vegetables, and mushrooms for the preparation of extracts with α-amylase and α-glucosidase inhibition properties: A review. Food Chem.. 2021;338:128119.
- [Google Scholar]
- Cannabidiol modulation of oxidative stress and signalling. Neuronal Signaling 2021:5.
- [Google Scholar]
- UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem.. 2004;25:1605-1612.
- [Google Scholar]
- Accurate ionization potentials and electron affinities of acceptor molecules I. Reference data at the CCSD (T) complete basis set limit. J. Chem. Theory Comput.. 2016;12:595-604.
- [Google Scholar]
- PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput.. 2013;9:3084-3095.
- [Google Scholar]
- Comparative assessment of antimicrobial, antiradical and cytotoxic activities of cannabidiol and its propyl analogue cannabidivarin. Sci. Rep.. 2021;11:1-13.
- [Google Scholar]
- Ferulic acid modulates dysfunctional metabolic pathways and purinergic activities, while stalling redox imbalance and cholinergic activities in oxidative brain injury. Neurotox. Res. 2019:1-12.
- [Google Scholar]
- OriginPro 9.1: scientific data analysis and graphing software-software review. J. Chem. Inf. Model.. 2014;54:1552.
- [Google Scholar]
- α-Glucosidase inhibitory activity of cannabidiol, tetrahydrocannabinol and standardized cannabinoid extracts from Cannabis sativa. Current Res. Food Sci.. 2022;5:1091-1097.
- [Google Scholar]
- Preliminary study: Comparison of antioxidant activity of cannabidiol (CBD) and α-tocopherol added to refined olive and sunflower oils. Molecules. 2019;24:3485.
- [Google Scholar]
- Unuofin, J.O., Lebelo, S.L., 2020. Antioxidant effects and mechanisms of medicinal plants and their bioactive compounds for the prevention and treatment of type 2 diabetes: an updated review. Oxidative Med. Cell. Longevity 2020.
- Wang, W., Wolf, R., J.; Caldwell, J.K., PA; Case DA, 2004. Development and testing of a general amber force field. J. Comput. Chem. 25, 92.
- Theoretical study on the structural and antioxidant properties of some recently synthesised 2, 4, 5-trimethoxy chalcones. Food Chem.. 2015;171:89-97.
- [Google Scholar]
- Phytochemical properties of black tea (Camellia sinensis) and rooibos tea (Aspalathus linearis); and their modulatory effects on key hyperglycaemic processes and oxidative stress. J. Food Sci. Technol.. 2020;57:4345-4354.
- [Google Scholar]
- A review of the molecular mechanisms of hyperglycemia-induced free radical generation leading to oxidative stress. J. Cell. Physiol.. 2019;234:1300-1312.
- [Google Scholar]
- Radical scavenging activity of puerarin: a theoretical study. Antioxidants. 2019;8:590.
- [Google Scholar]
- Cannabidiol improves metabolic dysfunction in middle-aged diabetic rats submitted to a chronic cerebral hypoperfusion. Chem. Biol. Interact.. 2019;312:108819.
- [Google Scholar]







