Translate this page into:
DNA binding, cleavage, catalytic, magnetic active; 2,2–bipyridyl based d-f hetero binuclear Gd(III), Cu(II) complexes and their Electrochemical, fluorescence studies
⁎Corresponding authors at: Faculty, International Ph.D. Program in Innovative Technology of Biomedical Engineering and Medical Devices, Ming Chi University of Technology, New Taipei City 243303, Taiwan (Mani Govindasamy). vijaywarmarajt@gmail.com (A. Vijayaraj), govindasamy420700@gmail.com (Mani Govindasamy)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
Several 2,2-bipyridyl-based d-f heterobinuclear [GdCuL1-5(bpy)2(NO3)2] complexes are present, where (Ligand 1) (9E)-N1-(2-Hydroxy-5-methylbenzylidene)–N2-((E)-2-(2-hydroxy-5-methyl benzylideneamino)ethyl)ethane-1,2-diamine. (Ligand 2) N1,N1-bis((E)-2-(2-hydroxy-5-methylbenzylideneamino)ethyl)ethane-1,2-diamine. (Ligand 3) (9E)-N1-(2-((E)-2-(2-hydroxy-5-methylbenzylideneamino)ethylamino)ethyl)–N2-(2-hydroxy-5-methylbenzylidene)ethane-1,2-diamine. (Ligand 4) (9E)-N1-(2-((E)-3-(2-hydroxy-5- methylbenzylideneamino) propylamino) ethyl)–N3-(2-hydroxy-5-methylbenzylidene)propane-1,3-diamine and (Ligand 5) (9E)-N-(2-hydroxy-5-methylbenzylidene)-3-(4-((E)-3-(2-hydroxy-5-methylbenzylideneamino)propyl)piperazin-1-yl)propan-1-amine. These compounds were described using spectroscopy and the elemental analysis method. Researches were conducted into the luminous, Genetic code, catalytic, magnetism, and breaking attributes of the [GdCuL1-5(bpy)2(NO3)2] complexes. In DMF with 0.1 M tetra-n-butylammonium perchlorate, the binuclear [GdCuL1-5(bpy)2(NO3)2] network complexes exhibit two one electron irreversible reduction events. VSM was used to calculate the complexes' magnetic susceptibility. There is ferromagnetic coupling in the [GdCuL1-5(bpy)2(NO3)2] complexes. The [GdCuL1-5(bpy)2(NO3)2] complexes' excited state lifetimes lengthen in the following order: [GdCuL5(bpy)2] [GdCuL1(bpy)2(NO3)2] [GdCuL3(bpy)2(NO3)2] [GdCuL4(bpy)2] and [GdCuL2(bpy)2(NO3)2]. The binuclear [GdCuL1-5(bpy)2(NO3)2] complexes' inceptive rate of progress for oxidizing 1,2-benzenediol to cyclohexa-3,5-diene-1,2-dione are longer chains with higher activity. Both the [GdCuL5(bpy)2(NO3)2] and [GdCuL4(bpy)2(NO3)2] complexes have strong DNA genetic code properties in the calf genus thymus. The complexes exhibit considerable singlet oxygen-mediated oxidative rift of circular recombinant plasmid pBR322 cloning vector in the existence of 2-sulfanylethanol.
Keywords
Gadolinium
Lifetime
Electrochemical properties
DNA Cleavage
Catecholase activity
DNA binding
1 Introduction
Numerous Schiff Base 2,2-bipyridyl based d-f hetero binuclear metal complexes can be employed for the construction of structural ferromagnetisms, as precursors for a range of Schiff Base processes, analogues of reaction centers of optical materials, luminous materials, DNA binding and dissociation reagents, metalloproteinase, etc. (Rahman et al., 2019) Transition metal complexes with diverse coordination environments and a range of spectroscopic and electrochemical properties substantially improve the customization options for species that are appropriate for catechol, DNA binding, and breakage activities (Chen et al., 1995). This has triggered an increase in curiosity in the synthesis of asymmetric and symmetric binuclear Gd(III)Cu(II) complexes in the past decade. Aside from their fascinating electrochemical and photo-physical properties, they also exhibit deterministic coordination tendencies, 2,2-bipyridines (bpy) have received a lot of research (Neidle et al., 1997). Since they may catalyst reduction and oxidation reactions while encapsulating a variety of substrates under visible light irradiation, d-f Gd(III)Cu(II) complexes have attracted a lot of attention.
The processes by which metalloproteinase function have been clarified thanks to spectroscopic tags, functional models, and metal complexes that include the active sites of proteins. Binucleating ligands are especially appropriate for the synthesis of such complexes as they're more persistent and the Gd(III), Cu(II) two metal ions are anchored firmly together. This has significant implications for metal–metal interactions and their activity. These attributes have allowed chemical nucleases to be adopted as nucleic acid targeting and separating agents (Arcamone et al., 1964) as well as adjuvants in PCR diagnostics (Finlay at al., 1951).
Gadolinium(III) copper(II) side-off unilateral and asymmetrical binuclear clusters have been produced. Various operating environments are used by the imine and 2,2-bipyridyl moieties to co - ordinate the Gd(III) and Cu(II) ions. Some of the Gd(III) ions is coupled to an acyclic imine compartment, and the Cu(II) ion aligns to 2,2-bipyridyl moieties. The mono nuclear acyclic Gd(III) complexes have already been reported (Suckling et al., 2012). Here, we outline the preparation of side-off asymmetric binuclear Gd(III)Cu(II) complexes (Scheme 1) and explore their luminescent, electro-chemical, catecholase activity, magnetic, DNA bonding, DNA breakage, and anti - microbial activities.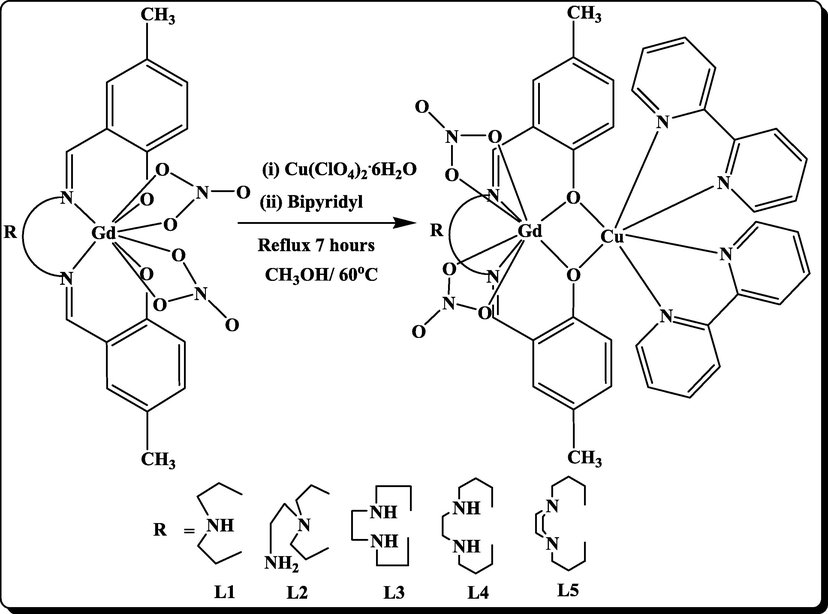
Schematic diagram for the synthesis of binuclear Complexes.
2 Experimentation techniques:
2.1 Solvent & starting material
The method outlined in the literature was utilized to produce 5-methylsalicylaldehyde (Kurmis et al., 2017). Gadolinium nitrate, 2,2-bipyridyl, bought from Aldrich. We bought acetonitrile, dimethyl form amide, and methanol of analytical quality from Qualigens. TBAP is used in electrochemical analysis as a supporting electrolyte and is acquired from Fluka and recrystallized with hot methanol. We procured N, N-bis-(3-aminopropyl) piperazine, N, N-bis-(3-aminopropyl) ethylene diamine, and tris-(2-aminoethyl) amine from Aldrich. We bought copper corbanet, diethylenediamine, and triethylenetetramine from Qualigens.
2.2 Physical and analytical measurements:
The complexes are elementally analyzed on a Heraeus CHN rapid analyzer. An Elico digital conductivity bridge modal CM-88 is employed to analyze the conductivity of the complexes using freshly prepared solutions of the complexes in DMF. The FT-IR spectrum was obtained using a Perkin Elmer FT-IR 8300 series spectrophotometer on KBr discs with thicknesses ranging from 4000 to 400 cm−1. On a Perkin-Elmer UV-320 spectrophotometer, electronic spectrum studies were captured between 200 and 1100 nm. ESI The samples had a concentration of around 1.0 mol dm-3, and mass spectral data were acquired using a JEOL DX-303 mass spectrophotometer. The diluted solution was electro-sprayed at a needle voltage of + 4.5 kV and a flow velocity of 510–6 dm−3 min−1.
2.3 Typical electrochemical study procedure
A three-electrochemical cell with a working electrode composed of glassy carbon, a reference electrode made of Ag-AgCl, and an auxiliary electrode made of platinum wire was used for the cyclic voltammetric trials on a CHI-600A electrochemical analyzer in an oxygen-free environment. Under the experimental conditions, the ferrocene/ferrocenium(Fc/Fc + ) couple's oxidation potential was 470 mV. As the inert electrolyte, tetra-n-butylammonium perchlorate (TBAP) was employed. At a temperature of 25 0.1 °C, experiments were conducted in a pure nitrogen environment. All of the complexes were present in 0.1 mol dm-3 TBAP (tetra-butyl ammonium perchlorate) DMF solutions at concentrations of 1.05 10–3 mol dm-3. Before applying the voltage, the solutions were deaerated for approximately 15 min. With measured uncertainties of 2 mV, the half wave potentials, E1/2, were roughly derived from (Epa + Epc)/2.
2.4 General procedure DNA cleavage studies
Using pBR 322, agarose gel electrophoresis has been used to examine the cleavage activity of complexes. In different concentrations (20 M, 40 M, 60 M, 80 M, and 100 M) supplemented with 5 M H2O2, 0.2 G/L of pBR 322 DNA and 100 M Tris-Hcl (pH 8.0), and the [GdCuL1-5(bpy)2(NO3)2] complexes were used to treat the DNA. All of the samples underwent a 2-hour incubation period at RT in both darkness and light. The DNA concentration was measured by a spectrometer following the incubation period, and the results were represented in g/l units. The remaining samples were loaded using 6X gel loading buffer and 1 % agarose gel with 1 g/ml ethidium bromide. In the TBE buffer, the electrophoresis was run for 1.5 h at 100 V. UV light was used to see the bands, and the Geldoc 100 (Biorad) equipment was used to take photos of them.
2.5 Synthesis of bipyridine based d-f hetero binuclear complexes
The d-f binuclear complex was created using a conventional process, a steady addition of Cu(ClO4)2·.6H2O in ethanol (0.0925 g, 0.25 m.mol) toward a rapidly agitated suspensions of the mononuclear complex in methanol (10 mL) resulted in a clear solution after 15 min. The aforementioned solution was then added dropwise to a methanolic solution, to which 5 mL of 2,2-bipyridyl (0.07809 g, 0.5 mmol) was blended. The resultant solution was then agitated for 3 h at 25 °C. When the solvent evaporated at room temperature, a solid with a dark green colour separated, and acetonitrile was reconstituted from this solid.
2.5.1 [GdCuL1(bpy)2(NO3)2](ClO4) yellowish green solid
The yield of 0.0790 g, or 59 %, of yield. Data for [C40H38ClN9O12CuGd] analysis; mass number: 1093.03 Determined (%): Found (%): C-48.12, H-4.15, N-12.54, Cu-6.24, Gd-15.67; C-48.26, H-4.05, N-12.66, Cu-6.38, and Gd-15.79. chosen IR (KBr, cm−1) 1637 s [v(C⚌N)]; 1522 s [(C—O)]; 1221 s [(NO3–)]; 1075 s [(ClO4)]; 1107 s [(ClO4-) uncoordinated]; 612 s [(M−O)]; ESI- MS (m/z) (%): [GdCuL1(bpy)2(NO3)2]+: 995.15 [MH + UV–vis [max, nm (M−1 cm−1)]: 589 (1 5 1) 358 (17735) 320 (19836) 270 (22336); Conductance (m, S cm2 mol−1); in DMF: 121.
2.5.2 [GdCuL2(bpy)2(NO3)2](ClO4)2 yellowish green solid
0.070 g, or 49 %, of yield. Data for [C42H44Cl2N9O13CuGd], M.Wt. 1174.57 Calculated (%) values were C-51.70; H-4.55; N-12.92; Cu-6.51; and Gd-16.12; while actual values were C-51.68; H-4.54; N; Cu; and Gd-16.10; specific IR (KBr, cm-1) 1637 s [v(C⚌N)], 1522 s [C—O], 1211 s [NO3–], 1075 s [ClO4], 1078 s [ClO4-) uncoordinated], and 612 s [M−O]; ESI- MS (m/z (%): [GdCuL2(bpy)2(NO3)] 2+: 487.83 [MH + ], calculated at 975.67 m/z; UV–vis [max, nm (M−1 cm−1)] in DMF-594 (1 4 3) 363 (19686) 276 (22036); Conductance (m, S cm2 mol−1) in DMF: 165.
2.5.3 [GdCuL3(bpy)2(NO3)2](ClO4)2 yellowish green solid
0.070 g (49 %), yield. Data for [C42H42Cl2N9O13CuGd] analysis; mass: 1172.54; Found (%): C-51.80, H-4.34, N-12.94, Cu-6.52, and Gd-16.13; calculated (%): C-51.81, H-4.35, N-12.95, Cu-6.53, and Gd-16.15. chosen IR (KBr, cm-1) 1637 s [v(C⚌N)], 1522 s [C—O]; 1203 s [NO3–]; 1075 s [ClO4]; 1100 s [ClO4-] uncoordinated; 612 s [M−O]; ESI- MS (m/z (%): [GdCuL3(bpy)2(NO3)] 484.73 [MH-], Calcd (m/z) 973.64; UV–vis [max, nm, M−1 cm−1] in DMF: 611 (1 3 8) 375 (16885) 303 (20686) 278 (21937); Conductance (m, S cm2 mol−1) in DMF: 182. 2 +.
2.5.4 [GdCuL4(bpy)2(NO3)2](ClO4)2 Yellowish-green solid
51 % yield, or 0.087 g. Data for [C44H46Cl2N9O13CuGd] analysis; mass: 1200.58 Calculated (%) values are 52.76C, 4.63H, 12.58 N, 6.34 Cu, and 15.70 Gd, while actual values are 52.74C, 4.62H, 12.56 N, 6.32 Cu, and 15.69 Gd. chosen IR (KBr, cm−1) 1638 s [v(C⚌N)], 1535 s [C—O], 1143 s [NO3–], 1075 s [(ClO4-) uncoordinated], and 622 s [(M−O)]; ESI- MS [GdCuL4(bpy)2] 499.87 [MH-], Calcd (m/z) 1001.68.; UV–vis [max, nm M−1 cm−1] in DMF: 635 (1 3 2) 383 (16485) 310 (20336) 282 (21737); Conductance (m, S cm2 mol−1) in DMF: 173. 2+:
2.5.5 [GdCuL5(bpy)2(NO3)2](ClO4)2 green solid
Yield: 0.087 g, or 51 %; analytical statistics for [C46H50Cl2N9O13CuGd]; M.Wt: 1228.64; calculated (%): C, 53.65; H, 4.89; N, 12.24; Cu, 6.17; Gd, 15.27; found (%): C, 53.64; H, 4.88; N, 12.23; Cu, 6.15; Gd, 15.26; chosen IR (KBr, cm−1) 1624 s [v(C⚌N)]; 1523 s [C—O]; 1134 s [NO3–]; 1047 w [(ClO4-) uncoordinated] 656 m [(M−O)]; ESI- MS (m/z) (%): [GdCuL5(bpy)2] 513.87 [MH-], Calcd (m/z) 1029.74.; UV–vis [max, nm, M−1 cm−1] in DMF: 650 (1 2 4) 390 (16134) 305 (20586) 285 (21587); Conductance (m, S cm2 mol−1) in DMF: 181. 2+:
3 Results and discussion
All the complexes obtained were crystalline powders that are stable in air and insensitive to light. d-f hetero binuclear Gadolinium(III), copper(II) complex is sparingly soluble in water but other complexes were insoluble. They were also insoluble in organic solvents such as ethanol, ether, and benzene, but soluble in DMSO. All the spectral data given the Tables 1-6.
No
Complexes
λmax (nm) (ε/M−1 cm−1)
d-d
charge transfer
1.
[GdCuL1(bpy)2(NO3)2]
589(1 5 1)
358(17735) 320(19836) 270(22336)
2.
[GdCuL2(bpy)2(NO3)2]
594(1 4 3)
363(17485) 323(19686) 276(22036)
3.
[GdCuL3(bpy)2(NO3)2]
611(1 3 8)
375(16885) 303(20686) 278(21937)
4.
[GdCuL4(bpy)2(NO3)2]
635(1 3 2)
383(16485) 310(20336) 282(21737)
5.
[GdCuL5(bpy)2(NO3)2]
650(1 2 4)
390(16134) 305(20586) 285(21587)
Complexes
τ1/ns
α1 (%)
τ2/ns
α2 (%)
τ3 /ns
α3 (%)
χ2
[GdCuL1(bpy)2(NO3)2]
0.953
21.84
2.150
44.58
8.253
33.58
0.979
[GdCuL2(bpy)2(NO3)2]
1.192
44.33
2.544
43.03
6.563
12.65
1.064
[GdCuL3(bpy)2(NO3)2]
0.892
24.71
2.753
48.96
6.933
26.33
1.000
[GdCuL4(bpy)2(NO3)2]
0.991
53.92
2.150
28.56
6.835
17.52
1.013
[GdCuL5(bpy)2(NO3)2]
0.453
18.12
1.434
56.68
4.903
25.20
0.928
Complexes
Epc1/V
Epc2/V
[GdCuL1(bpy)2(NO3)2]
1.62
2.20
[GdCuL2(bpy)2(NO3)2]
1.55
2.02
[GdCuL3(bpy)2(NO3)2]
1.43
1.90
[GdCuL4(bpy)2(NO3)2]
1.21
1.85
[GdCuL5(bpy)2(NO3)2]
1.10
1.80
Complexes
Rate constant (k) × 10- 3 min−1
Catecholase activity
[GdCuL1(bpy)2(NO3)2]
3.62
[GdCuL2(bpy)2(NO3)2]
4.12
[GdCuL3(bpy)2(NO3)2]
5.97
[GdCuL4(bpy)2(NO3)2]
6.45
[GdCuL5(bpy)2(NO3)2]
7.98
Complexes
Kb(M−1)
Kb(app)(M−1)
[GdCuL1(bpy)2(NO3)2]
2.96 × 105
1.67 × 106
[GdCuL2(bpy)2(NO3)2]
3.56 × 105
2.74 × 106
[GdCuL3(bpy)2(NO3)2]
2.83 × 105
1.56 × 106
[GdCuL4(bpy)2(NO3)2]
3.32 × 105
2.49 × 106
[GdCuL5(bpy)2(NO3)2]
3.31 × 105
2.48 × 106
S.No
Samples
O.D Value
Concentration (ng/µl)
1
Control
1.83
2500
2
[GdCuL1(bpy)2(NO3)2]
1.75
498.8
3
[GdCuL2(bpy)2(NO3)2]
1.71
477.9
4.
[GdCuL3(bpy)2(NO3)2]
1.63
455.5
5.
[GdCuL4(bpy)2(NO3)2]
1.57
444.6
6.
[GdCuL5(bpy)2(NO3)2]
1.49
389.6
3.1 IR spectra
The coordinated Schiff-base type (bpy) ligands' predicted bands are all present in the complexes' infrared (IR) spectra. Coordinated C⚌N- and C—O- stretching are ascribed to the spectrum at 1585 cm−1 and 1245 cm−1, respectively. Imine nitrogen and phenolic C—O—mediated coordination of such ligand toward the metal is predicted to significantly decrease the electron density in the imine and lower v(C⚌N) to 1623 cm−1 and 1239 cm−1, respectively, demonstrating that the ligand‘s imine nitrogen is firmly synchronized to the metal center (Munde et al., 2014). The absence of a moderately intense spectrum at 3330 cm−1 that was seen in the unobstructed ligand due to (OH) indicates that the ligand was deprotonated before it was coordinated (Ashley et al., 1942). Absorption at 420–456 cm−1 for the complexes could be attributed to the -(M−O) bond. Sharp peaks seen between 1335 and 1410 cm−1 are attributed to complexes containing v(Gd-NO3) (Arafa et al., 2011). The -(M−N) bond may be responsible for additional weak bands with lower frequency. Due to the aromatic C—H stretching vibration, bonds at 3155–3060 cm−1 dominate the spectra of all the complexes (Opperman et al., 2016). A single spectrum at 1110 cm−1 (3-antisymmetric stretching) that is not split and a peak around 650 cm−1 (4-antisymmetric bending) indicate the presence of unsynchronized perchlorate anions in all of the binuclear complexes. Fig. 1 displays the FT-IR spectra.![FT-IR spectrum of. [GdCuL1(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig2.png)
FT-IR spectrum of. [GdCuL1(bpy)2(NO3)2].
3.2 ESI mass spectral exploration
The parent ion peak (Mn + ) is visible in the ESI mass spectrum of the binuclear complexes [GdCuL3(bpy)2(NO3)2](ClO4)2 and [GdCuL4(bpy)2(NO3)2](ClO4)2 at (m/z) = 484.73 and 499.87, respectively. The complexes' diverse pieces are represented by certain noticeable peaks in the spectra. Figs. 2a and 2b display the ESI mass spectrum of the binuclear complexes [GdCuL3(bpy)2(NO3)] and [GdCuL4(bpy)2(NO3)2] respectively.![ESI Mass spectrum of. [GdCuL3(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig3.png)
ESI Mass spectrum of. [GdCuL3(bpy)2(NO3)2].
![ESI Mass spectrum of. [GdCuL4(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig4.png)
ESI Mass spectrum of. [GdCuL4(bpy)2(NO3)2].
3.3 Electronic spectral analysis of [Gd(III)Cu(II)L1-5(bpy)2(NO3)2] complexes
In DMF medium, electronic spectrum of all the complexes were collected. The dx2-y2 dxy yz; dz2; dxy linked with deformed octahedral geometry around the Cu(II) ion causes a single weak d-d band in the area 573–650 nm, which is visible in the electronic spectrum of all the complexes. It has been noted that an increase in chain length between imine nitrogen causes a red shift in the maximum value of the d-d band width. As the chain length increases, the distortion from planar geometry may be the cause of this red shift. The ligand to metal charge transfer transition is linked to the moderately intense band seen in the range of 358–394 nm (Kahvedzič et al., 2016). The intra ligand charge transfer transition π–π*, which has a peak in the 270–285 nm region, is thought to be responsible. Figs. 3–5 provide the electronic spectra for each hetero binuclear [GdCuL1-5(bpy)2(NO3)2] complex. Table 1 contains electronic spectrum information for all binuclear [GdCuL1-5(bpy)2(NO3)2] complexes.![Electronic spectra of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig5.png)
Electronic spectra of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
![Electronic spectra of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig6.png)
Electronic spectra of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
![Emission spectra of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig7.png)
Emission spectra of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
3.4 Fluorescence spectra
Fig. 5 displays the complex [GdCuL1-5(bpy)2(NO3)2] emission spectra under excitation at 430 nm. The connection between structure and attributes is what we find interesting. Thus, the complexes of fluorescence properties are found. The efficiency of the intramolecular transfer of energy between the triplet levels of such ligands as well as the emitting state of the ions, particularly relies on the energy gap between both levels, is associated with the luminescence of Ln3 + chelates. The ligand's triplet state energy is thought to have a significant role in the lanthanide ion's stimulation during intramolecular energy transfer (Lansiaux et al., 2002). There is a smaller emission band in the 400–700 nm range compared to compound 3 [GdCuL3(bpy)2(NO3)2]. indicating that the ligands 3 totally suppressed the fluorescence of the Gd, Cu ion in the cavity (Nagle et al., 2012). However, the [GdCuL1,2,4 and 5] emission band in the 430–700 nm region has a larger emission intensity. However, [GdCuL1-5(bpy)2(NO3)2] distinctive emission spectrum was noticed. The 6Dj level, which corresponds to the 8S7/2 6Dj transition, is the source of all emissions. The 8S7/2 6Dj transition generates the weak band at 490 nm. A change in the coordination environment has no impact on the band for 8S7/2 6Dj at 470 nm, which is magnetic-dipole permissible. The emission intensity of the strong band at 500 nm for 8S7/2 6Dj is dependent on the coordination environment of Gd and represents an electric-dipole allowed transition (III). This demonstrates that the ligand center had an impact on the fluorescence emission of Gd(III) ions within the complex.
3.5 Life time measurement [GdCuL1-5(bpy)2(NO3)2] complexes
Due to advancements in the construction and miniaturization of the lasers and electronics required for Time Correlated Single Photon Counting (TCSPC) measurements of fluorescence lifetime, the application of the time domain technique has become simpler. This fitting demonstrates unequivocally that the complexes are all tri exponentially fitted compare to (Nagle et al., 2014). The [GdCuL1-5(bpy)2(NO3)2] complexes' emission spectra and decay time observations allowed the credentials of the highest ligand triplet state depicted in Fig. 6. Compared to other mono and binuclear nuclear complexes, the [GdCuL2(bpy)2(NO3)2] complex has a high life time value. The life time is raised in order.![Life time decay plot of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig8.png)
Life time decay plot of thebinuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
[GdCuL5(bpy)2(NO3)2] < [GdCuL1(bpy)2(NO3)2] < [GdCuL3(bpy)2(NO3)2] < [GdCuL4(bpy)2(NO3)2] < [GdCuL2(bpy)2(NO3)2].
Table 2 presents the data. The amount of distinct luminous centers, energy transmission, host flaws, and contaminants all affect the exponential decay behavior.
3.6 Cyclic voltammetric behavior
The binuclear complex, [GdCuL1-5(bpy)2(NO3)2] cyclic voltammogram was obtained in DMF medium between 0.0 and 2.3 V at a scan rate of 0.05 V s-1 and is shown in (Fig. 7). Epc1 = 1.62 V and Epc2 = 2.20 V(Ag/AgCl) are the two well determined catholic peak potentials for the complex [GdCuL1]. 1.91 V was calculated to be the half wave potential E1/2 for the first reduction process, Cu(II)/Cu (I). The ligand-centered mono nuclear complex [Gd(III)L1(bpy)2(NO3)2] peak potential may correspond to the second reduction process, which was seen at the cathodic potentials Epc2 2.20 V. It is proposed that the total electrode reaction is LGd(III)Cu(II)/LGd(III)Cu(I)/L•Gd(III)Cu (I). The initial reduction process for all five d-f complexes is Cu(II)/Cu(I) exclusively reduced, and the second reduction potential corresponds to the ligand-centered peak potential of the Gd(III) mono nuclear complex. Table 3 contains the electrochemical data for each complex. Complex from [GdCuL1(bpy)2(NO3)2] to [GdCuL5(bpy)2(NO3)2] reduction potential value shift to more negative potential direction; this could be because the entire cyclic ring becomes more flexible, distorting the geometry of the gadolinium(III) complexes while also increasing the system's flexibility (Vinoth et al., 2022) (Govindasamy et al., 2017, 2019). In Fig.7. a cyclic voltammogram is displayed.![Cyclicvoltammograms of thebinuclear [gdcul1-5(bpy)2(NO3)2]complexes (Reduction process) (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig9.png)
Cyclicvoltammograms of thebinuclear [gdcul1-5(bpy)2(NO3)2]complexes (Reduction process) (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
3.7 Magnetic behavior
3.7.1 Magnetic properties of complex 5. [GdCuL5(bpy)2(NO3)2](ClO4)2
In the temperature range of 20 to 300 K, magnetic susceptibility's (m) temperature dependence is depicted in (Fig. 8). At 300 K, the effective magnetic moment (eff) is 7.78B expressed as MT vs T. The MT value is 7.85 cm3 K mol-1 at 180 K, which approximately equates to the value anticipated for the two uncoupled metal ions. The MT value rises with decreasing temperature, reaching 10.5 cm3 K mol-1 at 20 K. This number compares favorably to the (10.5 cm3 K mol-1) predicted value for the S = 4 spin state produced by the ferromagnetic coupling of Gd(III) (S = 7/2) and Cu(II) (S = 1/2) under the assumption that Gd = Cu = 2.0 is quite similar. Data were collected. On the basis of a spin-only equation generated from a spin Hamiltonian H= - JSCu SGd, a quantitative analysis can be done. The experimental data are based on the formula, taking into account the possibility that the two low-lying H = JSCu SGd levels E(4) = 0 and E(3) = 4 J may have distinct g values (O’Sullivan et al., 2014), [4 = (7Gd + Cu)/8 and g3 =(9 Gd -Cu)/8]. The parameters' calculated values are J = 4.2 cm-1, Cu = 2.05, Gd = 2.0, and R = [(obsT- calT)2/(obs T)2]. There is no question that the observed ferromagnetic behavior is a fundamental characteristic of the core at 3.910–5. The J values for the four structurally defined hetero binuclear (Cu, Gd) complexes are slightly lower than those previously published for GdO2Cu. The complex's structural analysis reveals that a third bridge connects an axial Gd site to an axial Cu site that, at best, has a very weak spin density (Winpenny et al., 1998). The measured J value (4.81 cm-1) is remarkably comparable to values discovered for complexes in which the magnetic interaction is mediated by a double bridge, such as CuO2Gd as in the preceding examples (Li and Liao et al., 1995). Additionally, the extremely good fit for (Fig. 8) that was achieved with the aid of the formula above, which corresponds to a binuclear Gd-Cu complex, conforms to the binuclear nature of the sample material.![Thermal dependence of χMT for [GdCuL5(bpy)2(NO3)2] at 0.5 T. The full line corresponds to the best data fit.](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig10.png)
Thermal dependence of χMT for [GdCuL5(bpy)2(NO3)2] at 0.5 T. The full line corresponds to the best data fit.
3.8 Kinetic studies
3.8.1 Oxidation of pyrocatechol (Catecholase Activity)
Pyrocatechol was used as a simple model substrate to test the catecholase activity of the [GdCuL1-5(bpy)2(NO3)2] complexes created in the current study in order to identify functional models for the metalloenzymes (Wermuth et al., 2011). For this, 100 equivalents of pyrocatechol were added to solutions of complexes in dimethylformamide at a concentration of 10-3 mol dm−3 in the presence of air. At regular intervals of 5 min, the reaction's progress was monitored spectrophotometric ally at 390 nm for about 45 min. The slope was calculated using the initial rates approach by observing the development of the product's O- 390 nm quinine’s band. First-order dependency on the complex concentration is revealed by a linear relationship between the starting rate and the complex concentration for [GdCuL1-5(bpy)2(NO3)2] complexes.
Fig. 9 displays the log (A∞/A∞-At) vs time plots for the catecholase activity of the [GdCuL1-5(bpy)2(NO3)2] complexes. The increase of the o-quinone chromophore in response to [GdCuL5(bpy)2(NO3)2] is depicted in Fig. 9′s inset over time. Table 4 provides the value of the [GdCuL1-5(bpy)2(NO3)2] complexes' observed initial rate constant. Catalytic activity is higher in the complex [GdCuL5(bpy)2(NO3)2] (7.98x10-3 min-1) than in the complex [GdCuL4(bpy)2(NO3)2] (6.45x10-3 min-1), which is higher than the complex [GdCuL1(bpy)2(NO3)2] (3.62x10-3 min-1). Below is a list of the complexes in order of activity.![Catecholase activity of binuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig11.png)
Catecholase activity of binuclear [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
[GdCuL5(bpy)2(NO3)2] > [GdCuL4(bpy)2(NO3)2] > [GdCuL3(bpy)2(NO3)2] > [GdCuL2(bpy)2(NO3)2] > [GdCuL1(bpy)2(NO3)2].
The findings show that when the chain length rises, the rate of catecholase oxidation to o-quinone has risen. Due to flexibility brought on by the coordination sphere's distortion, the catecholase activity of complexes with longer carbon chains in the imine compartment is higher than that of complexes with shorter carbon chains in the imine compartment. The geometry of the complexes is more deformed as the chain length increases. The increased reaction rate that has been seen may be favored by this shape.
3.9 DNA binding and cleavage studies
3.9.1 Absorption spectral studies
Complexes [GdCuL1(bpy)2(NO3)2], [GdCuL2(bpy)2(NO3)2], and [GdCuL3(bpy)2(NO3)2] have binding properties. The effects of [GdCuL4(bpy)2(NO3)2] and [GdCuL5(bpy)2(NO3)2] on the UV spectra of calf thymus (CT) DNA are evaluated. Due to the intercalative mode's strong stacking contact between an aromatic chromophore and the base pairs of DNA, complex binding with DNA by intercalation typically results in bathochromic (Edwards et al., 1992). The interaction of the binuclear [GdCuL1(bpy)2(NO3)2], [GdCuL2(bpy)2(NO3)2], [GdCuL3(bpy)2(NO3)2], [GdCuL4(bpy)2(NO3)2], and [GdCuL5(bpy)2(NO3)2] complexes with CT DNA has been studied in the current study. Using a constant concentration to which increments of the DNA (10 mM) stock solution were added gradually at 25 °C, absorption titration tests of the Gd(III)Cu(II) complexes in buffer were carried out. In the UV–vis absorption spectra, the complexes attachment to duplex DNA resulted in a reduction in absorption intensities and a little redshift. After intercalating DNA base pairs, the intercalated ligand's π* orbital may couple with the base pairs' orbital, lowering the π–π* transition energy and producing bathochromic (Trott et al., 2010).
For the purpose of quantitatively comparing the affinity of the complexes toward DNA, the binding constant K of the complexes to CT DNA was calculated by observing changes in the absorbance of the charge transfer spectral band near at 230 nm, 280 nm, and 350 nm for the complex [GdCuL2(bpy)2(NO3)2], near 233 nm, 275 nm, and 375 nm for the complex [GdCuL3(bpy)2(NO3)2] (b Fig. 10 depicted the complexes' spectroscopic titration. The degree of hypochromic serves as a gauge for the intercalative binding's strength. The order bipyridyl-based binuclear complexes follows the observed trend in hypochromic among the current complexes (GdCu). Following the equation, the intrinsic binding constants Kb of the four complexes with CT DNA were calculated.![Absorption spectra of complex [GdCuL2(bpy)2(NO3)2] (20 µM) in the presence of increasing quantity of CT DNA; 0–80 µM. The arrow shows the absorbance change upon increasing DNA concentrations. Inset shows the plot of (εa – εf)/(εb – εf) vs [DNA] for the titration of DNA with the complex.](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig12.png)
Absorption spectra of complex [GdCuL2(bpy)2(NO3)2] (20 µM) in the presence of increasing quantity of CT DNA; 0–80 µM. The arrow shows the absorbance change upon increasing DNA concentrations. Inset shows the plot of (εa – εf)/(εb – εf) vs [DNA] for the titration of DNA with the complex.
The apparent absorption coefficients a, f, and b correspond to Aobsd/[M2], the extinction coefficient for free complexes, and the extinction coefficient for complexes in the completely bound state, respectively, where [DNA] is the concentration of DNA in base pairs. Kb is determined by plotting [DNA] / (b-f) versus [DNA] and calculating the slope to intercept ratio. The complexes stated above have binding constant values Kb, which are computed and provided in Table 5.
3.9.2 Fluorescence spectral studies
Fluorescence spectroscopy is a useful technique for examining metal interactions with DNA. One of the most sensitive fluorescent probes that can connect with DNA is ethidium bromide (EB) (Yuan et al., 2020). After intercalating into DNA, EB exhibits a greater rise in fluorescence. The amount of EB-available DNA binding sites decreases as a result of the metal intercalating into the DNA, which lowers the fluorescence of the EB-DNA system (Vanpatten et al., 2018). Fig. 11. depicts the emission spectra of EB bound to DNA in the presence and absence of the complexes [GdCuL2(bpy)2(NO3)2], [GdCuL3(bpy)2(NO3)2], [GdCuL4(bpy)2(NO3)2], and [GdCuL5(bpy)2(NO3)2]. At various complex concentrations, 610 nm (510 nm excitation) fluorescence intensities were recorded. 40 M DNA and 0.66 M Ethidium Bromide (EB, at saturation binding level) were dissolved in a 2 mL solution and titrated with 0–80 M metal complexes at 25 °C. The fluorescence intensity considerably decreases when complex DNA that has been pretreated with EB is added, demonstrating that complex competes with EB for DNA binding. The degree to which the emission intensity decreases provides information on the complexes' propensity to bind DNA as well as their ability to stack (intercalate) between neighboring DNA base pairs (Salvador et al., 2012). The fluorescence quenching curve of DNA-bound EB by complexes shows that the linear Stern-Volmert equation and the quenching of EB bound to DNA by complexes are in good accord.![Emission spectrum of EtBr bound to DNA system ([EtBr] = 6.6 µM, [CD DNA] = 40 µM, [GdCuL2(bpy)2(NO3)2] = 0–80 µM, λemi = 510 nm). The arrow shows the intensitychange upon increasing [GdCuL2(bpy)2(NO3)2] concentrations. Inset shows the plot of emission intensity Io /I vs [complex]/[DNA].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig13.png)
Emission spectrum of EtBr bound to DNA system ([EtBr] = 6.6 µM, [CD DNA] = 40 µM, [GdCuL2(bpy)2(NO3)2] = 0–80 µM, λemi = 510 nm). The arrow shows the intensitychange upon increasing [GdCuL2(bpy)2(NO3)2] concentrations. Inset shows the plot of emission intensity Io /I vs [complex]/[DNA].
The slope-to-intercept ratio, or Ksv, in the linear fit plot of I0/I versus [complex]/[DNA], is the measure of the slope. I0 is the emission intensity of EB-DNA when the complex is not present, I is the emission intensity of EB-DNA when the complex is present, and [Q] is the concentration of the quencher. Table 5 provides the Kapp values for complexes.
3.9.3 DNA cleavage study for [GdCuL1-5(bpy)2(NO3)2] complexes
The incubation duration at room temperature was prolonged when using the [GdCuL1-5(bpy)2(NO3)2] complexes, which enhanced the cleavage efficiency. It has been stated that the complex's binding activity with DNA was effective. All chemical complexes exhibit a striking uniformity of expression in that they all cleave DNA by forming cleaved bands. We can therefore bring those compounds for additional therapeutic development. In this article, we show how in vitro screening based on DNA cleavage can be used effectively. When compared to other ligands, the complex [GdCuL1(bpy)2(NO3)2] exhibits significant bioactivity in in vitro infection experiments, regardless of the ligand's location or saturation condition (Lee et al., 2020). These results support the value of complex activity for harmful microbes. The comprehensive work on in vitro investigations and the identification of various bioactive chemical types is abundantly possible with this exploratory inquiry. These inhibiting studies will advance knowledge of the molecular interactions between [GdCuL1-5(bpy)2(NO3)2] complexes and bacterial vectors. The information is in Table 6.
3.9.4 Antimicrobial activity
Studies have been done on the antimicrobial activity of synthetic binuclear complexes [GdCuL1-5(bpy)2(NO3)2] against five investigated bacteria. According to the results, all [GdCuL1-5(bpy)2(NO3)2] binuclear complexes had distinct effects on various species, with Klebsiella sp. being the target of the greatest activity at 125 M. Against all species, the complexes [GdCuL1(bpy)2(NO3)2] and [GdCuL5(bpy)2(NO3)2] show the strongest antibacterial activity. [GdCuL3(bpy)2] shown the least activity, in comparison. This variation may be brought on by the varied structures of [GdCuL1-5(bpy)2(NO3)2] complexes and the activity side chains or by variations in the ligands' binding sites to the molecules (Zhu et al., 2021), comparing of the other Gadolinium complexes, d-f hetero binuclear Gd(III)Cu(II) more active. Tables 7.1–7.5 contains information on [GdCuL1-5(bpy)2(NO3)2] complexes' antimicrobial activity (see Figs. 12a and 12b).![Agarose Gel electrophoresis photograph of the super coiled pBR322 DNA treated with different complexes [GdCuL1-5(bpy)2(NO3)2]. Lane1. Control; Lane (a). pBR322 DNA treated with 100 µM of [GdCuL1(bpy)2(NO3)2] complex; Lane (b). pBR322 DNA treated with 100 µM of [GdCuL2(bpy)2(NO3)2] complex; Lane (c). pBR322 DNA treated with 100 µM of [GdCuL3(bpy)2(NO3)2] complex; Lane (d). pBR322 DNA treated with 100 µM of [GdCuL4(bpy)2(NO3)2] complex; Lane (e). pBR322 DNA treated with 100 µM of [GdCuL5(bpy)2(NO3)2] complex; Form I and II represents Supercoiled, Nicked circular and Linear form of pBR 322 DNA respectively.](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig14.png)
Agarose Gel electrophoresis photograph of the super coiled pBR322 DNA treated with different complexes [GdCuL1-5(bpy)2(NO3)2]. Lane1. Control; Lane (a). pBR322 DNA treated with 100 µM of [GdCuL1(bpy)2(NO3)2] complex; Lane (b). pBR322 DNA treated with 100 µM of [GdCuL2(bpy)2(NO3)2] complex; Lane (c). pBR322 DNA treated with 100 µM of [GdCuL3(bpy)2(NO3)2] complex; Lane (d). pBR322 DNA treated with 100 µM of [GdCuL4(bpy)2(NO3)2] complex; Lane (e). pBR322 DNA treated with 100 µM of [GdCuL5(bpy)2(NO3)2] complex; Form I and II represents Supercoiled, Nicked circular and Linear form of pBR 322 DNA respectively.
![Efficiency of the DNA cleavage for [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].](/content/184/2023/16/5/img/10.1016_j.arabjc.2023.104656-fig15.png)
Efficiency of the DNA cleavage for [GdCuL1-5(bpy)2(NO3)2]complexes (a)[GdCuL1 (bpy)2(NO3)2], (b)[GdCuL2(bpy)2(NO3)2], (c)[GdCuL3(bpy)2(NO3)2], (d)[GdCuL4(bpy)2(NO3)2] and(e) [GdCuL5(bpy)2(NO3)2].
4 Conclusion
The five d-f heterobinuclear Gd(III)Cu(II) complexes based on 2,2′ bi-pyridine have been synthesized and studied. According to the spin Hamiltonian seen, the complex's ferromagnetic behavior is an inherent quality of the core. Due to the lengthening of the chain, [Gd(III)Cu(II)L1-5(bpy)2(NO3)2] complexes' electronic spectra exhibit a red shift. Time correlated single photon counting measurements [GdCuL2(bpy)2(NO3)2] complex having longer life time value. Measurements of the [GdCuL2(bpy)2(NO3)2] complex's electronic absorption and fluorescence spectra show good CT DNA binding, most likely via intercalation. In vitro, the complex [GdCuL1(bpy)2(NO3)2] is more cleavage-active. Methyl group density around the metal improves the catalytic performance. The antibacterial activity of each and every d-f hetrobinuclear Gd(III)Cu(II) complex is enhanced against both gramme positive and gramme negative microorganisms.
CRediT authorship contribution statement
A. Vijayaraj: Conceptualization, Methodology, Writing – original draft. R. Prabu: Resources. R. Suresh: Data curation. R. Sangeetha Kumari: Writing – review & editing. V. Kaviyarasan: Data curation. V. Narayanan: Investigation. P. Tamizhdurai: Formal analysis. Fatmah Ali Alasmary: Methodology, Funding acquisition, Project administration. Umamaheswari Rajaji: Formal analysis, Methodology. Mani Govindasamy: Formal analysis, Investigation, Writing - original draft.
Acknowledgments
This work was funded by the Researchers Supporting Project Number (RSP2023R259) King Saud University, Riyadh, Saudi Arabia. This work was supported by Ming Chi University of Technology, New Taipei City 243303, Taiwan. And financial support from the CSIR New Delhi is gratefully acknowledged and National Centre for Ultrafast Process, University of Madras, for the fluorescence and lifetime measurements.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Molecular modeling study and synthesis of novel dicationic flexible triaryl guanidines and imidamides as antiprotozoal agents. Eur. J. Med. Chem.. 2011;46:5852-5860.
- [Google Scholar]
- A chemotherapeutic comparison of the trypanocidal action of some aromatic diamidines. J. Chem. Soc. 1942:103-116.
- [Google Scholar]
- Crystal structures of B-form DNA-RNA chimers complexed with distamycin. Nat. Struct. Biol.. 1995;2:733-735.
- [Google Scholar]
- Crystal structure of a pentamidine- oligonucleotide complex: implications for DNA-binding properties. Biochemistry. 1992;31:7104-7109.
- [Google Scholar]
- Netropsin, a new antibiotic produced by a Streptomyces. J. Am. Chem. Soc.. 1951;73:341-1334.
- [Google Scholar]
- Highly sensitive determination of non-steroidal anti-inflammatory drug nimesulide using electrochemically reduced graphene oxide nanoribbons. RSC adv. 2017;7:33043-33051.
- [Google Scholar]
- Ultrasound-assisted synthesis of tungsten trioxide entrapped with graphene nanosheets for developing nanomolar electrochemical (hormone) sensor and enhanced sensitivity of the catalytic performance. Ultrason. Sonochem.. 2019;56:134-142.
- [Google Scholar]
- Isouronium and N-hydroxyguanidinium derivatives as Cell growth inhibitors: a comparative study. Eur. J. Med. Chem.. 2016;117:269-282.
- [Google Scholar]
- pyrrole- imidazole polyamide is active against enzalutamide resistant prostate cancer. Cancer Res.. 2017;77:2207-2212.
- [Google Scholar]
- Distribution of furamidine analogues in tumor cells: influence of the number of positive charges. J. Med. Chem.. 2002;45:1994-2002.
- [Google Scholar]
- Inhibitory functions of maslinic acid, a natural triterpene, on HMGB1-mediated septic responses. Phytomedicine. 2020;69:153-200.
- [Google Scholar]
- Structure dependent inhibition of the ETS-family transcription factor PU.1 by novel heterocyclic diamidines. Nucleic Acid Res.. 2014;42:1379-1390.
- [Google Scholar]
- High DNA affinity of a series of amide linked aromatic dications. J. Med. Chem.. 2012;55:4397-4406.
- [Google Scholar]
- Unexpected DNA affinity and sequence selectivity through core rigidity in guanidinium-based minor groove binders. J. Med. Chem.. 2014;57:7663-7672.
- [Google Scholar]
- Cytotoxicity of bis(phenylamidinium)- furan alkyl derivatives in human tumour cell lines: Relation to DNA minor groove binding. Bioorg. Med. Chem.. 1997;7:1403-1408.
- [Google Scholar]
- DNA Targeting as a likely mechanism underlying the antibacterial activity of synthetic bis-indole antibiotics. Antimicrob. Agents Chemother.. 2016;60:7067-7076.
- [Google Scholar]
- Recent developments in compounds acting in the DNA minor groove. Med. Chem. Commun.. 2019;10:26-40.
- [Google Scholar]
- Salvador, J. A. R., Moreira, V. M., Goncalves, B. M. F., Leal, A. S., Jing, Y. 2012. Ursane-type pentacyclic triterpenoids as useful platforms to discover anticancer drugs. Nat. Prod. Rep. 29, 1463−1479.
- Suckling, C. 2012. From multiply active natural product to candidate drug? antibacterial (and other) minor groove binders for DNA. Future Med. Chem. 4, 971−989.
- Trott, O., Olson, A. J. 2010. Auto Dock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 31, 455−461.
- Van Patten, S., Al-Abed, Y. 2018. High Mobility Group Box-1 (HMGb1): Current Wisdom and Advancement as a Potential Drug Target. J. Med. Chem. 61, 5093−5107.
- Solvothermal synthesis of silver tungstate integrated with carbon nitrides matrix composites for highly sensitive electrochemical nitrofuran derivative sensing in biological samples. Anal. Chim. Acta. 2022;1192:339355.
- [Google Scholar]
- High mobility group box 1 (HMGB1): a pivotal regulator of hematopoietic malignancies. J. Hematol. Oncol.. 2020;13:91.
- [Google Scholar]
- Microbial transformation of pentacyclic triterpenes for anti-inflammatory agents on the HMGB1 stimulated RAW 264.7 cells by Streptomyces olivaceus CICC 23628.Bioorg. Med. Chem.. 2021;52:116-494.
- [Google Scholar]
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.104656.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







