Translate this page into:
Integrated HTS2, UPLC-MS/MS, and network pharmacology identifies Bruceae Fructus waste as potential source of flavonoids and quassinoids for inhibiting breast cancer
* Corresponding authors: E-mail addresses: dwang@cdutcm.edu.cn (D. Wang), 2528325529@qq.com (X. Bao)
-
Received: ,
Accepted: ,
Abstract
Shells are the main agro-industrial waste generated from industrial anti-cancer drugs manufacture using Bruceae Fructus. However, its potential for medicinal applications in cancer treatment has rarely been explored. In the present study, an integrated high throughput sequencing-based high throughput screening (HTS2), ultra performance liquid chromatography/tandem mass spectrometry (UPLC-MS/MS), and network pharmacology strategy was put forward to explore the possible utilization of waste in inhibiting breast cancer. The shell of Bruceae Fructus (BFS) inhibits proliferation and migration of breast cancer cells, while also inducing cell cycle arrest and apoptosis. The HTS2 assay revealed the possible molecular mechanisms activating the TP53 signaling pathway and apoptosis and inhibiting the cell cycle. Through network pharmacology analysis, PI3K/Akt signaling pathway was identified as another underlying mechanism. The HTS2 assay approved the downregulation of most genes in the PI3K/Akt pathway. UPLC-MS/MS identified flavones and quassinoids as the main ingredients in BFS. Molecular docking revealed the binding of these compounds to key targets including PI3K, AKT1, and PDK1. Our research shows that BFS may serve as a valuable source for discovering anti-cancer compounds and offers a novel approach for identifying potential drugs from presumed herb waste.
Keywords
Breast cancer
Bruceae Fructus waste
Flavonoids
HTS2
Quassinoids
1. Introduction
Brucea javanica (L.) Merr. belongs to the Brucea genus of Simaroubaceae family and it is traditionally used in countries and regions of southeast Asia. Bruceae Fructus (BF), ripe fruit of B. javanica, has been used clinically since Qing dynasty in China about 300 years ago. BF is black or brown, and oval with raised mesh wrinkles (Figure 1a). The shell of the BF (BFS) is hard, while the kernel of BF (BFK) is white and rich in oil (Figure 1a). The oil has been developed into oral liquids, injections, or capsules, for the adjuvant treatment of various cancers like lung cancer [1,2], brain metastasis of lung cancer, and gastrointestinal cancer [3,4]. There is also evidence proving that oil’s potential to treat breast cancer [5,6], leukemia [7], and colon cancer [8]. In Malaysia, BFK was also exploited for producing biodiesel because of its high oil content in the range of 35-42 wt% [9]. BFK is usually available for the extraction of oil; however, the BFS is often discarded as an agro-industrial residue due to traditional usage habits and poor understanding of phytochemical and pharmacological profiles. The weight of the BFS accounts for nearly half of the BF (Figure 1b). There still aren’t enough studies on the BFS although Liu et al. found brusatol in BFS [10]. Brusatol is a well-known NRF2 inhibitor [11] and exhibits anti-cancer [12-14] and anti-malaria effects [15,16]. Given that there is a growing demand for BF, it is worth exploring the chemical basis of BFS and potential pharmacological activities to improve the utilization of BF.
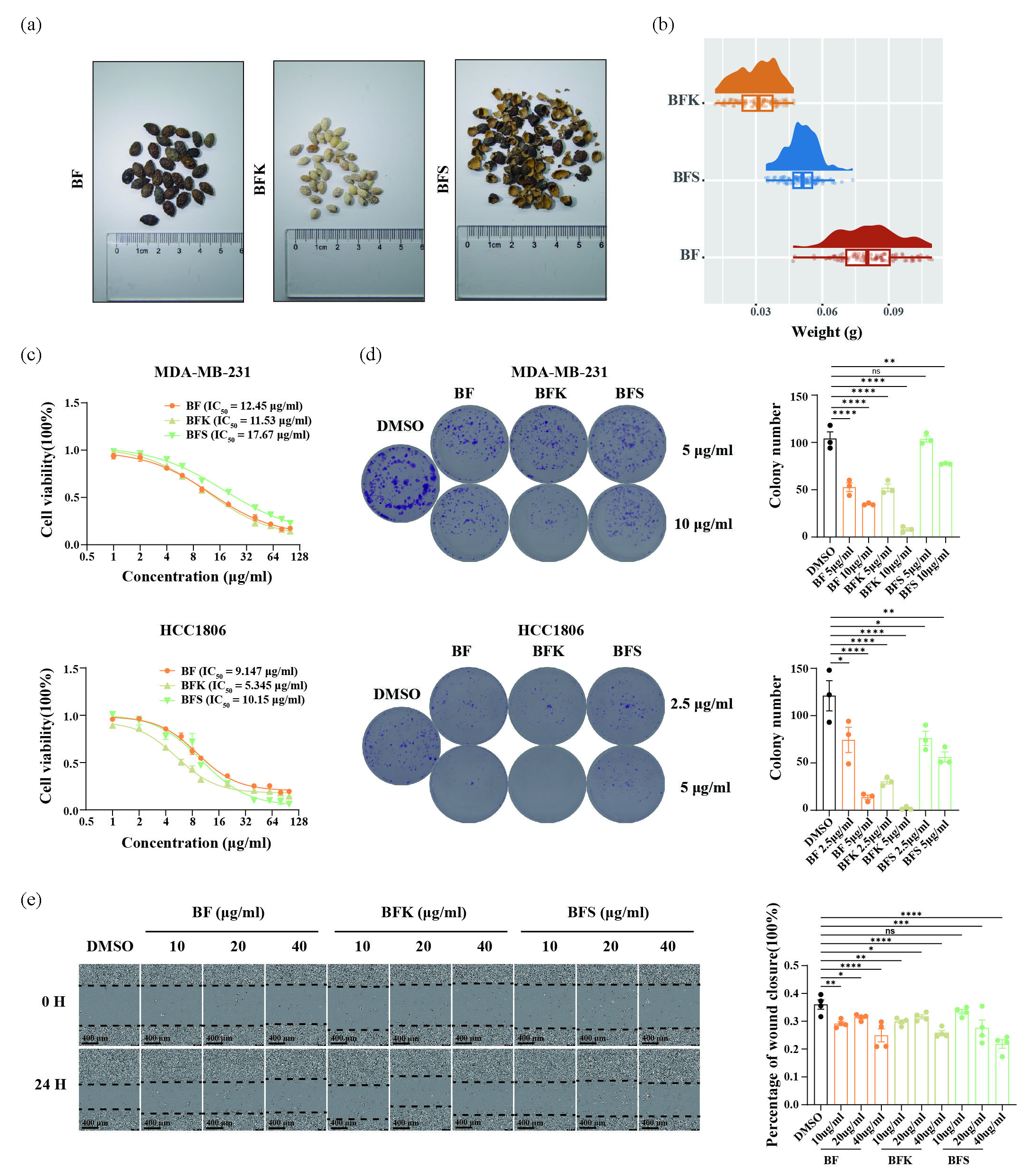
- BF, BFK, and BFS inhibited the proliferation and migration of breast cancer cells. (a) Different parts of Bruceae Fructus. (b) Weights analysis of BF, BFS, and BFK. (c) Ethanolic extract of BF, BFK, and BFS reduced MDA-MB-231 and HCC1806 cells viability, and (d) inhibited colony formation, and (e) migration. BF: Bruceae Fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus.
Breast cancer is the leading cause of cancer mortality among women in the vast majority of countries [17]. Although patients benefit from surgery, chemotherapy, targeted therapy, and new therapies, such as immunotherapy, these therapies could fail in the clinic or cause drug resistance. Therefore, it is urgent to discover effective drugs to treat breast cancer. The BF extracts or isolated compounds exert anti-cancer effects mainly through inhibiting cell proliferation, inducing apoptosis, inhibiting migration/invasion, inducing autophagy, and restraining angiogenesis [18]. Existing research showed that the ethanolic extract of BF inhibits breast cancer by restraining autophagy by activating the PI3K/Akt/mTOR pathway [19]. Su Jiyan et al. found that BF oil could also restrain autophagy by regulating gut microbiota-mediated amino acids [6]. However, whether BFS has effects on breast cancer and the chemical basis and molecular mechanisms of these effects are still unclear and need to be addressed.
Transcriptomics is a useful tool for exploring the potential pharmacological activities of medicine and its molecular mechanisms by analyzing multiple biological pathways or regulatory gene networks in cells after perturbation. Based on RNA-mediated oligonucleotide annealing, selection, and ligation with next-generation sequencing [20], high throughput sequencing-based high throughput screening (HTS2) technology permits quantitative profiling of the required sets of genes. Compared to microarray or RNA-seq, HTS2 costs less, detects accurately, and is more suitable for large-scale analysis [21]. Gene expression of immune cells perturbed by 578 herbal extracts was evaluated using HTS2. Simultaneously, the authors identified key targets of cytokine storm-related pathways and uncovered molecular mechanisms of their therapeutic effects on COVID-19 [22]. Therefore, this study will use HTS2 to explore the possible molecular mechanisms of BFS in inhibiting breast cancer.
Network pharmacology is widely used to study the chemical profiles and molecular mechanisms of herbal medicines treating diseases by predicting potential hub targets and pathways [23]. Potential molecular mechanisms were explored with experimental in vitro and in vivo validation. Bergaptol was identified as an active compound in Psoreleae Fructus (“Buguzhi” in Chinese) using ultra-performance liquid chromatography mass spectrometry/mass spectrometry (UPLC-MS/MS) Bergaptol promotes melanogenesis by regulating the p-P38 and phosphorylated extracellular regulated protein kinases (p-ERK) signaling pathways [24]. Jinhua Qinggan granules alleviated lung damage in mice models by inhibiting the expression and activation of inflammation-related proteins like p-ERK, p-STAT3, and NF-κB [25]. When integrated with HTS2, Yupingfeng San was found to treat hepatocellular carcinomas by inhibiting the mitogen-activated protein kinase (MAPK) signaling pathway [26]. Public databases provide limited phytochemical information on BF. It is necessary to profile the chemical basis of BF with a data-mining approach and then conduct network pharmacology research.
This study aims to explore the potential uses of BFS through HTS2, UPLC-MS/MS, and network pharmacology. Firstly, the influences of the BFS on breast cancer cells was studied through cell phenotype-related assays. Then, possible molecular mechanisms and key targets were explored using HTS2 and network pharmacology. Employing UPLC-MS/MS, the phytochemical composition of BFS was exhibited. Furthermore, molecular docking illustrated how these compounds interacted with key targets. Overall, this research provides insights into phytochemical and pharmacological profiles of BFS and suggests it to be a candidate source for discovery of compounds against breast cancer. Moreover, this study highlights the applicability of this research strategy in discovering utilization of other agro-industrial residues.
2. Materials and Methods
2.1. Preparation and extraction of BF, BFS, and BFK
BF (NO. 20210621) was purchased from the Bozhou Traditional Chinese Medicine market in Anhui province, China. The ripe fruits of B. javanica were identified by Prof. Guangzhi Wang from Chengdu University of Traditional Chinese Medicine. To obtain BFS and BFK, we hammered BF and separated the kernels from the mixture to obtain BFK as per Chinese Pharmacopoeia instructions. Meanwhile, the shells were also gathered as BFS. They were all stored in dry places, at room temperature, and away from lights. For extraction, BF, BFS, and BFK were smashed and extracted in a 95% ethanol solution using a Soxhlet extractor under heat for 8 hrs . After being evaporated, the remaining extracts were stored at -30°C.
2.2. Cell culture
MDA-MB-231 and HCC1806 cells were obtained from Zhejiang Meisen Cell Technology Co., Ltd. (Hangzhou, China). The cells were cultured in a 5% CO2 humidified incubator at 37°C in medium supplied with 10% fetal bovine serum (FBS) (ExCell) and 100 units/mL of streptomycin and penicillin (Cytiva). MDA-MB-231 cells were cultured in DMEM (Gibco), while HCC1806 cells were cultured in RPMI1640 medium (Gibco).
2.3. Cell viability assay
The cells were seeded in 96-well plates (3,000 cell/well for HCC1806 and 5,000 cell/well for MDA-MB-231) and cell viability was measured using the CCK-8 assay. After being cultured for 24 hrs, the cells were treated with several different concentrations of the ethanolic extract of BF, BFS, or BFK for another 48 hrs (3 biological replicates for each concentration). The CCK-8 regent (Bioground) was then added to each well and incubated for 1 hr at 37°C. The absorbance of each well at 450 nm was measured using Varioskan LUX multimode microplate reader (Thermo Fisher Scientific Inc., MA, USA).
2.4. Colony formation and wound healing assay
For colony formation assay, about 1,000 MDA-MB-231 or HCC1806 cells were seeded in 6-well plates. The ethanolic extracts of BF, BFS, or BFK (5, 10 μg/mL for MDA-MB-231; 2.5, 5 μg/mL for HCC1806) were added in the culture medium, and the cells were incubated for 2 weeks (3 biological replicates for each concentration). Then the cells were washed twice with phosphate buffered saline (PBS) and fixed with 4% PFA at room temperature. The fixed cells were stained with a crystal violet staining solution (Beyotime) and the colony numbers were counted via ImageJ 1.54f [27]. For the wound healing assay, about 10,000 MDA-MB-231 cells were seeded in 96-well plates. The wound was made using Incucyte Woundmaker Tool (Sartorius, Germany). After washing with PBS, medium free of FBS was added with or without extracts at concentrations of 10, 20, and 40 μg/mL (4 biological replicates for each concentration). Migration of cells was monitored and pictured in IncuCyte Live-Cell Analysis System (Sartorius, Germany). The size of closure area was analyzed in ImageJ.
2.5. Cell cycle
About a million of MDA-MB-231 or HCC1806 cells were seeded in 6 cm dishes. After being cultured for 24 hrs, the cells were incubated with ethanolic extracts of BF, BFS, or BFK (5, 10 μg/mL for MDA-MB-231; 10, 20 μg/mL for HCC1806) for another 24 hrs (3 biological replicates for each concentration). After digesting with 0.25% trypsin (Servicebio), the cells were collected and washed with cold PBS. Then the cells were fixed in 75% ethanolic solution at -30°C overnight. The fixed cells were washed with PBS, stained using a PI/RNase Staining Buffer (BD), and detected in a FACSVerse flow cytometer (BD, NJ, USA). ModFit LT 6.0 was adopted for analyzing the distributions of cell cycle.
2.6. Apoptosis
Annexin V-fluorescein isothiocyanate (FITC)/ Propidium iodide (PI) Apoptosis Detection Kit (Vazyme) was applied to detect apoptosis in cells. Briefly, a million of MDA-MB-231 or HCC1806 cells were seeded in 6 cm dishes for 24 hrs and then treated with ethanolic extracts of BF, BFS, or BFK (10, 20 μg/mL) for 24 hrs (3 biological replicates for each concentration). The cells were gathered after digesting with 0.25% trypsin without ethylenediaminetetraacetic acid (EDTA) (Servicebio), then washed twice with cold PBS, and resuspended with 100 μL binding buffer. The cells were stained with Annexin V-FITC and PI staining solution for 10 min at room temperature. Additional 400 μL binding buffer was added before detection in FACSVerse flow cytometer (BD, NJ, USA). The data was analyzed in FlowJo 10.6.2.
2.7. HTS2 assay
About 3,000 MDA-MB-231 cells were seeded in each well of 384-well plates. After incubation for 24 hrs, the cells were treated with ethanolic extracts of BF, BFS, or BFK at two concentrations (15 and 25 μg/mL, based on CCK-8 assay in 2.3) for another 24 hrs (3 biological replicates for each concentration). For choosing the concentrations of BF, BFK, and BFS in HTS2, we first conducted a CCK-8 assay to test their impact on the cell viability of MDA-MB-231. IC50 values of BF, BFK, and BFS were 12.45, 11.53, and 17.67 μg/mL separately (perturbed for 48 hrs). But the perturbation time in HTS2 was 24 hrs . At last, we chose 15 and 25 μg/mL for HTS2 assay. Then, the cells were lysed and transferred to the HTS2 platform for establishing cDNA library of transcripts as described [20]. The library was sequenced using NovaSeq 6000 (Illumina), and the mRNA level of the target gene was quantified as described [28]. The differentially expressed genes (DEGs) were calculated using DESeq2 [29] and the cutoff was set as |fold change| > 1.5 and P < 0.05. Additionally, the DEGs were imported to the ConnectivityMap (CMAP, https://clue.io/) [30] to query possible molecular mechanisms of functions.
2.8. Gene enrichment analysis
Gene ontology (GO) enrichment analysis, Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis, and Gene Set Enrichment Analysis (GSEA) were conducted in R 4.2.3 [31] using packages including clusterProfiler 4.6.2 [32], fgsea 1.24.0 [33] and tidyverse 2.0.0 [34].
2.9. UPLC-MS/MS analysis
Compounds were separated and identified using the Vanquish UPLC equipped with Q Exactive Orbitrap HRMS (Thermo Fisher Scientific Inc., MA, USA). Detailed chromatographic separation was conducted using AccucoreTM C18 column (3 mm × 100 mm, 2.6 μm, Thermo Fisher Scientific Inc., MA, USA) at a temperature of 30°C. The mobile phase consisted of eluent A (0.1% formic acid aqueous solution, v/v) and eluent B (0.1% formic acid acetonitrile, v/v), and gradient elution was carried out as follows: 0-40 mins, 5%-95% B, 40-45 mins, 95% B, post time 5 mins. The flow rate was 0.3 mL/min and injection volume was 3 μL. The detection was under both positive and negative iron mode with HESI as an ionization source. Other conditions as follows: scan range, 100-1,500 m/z; spray voltage, 3,500 V (positive iron mode), 3,000 V (negative iron mode); capillary temperature, 320°C; probe heater temperature, 350°C; sheath gas, 35 arb; aux gas, 10 arb; scan mode, full MS/dd-MS2; resolution, 70,000 (full MS), 17,500 (dd-MS2); stepped collision energies were 20, 40, 60 eV. Raw data was processed using Compound Discoverer 3.3 (Thermo Fisher Scientific Inc., MA, USA) with self-built analysis template for identifying unknown compounds. As for validation strategy for UPLC-MS/MS compound identification, we compared MS/MS data with that in databases such as mzCloud, mzVault, Chemspider, and local chemicals libraries. We set filter criteria of MS/MS data (mzCloud Best Match or mzVault Best Match > 80) to obtain possible compounds. To confirm these compounds more robustly, we next searched MS/MS data of each compound in databases like SCIFinder and Spectral Database for Organic Compounds (SDBS). Only compounds with correct molecular ion peak and reasonable fragment ion peak were thought to be the compounds in BF, BFK or BFS. And the list was verified next with reported data.
2.10. Gathering chemical information of BF and prediction of potential targets
To collect compounds isolated from BF, we searched in electronic databases including SciFinder, PubMed, Wiley, Springer, CNKI, and Google Scholar using keywords “Brucea javanica,” “Bruceae Fructus,” and “yadanzi.” The names of compounds were recorded, and the structures were drew using ChemDraw 19.0. Using SMILES of each compound, we predicted their targets in SwissTargetPrediction (http://www.swisstargetprediction.ch/) [35].
2.11. Collection of breast cancer targets and generation of network
We searched for targets of breast cancer in public websites such as OMIM (https://www.omim.org/, updated February 23rd, 2024) [36], TTD (https://db.idrblab.net/ttd/) [37], PHARMGKB (https://www.pharmgkb.org/) [38], and Drugbank (https://go.drugbank.com/) [39]. Then we picked overlapped genes with that in which 2.9 as BF’ targets breast cancer. The overlapped genes were also searched in STRING (https://cn.string-db.org/, version 12.0) [40] to get the hub genes, and were visualized in Cytoscape 3.10.0.
2.12. Molecular docking
The crystal structures of proteins in this study were obtained from the RCSB Protein Data Bank (RCSB PDB, https://www.rcsb.org/) [41]. And molecular docking was conducted using Schrodinger’s Maestro (Schrödinger Release, 2019–1). In brief, the protein was prepared using Protein Preparation module. Using LigPrep module, the possible 3D structures of compounds were prepared for docking. According to the known active binding sites, Glide was constructed for next molecular docking calculations and simulations [42].
2.13. Statistical analysis
Data in this study is presented as mean ± SEM. Comparation between two groups involved two-tailed unpaired Student’s t-test, and one way ANOVA tests followed by Tukey’s multiple comparisons were used for multiple groups. The significance level was set at *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
3. Results and Discussion
3.1. Comparing different pharmacological activities of BF, BFK, and BFS
3.1.1. Effects on cell viability, colony formation, and migration of TNBC cells
Triple-negative breast cancer (TNBC) represents one of the subtypes with the poorest prognosis and currently lacks efficient therapy. Therefore, we selected MDA-MB-231 and HCC1806, two typical TNBC cell lines, as models to investigate pharmacological activities. CCK-8 assays were conducted in these two cell lines to assess the effects on cell-viability. BFK was the strongest inhibitor of cell viability in both cell lines, and the IC50 value was 11.53 μg/mL in MDA-MB-231 and 5.345 μg/mL in HCC1806. IC50 values of BF and BFS were 12.45 μg/mL and 17.67 μg/mL, respectively in MDA-MB-231. IC50 values were 9.147 μg/mL and 10.15 μg/mL in HCC1806 (Figure 1c). The results of colony formation assay were consistent with the inhibition of cell viability (Figure 1d). In wound healing assays, BF, BFK, and BFS inhibited the migration of MDA-MB-231 (Figure 1e). BFS inhibits growth, proliferation, and migration of TNBC cells.
3.1.2. Effects on cell cycle and apoptosis of TNBC cells
Considering BF, BFK, and BFS could inhibit the growth, proliferation, and migration of TNBC cells at relatively low concentrations, we explored whether it was relevant to cell cycle and apoptosis. With analysis of DNA content, we found that BF arrests the cell cycle of MDA-MB-231 cells at the G0/G1 phase, and HCC1806 at the S stage (Figure 2a). The activity of inducing apoptosis was also observed in MDA-MB-231 and HCC1806 (Figure 2b). Both BFK and BFS exhibited similar effects of arresting the cell cycle and inducing apoptosis.
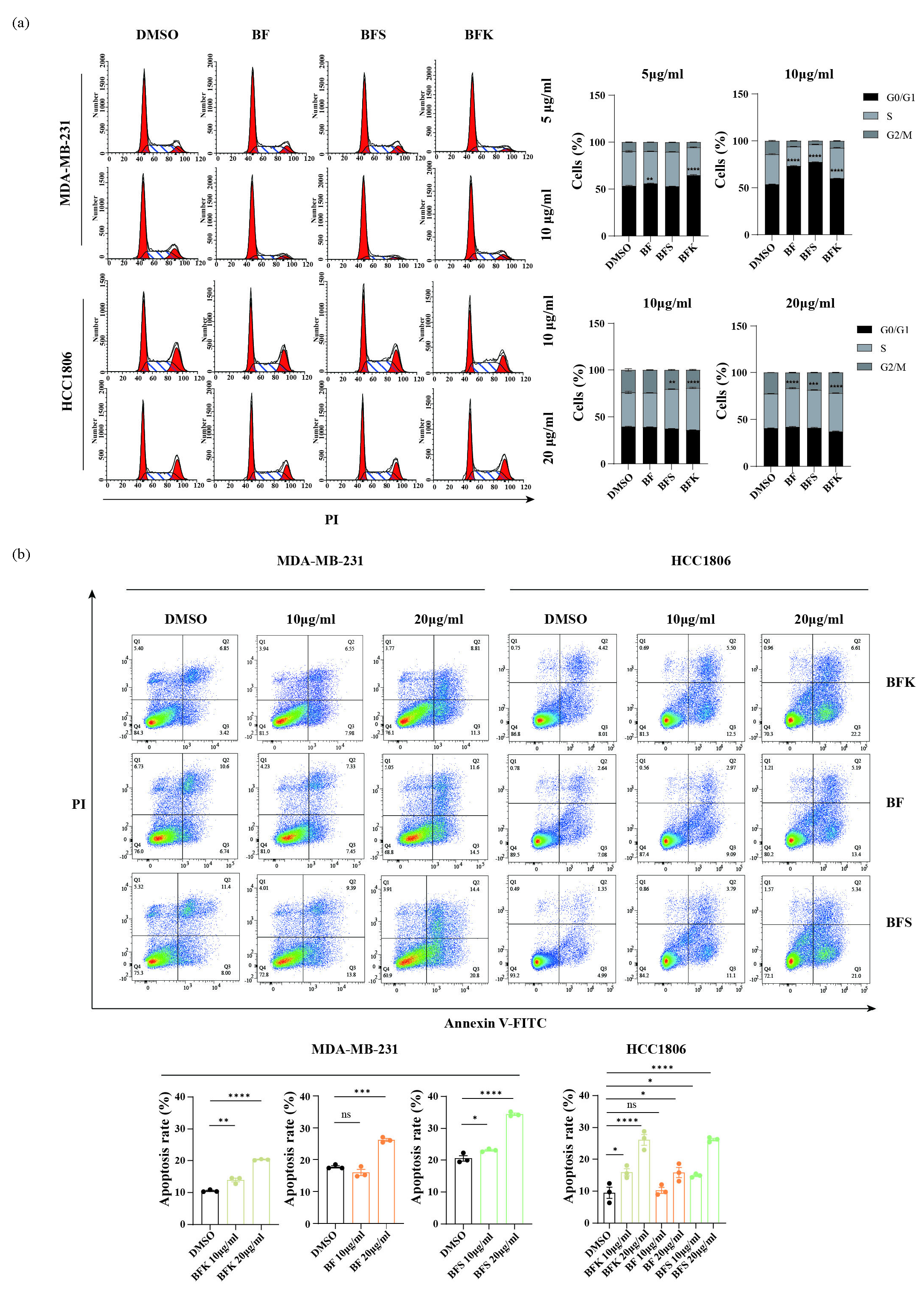
- The ethanolic extracts of BFK, BF, and BFS induce cell cycle arrest and apoptosis of MDA-MB-231 and HCC1806 cells. (a) DNA content analysis in MDA-MB-231 and HCC1806 using flow cytometry. (b) Apoptosis analysis in MDA-MB-231 and HCC1806 using flow cytometry. BF: Bruceae Fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus. DMSO: Dimethyl sulfoxide.
3.2. HTS2 analysis
3.2.1. BFS regulates similar number of DEGs in MDA-MB-231 with BF or BFK
Using dimethyl sulfoxide (DMSO) (negative control) and JQ1 (positive control), we treated MDA-MB-231 cells with two concentrations of ethanolic extracts of BF, BFK, and BFS. The correlation between each pair of samples in the DMSO or JQ1 group was found to be greater than 0.94, indicating excellent repeatability within the experimental groups (Figure A1a, b). Samples in DMSO and JQ1 group were separated into two different clusters using uniform manifold approximation and projection (UMAP) analysis (Figure A1c), suggesting the reliability of HTS2 assay.
Next, we visualized the gene regulations in MDA-MB-231 cells treated with different extracts using volcano plots (Figure 3a). A Venn plot revealed a total of 240 DEGs identified in the BFK group, 182 in the BF group, and 181 in the BFS group at a concentration of 15 μg/mL. At a concentration of 25 μg/mL, 470 DEGs were identified in the BFK group, 446 DEGs in the BF group, and 437 DEGs in the BFS group (Figure. 3b). With respect to gene perturbation, BFS exhibited similar pattern of gene regulation with BF or BFK in MDA-MB-231 cells.
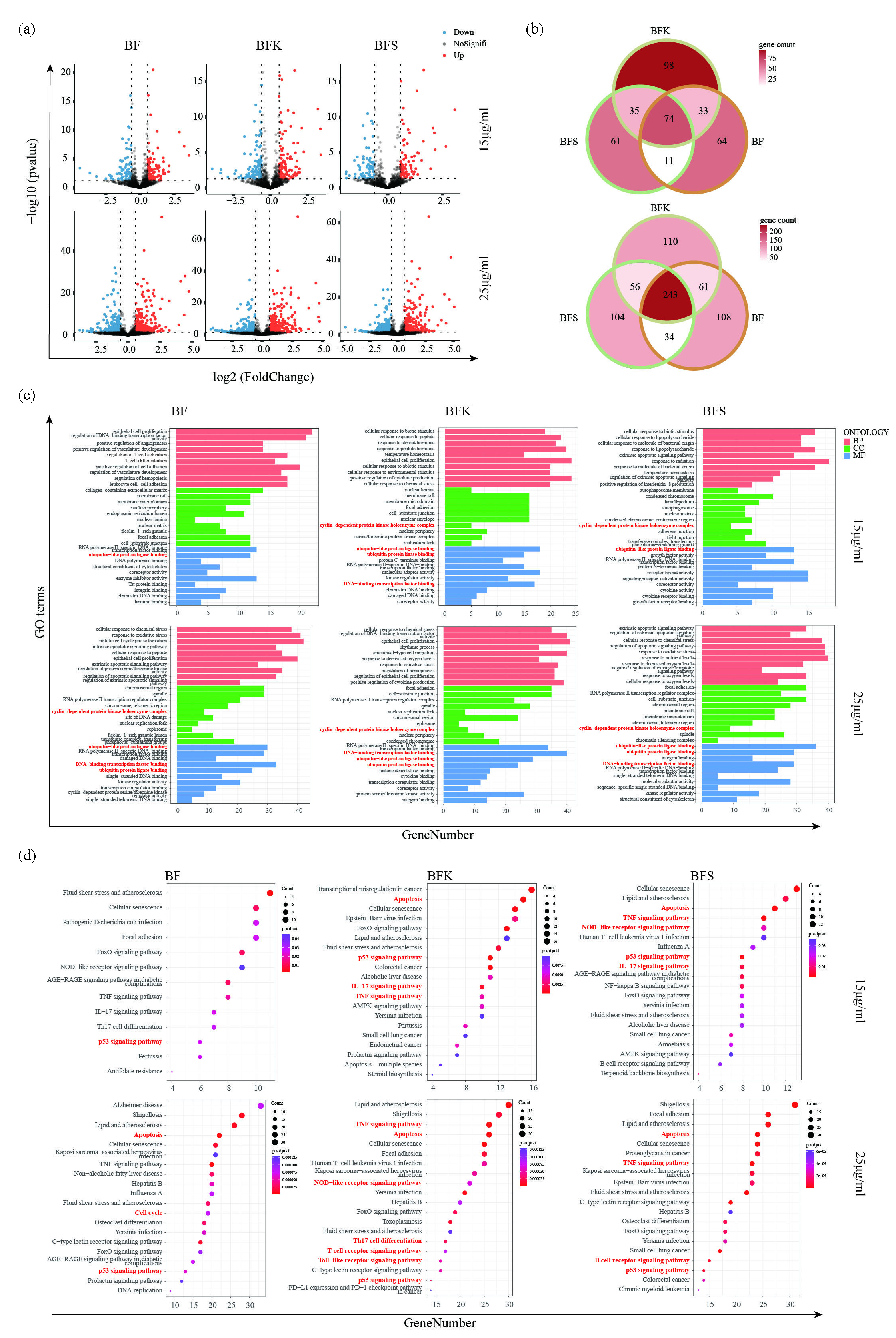
- HTS2 technology illustrated mechanisms of BF, BFK, and BFS treating breast cancer. (a) Volcano plot profiling DEGs and (b) Venn diagrams of DEGs in MDA-MB-231 treated with ethanolic extracts of BF, BFK, and BFS. (c) GO enrichment analysis and (d) KEGG pathway enrichment analysis of DEGs in MDA-MB-231 treated with ethanolic extracts of BF, BFK, and BFS. BF: Bruceae Fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus, GO: Gene ontology, KEGG : Kyoto encyclopedia of genes and genomes, HTS2: High throughput screening, DEGs: Differentially expressed genes.
3.2.2. GO and KEGG enrichment analysis
HTS2 results showed that “cyclin−dependent protein kinase holoenzyme complex” was enriched in the GO enrichment analysis, while “p53 signaling pathway”, “Apoptosis”, and “Cell cycle” were enriched in KEGG enrichment analysis. We also found that cell cycle related pathways were inhibited according to GSEA (Figure A2). Besides, other GO terms like “RNA polymerase II transcription regulator complex”, “DNA−binding transcription factor binding”, “ubiquitin−like protein ligase binding”, and “ubiquitin protein ligase binding” were also enriched significantly. It inspired us that both BFK and BFS probably exerted anti-cancer effects via inhibiting gene transcription or promoting ubiquitin-dependent protein degradation (Figure 3c).
In addition, some inflammation related KEGG terms such as “NOD−like receptor signaling pathway,” “TNF signaling pathway,” and “IL−17 signaling pathway” were enriched in both BFK and BFS group. Differently, “T cell receptor signaling pathway” and “Toll−like receptor signaling pathway” were enriched in BFK group, meanwhile “B cell receptor signaling pathway” was enriched in BFS group (Figure 3d). That means BF might have influence on the immune microenvironment in breast cancer and recruit anti-cancer immune cells. In summary, pathways predicted in network pharmacology were validated through HTS2 analysis. Additionally, other intriguing pathways were enriched, shedding light on how BF could potentially treat breast cancer.
3.2.3 Results of GSEA
Cell cycle and apoptosis were enriched in GO and KEGG enrichment analysis. Previous experiments have validated the ability of BF, BFK, and BFS to arrest the cell cycle and induce apoptosis in TNBC cells. We performed GSEA using HTS2 data to illustrate how they influence these pathways at molecular level. As a result, most genes in “KEGG CELL CYCLE” were downregulated in MDA-MB-231 treated with extracts of BF, BFK, and BFS. Some genes encoding cell cycle checkpoint proteins like CDC6, CDK1, CCNB1, and CCNB2 were downregulated. In Figure 4 SKP2, a regulator of p21 and p27 [43], was also downregulated (Figure 4a). Additionally, most genes in “HALLMARK P53 PATHWAY” were upregulated, indicating activation of the pathway after treatment (Figure 4c). The activation of p53 might result in the cell-cycle arrest of TNBC cells [44,45]. Genes in “HALLMARK APOPTOSIS” were upregulated after treatment with extracts of BF, BFK, and BFS. For example, in the BFK group, CASP7, an activator of apoptosis [46], was upregulated. The increasing expression of TXNIP, DDIT3, and GADD45A suggested the accumulation of reactive oxygen species (ROS) and DNA damage [47-49]. The upregulations of these genes were also observed in the BF and BFS groups (Figure 4b).
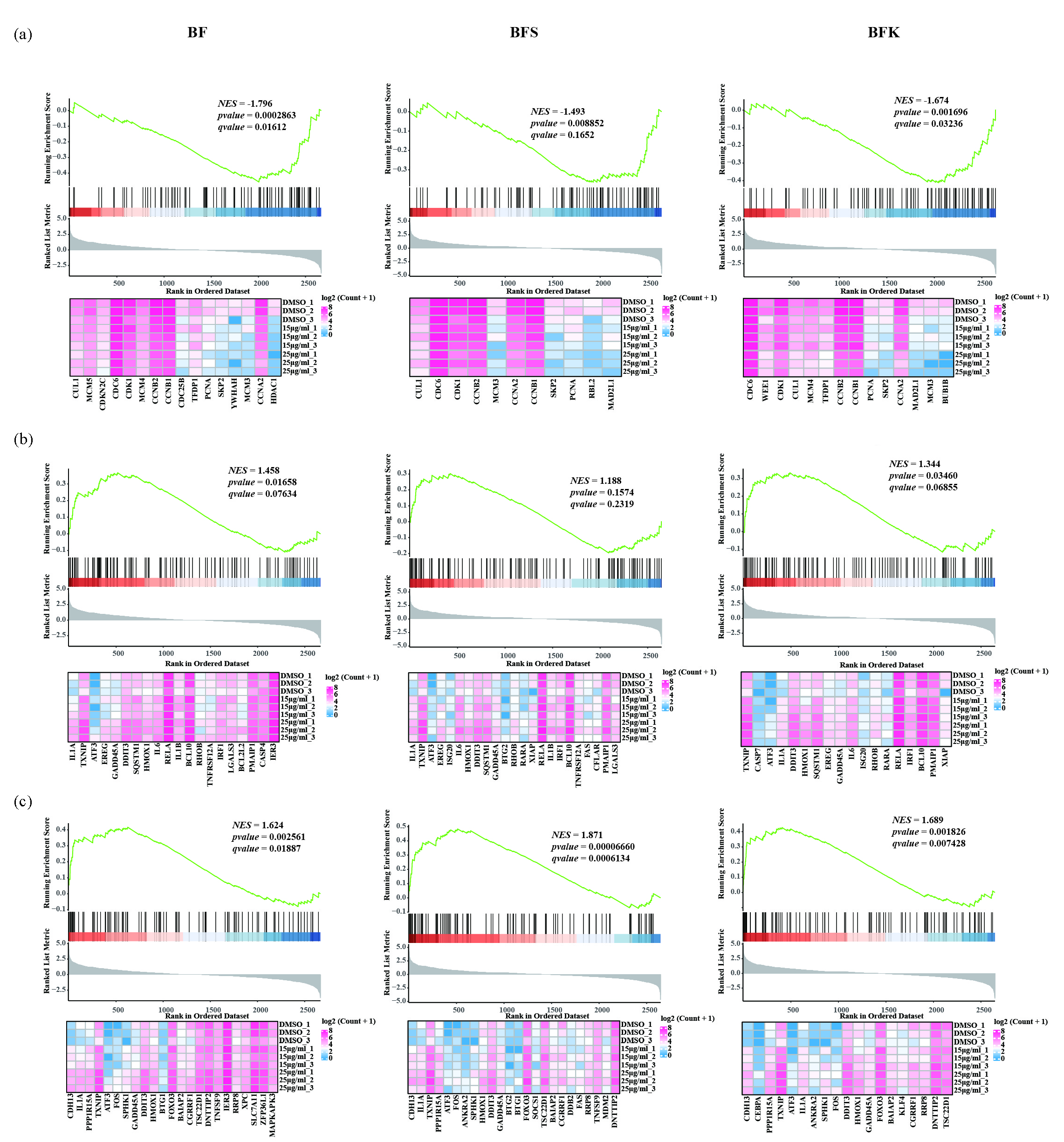
- BF, BFS, and BFK inhibited the pathway of (a) “KEGG CELL CYCLE”, and activated pathways of (b) “HALLMARK APOPTOSIS” and (c) “HALLMARK P53 PATHWAY”. The lower heatmap shows an expression of the core gene in the pathway in the form of log2 (count + 1). BF: Bruceae Fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus, NES: Normalized enrichment score, KEGG: Kyoto encyclopedia of genes and genomes.
3.3. The chemical composition of BF, BFS, or BFK varies
BFK is traditionally used in clinical practice, unlike BF or BFS. Therefore, we compared the differences in chemical composition among them using UPLC-MS/MS (Figure 5a, b). We identified a total of 60 ingredients in BF, with 31 discovered in negative mode and 29 in positive mode. Quassinoids such as bruceine H were detected only in BFK, while yadanziolide B was only found in BFS. Although peak areas were smaller in BFS than in BFK, compounds like bruceolide, brusatol, bruceine A, and bruceine D were present in both parts of BF. Additionally, organic acids and flavones such as caffeic acid, isovanillic acid, pimelic acid, ferulic acid, quercetin, quercetin-3β-D-glucoside, kaempferol, and luteolin were more readily found in BFS. The MS/MS data and peak areas of each compound have been presented in Table 1. These findings suggest that BFS contains lower content of quassinoids, but more flavones compared to BFK.
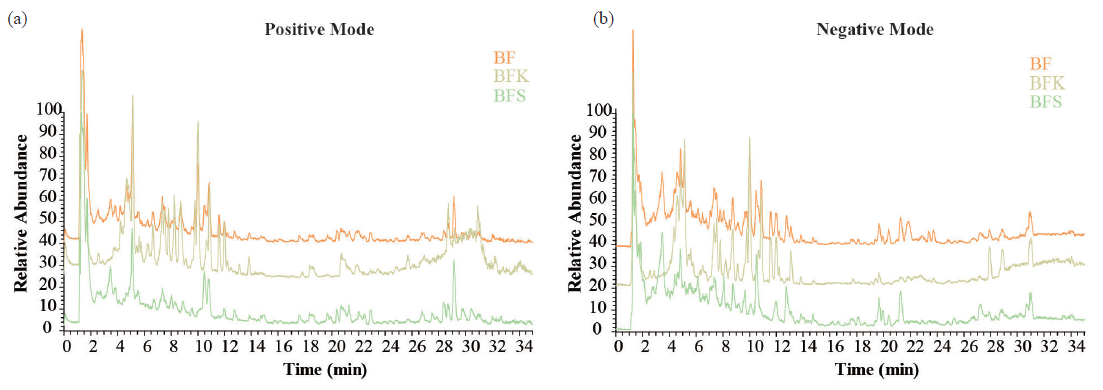
- UPLC-MS/MS chromatograms of ethanolic extracts of BF, BFS, and BFK in (a) positive and (b) negative modes. BF: Bruceae Fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus, UPLC: Ultra performance liquid chromatography, MS: Mass spectrometry.
| Number | RT (min) | Name | Formular | Adduct | MS | Error (ppm) | MS/MS | Area. BF | Area. BFK | Area. BFS |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1.211 | DL-Arginine | C6H14N4O2 | [M+H] +1 | 175.11891 | 0.06 | 158.09232, 130.09749, 116.07081, 70.06573, 60.05629 | 2.46E+09 | 2.87E+08 | |
| 2 | 1.245 | L-Iditol | C6H14O6 | [M-H]-1 | 181.07140 | -1.40 | 163.06062, 149.04497, 115.03910, 101.02353, 89.02348, 71.01287, 59.01282 | 4.52E+08 | 9.01E+08 | 3.31E+08 |
| 3 | 1.288 | Trigonelline | C7H7NO2 | [M+H] +1 | 138.05482 | -0.47 | 110.06028, 94.06552, 67.05473 | 1.13E+10 | 1.87E+09 | 6.51E+09 |
| 4 | 1.327 | Adenine | C5H5N5 | [M+H] +1 | 136.06172 | -0.23 | 119.03545, 94.04031 | 3.96E+08 | 5.53E+08 | 3.25E+08 |
| 5 | 1.405 | D- (+) -Malic acid | C4H6O5 | [M-H]-1 | 133.01353 | -4.87 | 115.00288, 89.02347, 72.99214, 71.01286 | 4.55E+09 | 6.14E+08 | 4.57E+09 |
| 6 | 1.426 | Gluconic acid | C6H12O7 | [M-H]-1 | 195.05078 | -1.49 | 177.03963, 159.02962, 129.01860, 99.00793, 75.00780 | 1.98E+09 | 2.16E+09 | 1.50E+09 |
| 7 | 1.474 | 3-Hydroxy-3-methylglutaric acid | C6H10O5 | [M-H]-1 | 161.04495 | -3.06 | 143.03423, 101.02352, 99.04425, 57.03354 | 8.70E+08 | 7.01E+07 | 1.55E+09 |
| 8 | 1.717 | Nicotinic acid | C6H5NO2 | [M+H] +1 | 124.03932 | 0.98 | 96.04474, 80.05000 | 1.87E+09 | 3.63E+08 | 2.27E+09 |
| 9 | 1.725 | D- (+) -Pyroglutamic Acid | C5H7NO3 | [M+H] +1 | 130.04988 | 3.62 | 102.05543, 84.04491, 56.05017 | 2.19E+09 | 8.91E+08 | 1.73E+09 |
| 10 | 1.728 | Uridine | C9H12N2O6 | [M-H]-1 | 243.06335 | 1.40 | 200.05627, 152.03452, 110.02396 | 1.03E+08 | 4.48E+07 | 1.33E+08 |
| 11 | 1.730 | 3-Hydroxypicolinic acid | C6H5NO3 | [M+H] +1 | 140.03407 | -1.25 | 122.02402, 112.03951, 94.02926 | 1.64E+09 | 2.46E+09 | |
| 12 | 1.732 | Adenosine | C10H13N5O4 | [M+H] +1 | 268.10379 | -0.73 | 136.06169, 119.03522 | 1.49E+09 | 4.51E+08 | 1.99E+09 |
| 13 | 2.093 | Isovanillic acid | C8H8O4 | [M-H]-1 | 167.03447 | -2.21 | 152.01089, 123.04432, 108.02060 | 1.08E+08 | 1.55E+08 | |
| 14 | 2.367 | Glutaric acid | C5H8O4 | [M-H]-1 | 131.03424 | -4.66 | 113.02362, 87.04421, 69.03358 | 2.21E+08 | 6.56E+07 | 2.66E+08 |
| 15 | 2.999 | Methylsuccinic acid | C5H8O4 | [M-H]-1 | 131.03424 | -4.77 | 113.02357, 87.04421, 69.03358 | 3.33E+08 | 4.68E+07 | 5.04E+08 |
| 16 | 3.459 | 2-Hydroxyquinoline | C9H7NO | [M+H] +1 | 146.05992 | 0.21 | 128.04951, 118.06529, 91.05470 | 1.01E+10 | 3.38E+08 | 1.66E+10 |
| 17 | 3.560 | Bruceine H | C20H26O10 | [M+H] +1 | 427.15765 | 0.46 | 409.14920, 391.13785, 373.12762, 347.14896, 299.12677, 269.11682 | 8.66E+07 | 1.21E+08 | |
| 18 | 3.567 | Caffeic acid | C9H8O4 | [M+H] +1 | 181.04924 | -1.68 | 163.03882, 135.04404, 117.03368, 89.03902 | 8.74E+07 | 1.32E+08 | |
| 19 | 3.852 | trans-3-Indoleacrylic acid | C11H9NO2 | [M+H] +1 | 188.07056 | 0.10 | 170.06004, 146.05998, 144.08076, 118.06532 | 3.38E+09 | 4.58E+08 | 2.33E+09 |
| 20 | 3.892 | 2-Anisic acid | C8H8O3 | [M-H]-1 | 151.03944 | -4.06 | 136.01585, 123.04435, 107.04939, 93.03366 | 8.90E+07 | 1.55E+08 | |
| 21 | 4.041 | N-Acetyl-L-tyrosine | C11H13NO4 | [M-H]-1 | 222.07703 | 0.40 | 180.06618, 163.03940, 119.04948, 107.04940, 58.02881 | 1.83E+08 | 1.11E+07 | 2.93E+08 |
| 22 | 4.824 | Bruceine D | C20H26O9 | [M+H] +1 | 411.16443 | -0.40 | 393.15375, 375.14380, 345.13300, 299.12805, 271.13257 | 8.04E+09 | 9.04E+09 | 8.88E+07 |
| 23 | 5.125 | Indole-3-acrylic acid | C11H9NO2 | [M+H] +1 | 188.07057 | -0.76 | 170.05998, 146.05997, 142.06509, 118.06534 | 2.20E+08 | 3.34E+08 | |
| 24 | 5.201 | Pimelic acid | C7H12O4 | [M-H]-1 | 159.06572 | -2.98 | 141.05515, 115.07562, 97.06499, 95.04934 | 9.86E+07 | 1.63E+08 | |
| 25 | 5.553 | Norharman | C11H8N2 | [M+H] +1 | 169.07587 | 0.05 | 115.05437, 65.03916 | 2.20E+09 | 7.11E+07 | 2.91E+09 |
| 26 | 6.128 | 1,3,5-trihydroxy-4-{[(2E)-3-(3-hydroxy-4-methoxyphenyl) prop-2-enoyl] oxy} cyclohexane-1-carboxylic acid | C17H20O9 | [M-H]-1 | 367.10336 | 1.20 | 193.05037, 173.04506, 134.03662, 93.03368 | 1.65E+09 | 2.30E+07 | 3.63E+09 |
| 27 | 6.128 | Ferulic acid | C10H10O4 | [M+H] +1 | 195.06670 | 0.45 | 177.05452, 145.02834, 117.03365, 89.06017 | 3.32E+07 | 6.88E+07 | |
| 28 | 6.447 | 2-(Acetylamino)hexanoic acid | C8H15NO3 | [M-H]-1 | 172.09741 | -1.98 | 130.08659, 128.10716, 82.06524 | 1.70E+08 | 1.03E+08 | 1.02E+08 |
| 29 | 6.713 | Vanillin | C8H8O3 | [M+H] +1 | 153.05455 | -0.06 | 125.05978, 111.04430, 93.03391, 65.03922 | 1.65E+09 | 9.81E+07 | 2.27E+09 |
| 30 | 7.011 | Suberic acid | C8H14O4 | [M-H]-1 | 173.08145 | -1.89 | 155.07103, 129.09146, 111.08070, 83.04929 | 3.41E+08 | 8.45E+07 | 5.12E+08 |
| 31 | 7.072 | Ellagic acid | C14H6O8 | [M-H]-1 | 300.99927 | 1.32 | 283.99643, 229.01411, 185.02385 | 3.80E+09 | 3.69E+07 | 4.51E+09 |
| 32 | 7.206 | Quercetin-3β-D-glucoside | C21H20O12 | [M-H]-1 | 463.08875 | 1.06 | 300.02786, 271.02521, 255.03014 | 5.06E+08 | 9.91E+08 | |
| 33 | 7.207 | Quercetin | C15H10O7 | [M+H] +1 | 303.04947 | -1.05 | 229.04929, 201.05443, 153.01810, 137.02332 | 1.50E+08 | 3.09E+08 | |
| 34 | 7.306 | 5,7-Dihydroxy-2-(4-hydroxyphenyl)-4-oxo-4H-chromen-3-yl 6-O-(6-deoxyhexopyranosyl) hexopyranoside | C27H30O15 | [M-H]-1 | 593.15204 | 0.51 | 447.09653, 285.04086 | 6.52E+08 | 9.74E+08 | |
| 35 | 7.383 | Kaempferol | C15H10O6 | [M+H] +1 | 287.05463 | -2.49 | 269.04416, 153.01816, 135.04407 | 3.35E+08 | 6.69E+08 | |
| 36 | 7.389 | Indole-3-acetic acid | C10H9NO2 | [M-H]-1 | 174.05559 | -2.02 | 146.06049, 130.06551 | 3.69E+08 | 1.76E+07 | 6.86E+08 |
| 37 | 7.780 | Bruceolide | C21H26O10 | [M+H] +1 | 439.15204 | 0.03 | 421.14868, 403.13785, 375.14288, 343.11716, 299.12738, 201.09094 | 7.24E+07 | 4.83E+07 | 1.28E+07 |
| 38 | 7.905 | 2-(acetylamino)-3-(1H-indol-3-yl) propanoic acid | C13H14N2O3 | [M-H]-1 | 245.09369 | 0.80 | 203.08240, 116.04974, 74.02378 | 6.65E+07 | 2.88E+06 | 1.02E+08 |
| 39 | 8.063 | 9-(2,3-dihydroxypropoxy)-9-oxononanoic acid | C12H22O6 | [M-H]-1 | 261.13461 | 1.16 | 187.09724, 125.09644 | 1.02E+08 | 2.24E+07 | 1.44E+08 |
| 40 | 8.455 | 4-Indolecarbaldehyde | C9H7NO | [M+H] +1 | 146.05998 | -1.24 | 118.06531, 91.05470 | 8.41E+08 | 4.66E+07 | 1.55E+09 |
| 41 | 8.727 | brusatol | C26H32O11 | [M+H] +1 | 521.20239 | -2.05 | 485.35779, 421.14859, 403.13889, 299.12811 | 3.67E+09 | 3.80E+09 | 6.92E+07 |
| 42 | 9.469 | Salicylic acid | C7H6O3 | [M-H]-1 | 137.02374 | -4.17 | 93.03368, 65.03853 | 5.96E+08 | 9.99E+07 | 9.78E+08 |
| 43 | 9.558 | 4-Nitrophenol | C6H5NO3 | [M-H]-1 | 138.01889 | -4.78 | 108.02083 | 2.58E+08 | 2.52E+07 | 3.90E+08 |
| 44 | 10.090 | ethyl 9H-beta-carboline-3-carboxylate | C14H12N2O2 | [M+H] +1 | 241.09706 | -1.46 | 213.06570, 185.07089, 167.06032 | 8.64E+07 | 3.12E+06 | 1.72E+08 |
| 45 | 10.493 | Luteolin | C15H10O6 | [M-H]-1 | 285.04071 | -0.58 | 241.05075, 133.02873 | 8.39E+09 | 3.73E+07 | 1.34E+10 |
| 46 | 10.642 | yadanziolide B | C20H26O11 | [M+H] +1 | 443.15417 | -1.11 | 425.14172, 407.13257, 389.12283, 361.12787, 285.11172, 243.10147, 213.09084, 201.09079 | 1.11E+08 | 2.76E+07 | 7.52E+07 |
| 47 | 10.868 | Bruceine A | C26H34O11 | [M+H] +1 | 523.21625 | -1.71 | 421.14682, 403.13736, 375.14297, 201.09084, 85.06529 | 2.79E+09 | 2.64E+09 | 6.79E+07 |
| 48 | 11.574 | bruceantinol | C30H38O13 | [M+NH4] +1 | 624.26392 | -2.54 | 547.21686, 439.15924, 201.09091, 127.07540, 109.06505, 81.07040 | 1.47E+08 | ||
| 49 | 13.543 | Dodecanedioic acid | C12H22O4 | [M-H]-1 | 229.14450 | 0.65 | 211.13377, 167.14365 | 1.06E+08 | 1.09E+07 | 2.49E+08 |
| 50 | 13.927 | Hispidulin | C16H12O6 | [M-H]-1 | 299.05652 | 0.80 | 284.03287, 256.03732 | 9.39E+06 | 2.67E+07 | |
| 51 | 16.213 | 6-Gingerol | C17H26O4 | [M-H]-1 | 293.17651 | 1.71 | 236.10542, 221.15460 | 4.94E+07 | 8.76E+07 | 5.05E+07 |
| 52 | 17.127 | (+/-)12(13)-DiHOME | C18H34O4 | [M-H]-1 | 313.23877 | 1.25 | 295.22827, 183.13861, 129.09137, 99.08064 | 1.78E+08 | 8.99E+07 | 2.52E+08 |
| 53 | 18.091 | (+/-)9,10-dihydroxy-12Z-octadecenoic acid | C18H34O4 | [M-H]-1 | 313.23856 | 0.99 | 295.22800, 201.11302 | 8.71E+08 | 1.96E+08 | 1.73E+09 |
| 54 | 19.393 | Hexadecanedioic acid | C16H30O4 | [M-H]-1 | 285.20749 | 1.15 | 267.19687, 223.20665 | 7.28E+08 | 5.05E+08 | 7.24E+08 |
| 55 | 19.736 | Progesterone | C21H30O2 | [M+H] +1 | 315.23175 | -1.40 | 297.22189, 109.06511, 97.06524 | 2.80E+08 | 6.11E+08 | |
| 56 | 23.464 | 2-Hydroxymyristic acid | C14H28O3 | [M-H]-1 | 243.19670 | 0.84 | 225.18541, 197.19078 | 4.69E+07 | 1.71E+07 | |
| 57 | 26.638 | 1-Linoleoyl glycerol | C21H38O4 | [M+H] +1 | 355.28394 | -1.81 | 337.27319, 263.23676, 245.22621, | 1.10E+08 | 7.19E+07 | 1.14E+08 |
| 58 | 27.277 | 16-Hydroxyhexadecanoic acid | C16H32O3 | [M-H]-1 | 271.22827 | 1.34 | 253.21617, 225.22237 | 3.98E+09 | 4.87E+08 | 7.51E+09 |
| 59 | 28.741 | Oleamide | C18H35NO | [M+H] +1 | 282.27890 | -1.80 | 265.25214, 247.24182 | 1.04E+09 | 2.38E+09 | |
| 60 | 30.940 | Erucamide | C22H43NO | [M+H] +1 | 338.34125 | -1.05 | 321.31485, 303.30447 | 4.07E+08 | 4.25E+08 |
BF: Bruceae fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus, RT: Retention time, ppm: Parts per million, MS: Mass spectrometry
3.4. Results of network pharmacology
3.4.1. Compounds collection and their targets prediction
To date, approximately 167 chemical ingredients have been identified in BF, encompassing 97 quassinoids, 14 terpenoids, 16 flavones, 13 nitrogen-containing compounds, and others. Subsequently, we used SwissTargetPrediction to predict the potential targets of these compounds, resulting in a total of 807 targets. Detailed information regarding these chemicals and their predicted targets has been provided in Table A1.
3.4.2. Collection of breast cancer targets and PPI network analysis
We collected therapeutic targets of breast cancer from OMIM, TTD, PHARMGKB, and Drugbank (Table A2). Ultimately, a total of 114 targets overlapped with those identified previously (in section 3.4.1) and were considered therapeutic targets (Figure 6a). Subsequently, we constructed a compound-target network (Figure 6b), which revealed that 62 quassinoids exerted significant anti-cancer effects. Analysis of the protein-protein interaction (PPI) network (Figure 6c) and topological analysis identified hub genes including TP53, SRC, PIK3CA, PIK3CB, PIK3R1, ESR1, AKT1, PIK3CD, CTNNB1, and CYP3A4 (Table 2). Given the higher abundance of quassinoids and lower presence of flavones in BFK compared to BFS, we compared the different targets of these compounds. PPI network analysis (Figure 6d, e) revealed that quassinoids influenced hub genes such as EGFR, SRC, AKT1, PIK3CA, and PIK3CB, while flavones acted on TP53, EGFR, SRC, AKT1, and ESR1. Together, BF may exert anti-breast cancer effects through modulation of these hub genes.
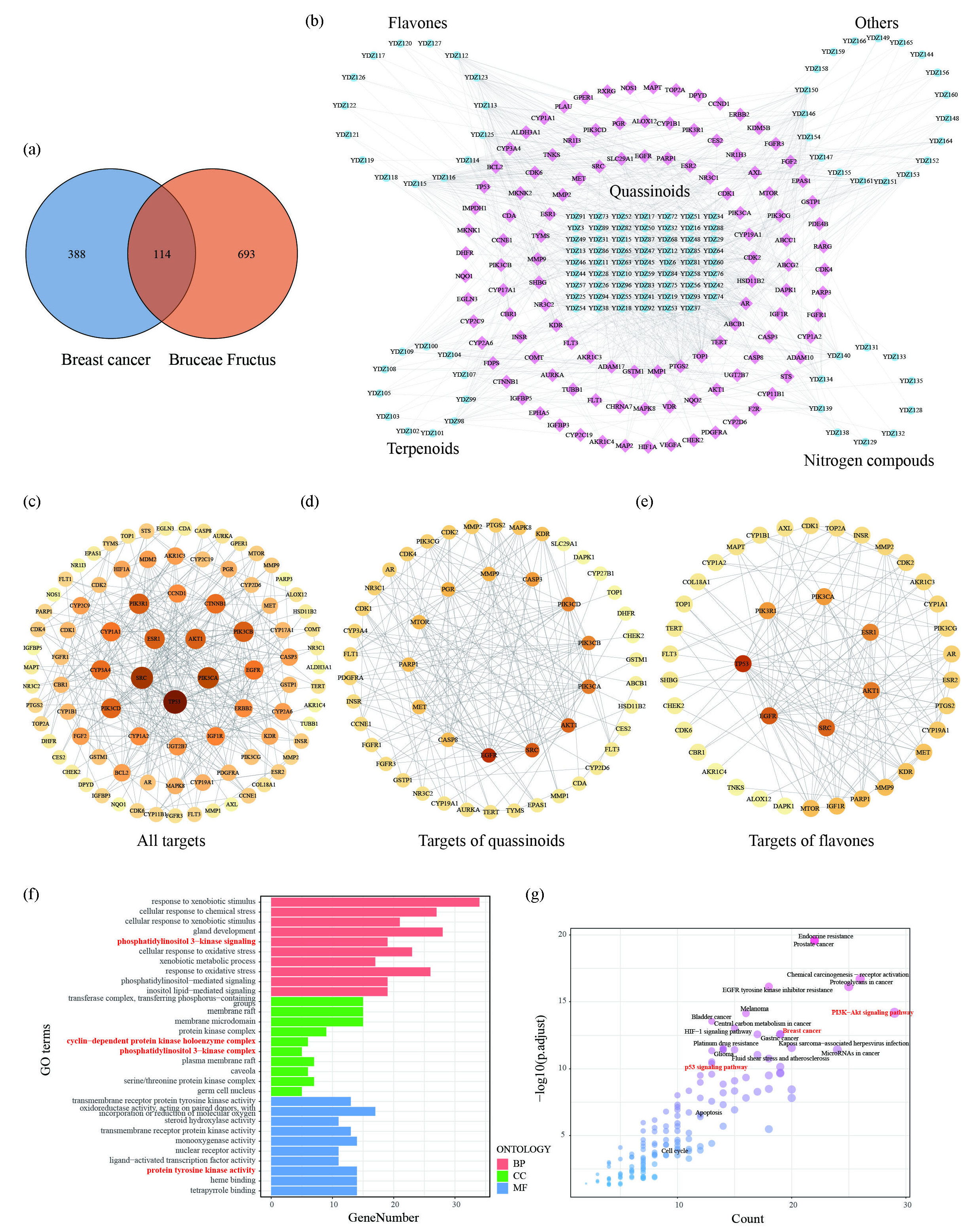
- Network pharmacology illustrating mechanisms of BF treating breast cancer. (a) Venn diagram of targets of BF and breast cancer. (b) Compound-target network was built using 114 overlapped genes. The round nodes are chemicals in BF while the square ones are therapeutical targets of breast cancer. (c) PPI network of 114 overlapped targets, (d) targets of quassinoids, and (e) targets of flavones. The size or color shade of the node is positively associated with the degree value. (f) GO enrichment analysis of 114 targets of BF treating breast cancer. (g) KEGG pathway enrichment analysis of 114 targets of BF treating breast cancer. BP: Biological process; CC: Cellular component; MF: Molecular function, GO: Gene ontology, KEGG : Kyoto encyclopedia of genes and genomes.
| GENE | Degree centrality | GENE | Betweenness centrality | GENE | Closeness centrality |
|---|---|---|---|---|---|
| TP53 | 27 | TP53 | 0.314625726 | TP53 | 0.461139896 |
| SRC | 23 | ESR1 | 0.20892383 | ESR1 | 0.458762887 |
| PIK3CA | 21 | CYP19A1 | 0.19645871 | MAPK8 | 0.419811321 |
| PIK3CB | 18 | SRC | 0.134042024 | AKT1 | 0.41588785 |
| PIK3R1 | 18 | MAPK8 | 0.107051319 | CTNNB1 | 0.413953488 |
| ESR1 | 18 | GSTP1 | 0.084028189 | PIK3CA | 0.412037037 |
| AKT1 | 18 | CYP3A4 | 0.083869476 | EGFR | 0.402714932 |
| PIK3CD | 17 | AKT1 | 0.082982932 | SRC | 0.395555556 |
| CTNNB1 | 16 | CASP3 | 0.072349842 | ERBB2 | 0.39380531 |
| CYP3A4 | 16 | PTGS2 | 0.070442016 | CYP19A1 | 0.388646288 |
| EGFR | 15 | PIK3CA | 0.069218826 | CCND1 | 0.388646288 |
| CYP1A1 | 15 | CYP1A1 | 0.064888167 | BCL2 | 0.381974249 |
| IGF1R | 13 | CTNNB1 | 0.045637558 | MDM2 | 0.38034188 |
| ERBB2 | 12 | HIF1A | 0.045168344 | CASP3 | 0.378723404 |
| CCND1 | 12 | TOP2A | 0.039542082 | HIF1A | 0.378723404 |
| CYP1A2 | 12 | UGT2B7 | 0.035694459 | ESR2 | 0.377118644 |
| UGT2B7 | 11 | EGFR | 0.03368072 | PIK3R1 | 0.375527426 |
| CYP2A6 | 10 | CCND1 | 0.031433278 | AR | 0.375527426 |
| CYP2C9 | 10 | PIK3R1 | 0.03136549 | PTGS2 | 0.372384937 |
| BCL2 | 10 | CYP2C9 | 0.031137693 | IGF1R | 0.367768595 |
PPI: Protein protein interaction.
3.4.3. PI3K/Akt signaling pathway was enriched in GO and KEGG analysis
In the results of the GO enrichment analysis, terms such as “phosphatidylinositol 3-kinase signaling” were enriched in Biological Processes (BP); “phosphatidylinositol 3-kinase complex” and “cyclin-dependent protein kinase holoenzyme complex” were enriched in Cellular Components (CC); and “protein tyrosine kinase activity” was enriched in Molecular Functions (MF) (Figure 6f). KEGG enrichment analysis revealed that terms including “PI3K-Akt signaling pathway”, “Breast cancer”, and “p53 signaling pathway” were among the top 20 terms arranged by gene count (Figure 6g).
3.5. PI3K/Akt signaling pathway was inhibited in MDA-MB-231
Since the PI3K/Akt signaling pathway was enriched significantly in network pharmacology, we checked the gene expression in this pathway. As shown in Figure 7a, most genes were downregulated including PIK3CB, PIK3CD, PIK3R2, PIK3R3, AKT1, and AKT2, while upstream regulators like ITGB1, ITGB4, ITGA6, ITGA2, and ITGAV were downregulated as well. We also observed that BF, BFK, and BFS groups had different DEGs at high concentrations, though they shared some of them (Figure 3b). To explore the effects of different DEGs, these genes were analyzed in CMAP. As a result, the common gene regulation profile of the three groups was similar to proteasome inhibitor, NFκB inhibitor, HDAC inhibitor, topoisomerase inhibitor, PI3K inhibitor, MTOR inhibitor, and CDK inhibitor (Figure 7b). The work pattern of DEGs regulated only in BF or BFK group was the same as common DEGs (Figure 7c, d). However, profiling of DEGs only in the BFS group behaved more like a protein synthesis inhibitor (Figure 7e). The above indicated that BF, BFK, and BFS all inhibited the PI3K/Akt signaling pathway.
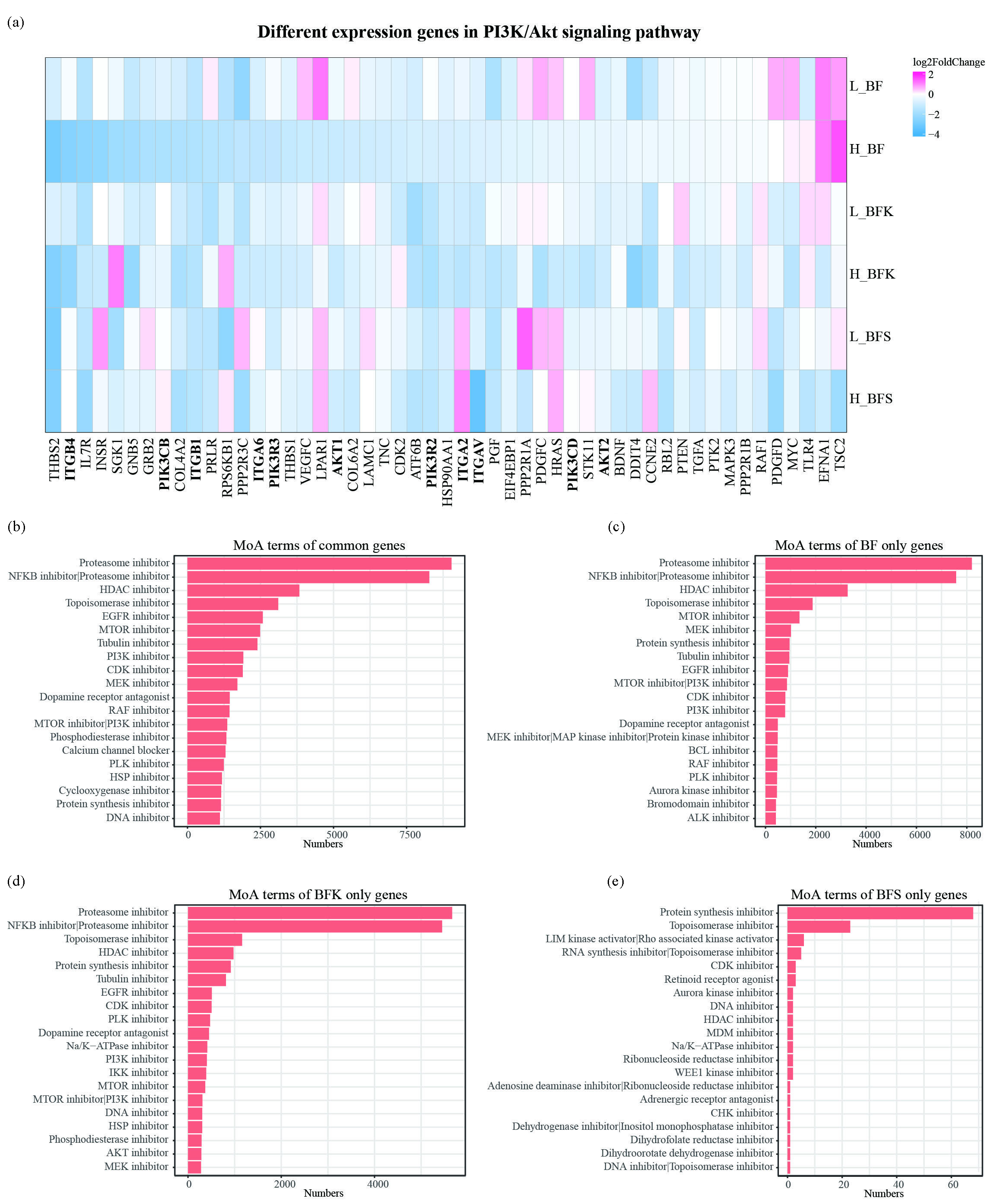
- BF, BFK, and BFS inhibited the PI3K/Akt signaling pathway. (a) DEGs in PI3K/Akt signaling pathway. L means the low concentration set at 15 μg/mL, while H means the high concentration at 25 μg/mL. Top 20 Mode of action analysis (MoA) terms analyzed in CMAP using common DEGs or (b) particular DEGs in groups treated with ethanolic extracts of (c) BF, (d) BFK, and (e) BFS at the concentration of 25 μg/mL. BF: Bruceae Fructus, BFK: Kernels of Bruceae Fructus, BFS: Shell of Bruceae Fructus, DEGs: Differentially expressed genes, PI3K/Akt: Phosphoinositide 3-kinase/protein kinase B, CMAP: Connectivity map, MoA: Mode of action.
3.6. Quassinoids and flavones in BF bind to targets in PI3K/Akt pathway
According to the results of HTS2 and network pharmacology analysis, the PI3K/Akt signaling pathway might be key in treating breast cancer. The PI3K/Akt signaling pathway is thought to promote cell growth, survival, and cell cycle progression [50]. However, it is aberrantly activated in various cancers, which results in bad diagnoses and prognoses [51]. To explore what compounds affect the PI3K/Akt pathway, we chose several key proteins of this pathway to conduct molecular docking. Crystal structures of PI3Kα (PDB ID: 6PYS), PI3Kγ (PDB ID: 4ANV), AKT1 (PDB ID: 7NH4), and PDK1 (PDB ID: 3QC4) were downloaded from RCSB PDB, and 3D structures of compounds in BF were prepared.
With a threshold of gscore < -6, we identified 112 compounds probably binding to PI3Kα, 92 to PI3Kγ, 117 to AKT1, and 77 to PDK1 (Table A3). Except for flavones like rutin and isovitexin, several quassinoids could bind to these targets. Yadanzigan binds to PI3Kα (docking gscore: -10.637), PI3Kγ (docking gscore: -10.133), or AKT1 (docking gscore: -11.645) by hydrogen-bonding with amino acid residues of the protein. In the interaction with PI3Kα, the hydroxyl group at C1, C11, C14, C4’, and C6’ of yadanzigan interacted with Ser774, Gln859, Lys802, and Asp933 respectively, while the ketone group at C16 worked with Thr856 (Figure 8a). With PI3Kγ, the hydroxyl group at C1, C1’, C12, and C14 formed hydrogen-bonds with Ala805, Ser806, and Asp954, and that at C6’ with Lys802 and Val803 (Figure 8b). As for AKT1, the hydroxyl group at C1, C11, C12, C14, and C3’ interacted with Lys276, Asn279, Glu278, Leu156, TYR18, and Asp274 (Figure 8c). And glycosyl of bruceoside B (docking gscore: -8.595) inserted into the PDK1 pocket by hydrogen-bonding with Ser160 and Ala162 (Figure 8d). Together, our analysis suggested that quassinoids and flavones in BF might bind to key targets in the PI3K/Akt signaling pathway.
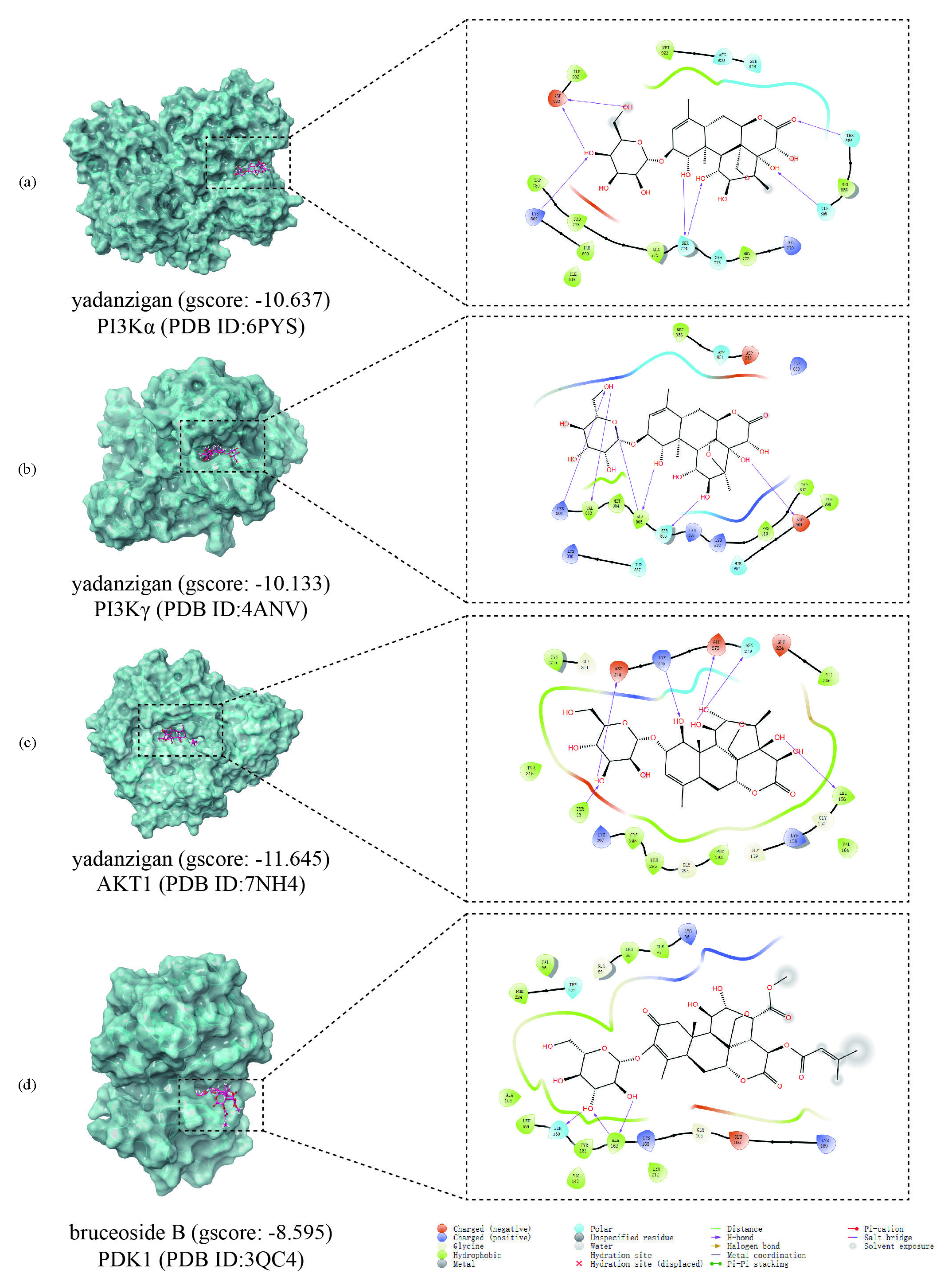
- Representative molecular docking results of breast cancer targets. (a) The patterns of yadanzigan binding to PI3Kα, (b) PI3Kγ, (c) AKT1, and (d) bruceoside B binding to PDK1.
3.7 Discussion
Traditional Chinese medicine (TCM) has been used in China for more than 2000 years. Although animals, insects, or minerals can be the sources of TCM, most TCM uses plants as sources. Long-term clinical practice has developed the principles of certain positions in a plant exerting particular therapeutical effects. For example, peels and seeds of Trichosanthis Fructus (“Gualou” in Chinese) are always used separately. The seeds can loosen the bowels and relieve constipation, while the pericarps prefer to clear heat, resolve sputum, and promote the flow of qi to soothe chest oppression according to the TCM theory [52]. As a result, non-medicinal parts of plants are usually abandoned in the process of actual production. It is no doubt a waste of great amounts of plant biomass. Incineration, stacking, and burying are traditional methods to dispose the waste, meanwhile ethanol and biogas preparation, biofuel and bioenergy production, paper making, and bioactive components extraction are also ways of proper utilization of the waste in industry [53,54]. However, potential medicinal values should be assessed first in terms of TCM residues to fulfill sustainable practices in pharmaceutical development. This study practiced a strategy that data mining and UPLC-MS/MS reveal the phytochemical profiles together with network pharmacology, HTS2, and experimental validation exploring pharmacological activities. BFS was proven to exert an anti-breast cancer effect by inhibiting proliferation and migration, arresting the cell cycle, and inducing apoptosis. The bioactive compounds are mainly flavones and quassinoids, which target the PI3K/Akt pathway. These compounds could be candidates for exploring PI3K/Akt inhibitors in treating breast cancer. Regarding other agro-industrial residues or underutilized medicinal plants, the integrative strategy could begin by identifying their chemical composition using UPLC-MS/MS. Based on traditional uses or areas of interest, potential disease models and cell models could then be selected. The gene expression profiles of cells treated with extracts from these residues or plants could be obtained through the HTS2 assay. Ultimately, this approach may lead to the discovery of potential compounds targeting specific diseases.
B. javanica is also called “Kusum” in Malaysia, “Kuwalot” in Indonesia, “balaniog” in Philippines, and “ratchadat” in Thailand. It is used for treating malaria [55,56]. In Cambodia, the roots are anthelmintic and can be used in the treatment of malaria and dysentery [57]. And the leaves have effects on acne [58]. However, there are no relevant products in use, manufactured with these parts. The life span of B. javanica is only a few years. Thus, it is worth studying the usage of the roots, stems, barks, and leaves without worrying about the production of BF. Perhaps, the strategy proposed in this study could help discover the economic value of these parts. Certainly, it would provide new insights into discovering proper utilization of other agro-industrial residues. By now, over one hundred quassinoids of great pharmacological activities such as anti-cancer, anti-inflammatory, and anti-malaria have been isolated from this plant [12,13,15,59]. But none have been approved for clinical uses because of severe side effects [60-62]. In the future, the chemicals isolations from this plant, or structure modification of quassinoids should be carried out.
With a data mining approach to profiling the phytochemical basis of BF, about 167 isolated compounds were collected, of which most were quassinoids and flavones. The results of network pharmacology and molecular docking also suggested that quassinoids and flavones were the main active ingredients targeting the PI3K/Akt pathway. PI3K is a critical protein mediating tumor cell growth, proliferation, and metabolism [63]. The PI3K/Akt signaling pathway is thought to promote cell growth, survival, and cell cycle progression [50]. However, it is aberrantly activated due to multiple genomic alterations of PIK3CA and AKT across various cancers leading to bad diagnosis and prognosis [51,64]. PIK3CA mutations have been identified in approximately 10% of TNBC cases. By now, only three PI3K/Akt inhibitors alpelisib, capivasertib, and everolimus have been approved by FDA for breast cancer treatment. Therapeutic resistance of PI3K/AKT inhibitors is associated with receptor tyrosine kinase reactivation, acquired mutation and/or amplification of PIK3CA or PIK3CB, and PI3K reactivation [50]. As Figure 7(a) exhibits, most genes in the PI3K/Akt pathway were downregulated by BF, BFK, and BFS, which illustrates that the PI3K/Akt signaling pathway was inhibited more effectively.
UPLC-MS/MS revealed that BFS contains more flavones like quercetin, quercetin-3β-D-glucoside, kaempferol, and luteolin than BFK. Quercetin is a common dietary flavone and has been proven to exert anti-breast cancer effects by inducing apoptosis and arresting the cell cycle [65]. Kaempferol suppresses the growth of breast cancer cells by upregulating the expression of p21 and bax [66]. In addition, it effectively decreases the incidence of breast precancerous lesions by activating the LKB1/AMPK pathway and inducing excessive mitochondrial fission and lethal mitophagy [67]. Another flavone, luteolin, could suppress the proliferation and metastasis of breast cancer cells by inactivating the AKT/mTOR pathway [68]. There are also other flavones exerting anti-TNBC effects. Sophoraflavanone G was reported to inhibit proliferation and metastasis by inactivating the EGFR/PI3K/Akt signaling pathway [69]. And both fisetin and chrysin could inhibit metastasis of TNBC [70,71]. Flavones like quercetin, kaempferol, and luteolin were seldom reported to induce toxicity or side effects. However, a high dose of quercetin might induce kidney-related issues, headaches, and stomach aches [72]. Although these flavones exert outstanding anti-cancer effects, their oral bioavailability is extremely poor, which limits their application in clinical settings [73,74]. Besides, quassinoids like brusatol, bruceine A, bruceine D, and bruceine H also exhibit outstanding anti-cancer effects in various types of cancers. The common dosage of these quassinoids in animal models is 1 mg/kg or 2 mg/kg (i.p or i.v). Although their anti-cancer effects are obvious, short t1/2 and MRT might increase times of doses and induce severe side effects [75,76]. Bruceantin was considered in clinical trials treating metastatic breast carcinoma and advanced malignant melanoma [60,62] but failed because of poor effects and severe side effects, including nausea, vomiting, mild hypotension, and fever. Their abilities of anti-breast cancer have seldom been evaluated. Approximately 97 quassinoids have been identified from BF, but the pharmacological activities of only several compounds were explored. Thus, the research of BF in the future could focus on these aspects.
TNBC is defined as a “cold tumor” characterized by a lack of effective immune cell infiltration within the tumor microenvironment. This feature poses a significant challenge to immunotherapy approaches, including PD-1/PD-L1 inhibitors. Only 10% of TNBC patients could response to checkpoint blockade immunotherapies like PD-1/PD-L1 inhibitors [77]. Scientists make efforts to turn “cold tumors” into “hot tumors” to treat breast cancer. Some inhibitors were identified to promote infiltration of effective CD8+ T cells and work better with immune checkpoint inhibitors [77,78]. PI3Kβ, encoded by PIK3CB, was found to promote immune escape in PTEN-null tumors [79]. In this study, BF treatment could decrease the expression of PIK3CB (Figure 7a), suggesting that BF might inhibit PI3Kβ and promote anti-tumor immune responses. Except for PI3K/Akt signaling pathway, cell cycle, and apoptosis, we identified other interesting pathways that might treat breast cancer. In this study, “IL−17 signaling pathway” was enriched. IL-17E, also known as IL-25, is a subtype in the IL-17 family. Studies have shown that IL-17E can induce apoptosis of breast cancer cells, and related treatment can reduce tumor growth [80,81]. Also, it can attract lymphocytes [82]. In addition, some immune pathways like “NOD-like receptor signaling pathway,” “T cell receptor signaling pathway,” “Toll-like receptor signaling pathway,” and “B cell receptor signaling pathway” were significantly enriched (Figure 3c and d), suggesting that BFS might communicate with immune cells in the immunosuppressive microenvironment. Thus, it is worth exploring whether BFS could attract more effective immune cells like T-cells or NK cells to the environment.
4. Conclusions
In summary, BFS was proven to exhibit an anti-breast cancer effect through integrating network pharmacology, HTS2 technology, and experimental validation. Meanwhile, UPLC-MS/MS revealed that BFS contains quite a few flavones and quassinoids, which showed binding abilities to key targets in the PI3K/Akt pathway according to the results of molecular docking. Transcriptome data suggested that BF, BFK, and BFS could inhibit the proliferation and migration of breast cancer cells, arrest the cell cycle, and induce apoptosis. And these pharmacological activities were all validated in breast cancer cells. As for molecular mechanisms, the PI3K/Akt pathway was predicted through network pharmacology and validated via HTS2 technology. BF, BFK, and BFS all had strong abilities of inhibition effects on this pathway. Also, together with the downregulation of genes in the cell cycle, the upregulation of genes in the TP53 pathway and apoptosis, were also proved using HTS2 technology. What’s more, this research strategy provided new insights into the utilization of agro-industrial residues.
Acknowledgment
This work was supported by the National Key R&D Program of China (No. 2023YFF0720300), Key Projects of Science and Technology Plan of Inner Mongolia Autonomous Region (Grant No. 201802115), the National Natural Science Foundation of China (Grant No. 82172723), Science and Technology Department of Sichuan Province (Grant No. 2021ZYD0079), Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (No. ZYYCXTD-D-202209).
CRediT authorship contribution statement
Xiankuo Yu: original draft writing, methodology, investigation, validation, conceptualization, visualization. Lei Xiang: review and edit the draft, methodology, investigation. Jun An: review and edit the draft, methodology, validation. Shengrong Li: methodology, investigation, validation. Chao Hu: methodology, validation. Yu Gui: methodology, validation. Yumei Wang: review and edit the draft, validation. Xilinqiqige Bao: review and edit the draft, supervision, funding acquisition, conceptualization. Dong Wang: review and edit the draft, supervision, funding acquisition, conceptualization. All authors have read and approved the final manuscript.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Declaration of generative AI and AI-assisted technologies in the writing process
The authors confirm that there was no use of artificial intelligence (AI)-assisted technology for assisting in the writing or editing of the manuscript and no images were manipulated using AI.
Supplementary data
Supplementary file: Table A1. Chemical ingredients isolated from BF and their targets predicted in SwissTargetPrediction. Table A2. Information of therapeutic targets of breast cancer. Table A3. Molecular docking results of compounds in BF binding to PI3Kα, PI3Kγ, AKT1, and PDK1. Figure A1. Quality evaluation of HTS2 data. Figure A2. GSEA in different gene sets.
References
- Analysis of curative effect of Brucea Javanica oil emulsion combined with TP chemotherapy in treatment of small cell lung cancer. Liaoning Journal of Traditional Chinese Medicine. 2016;23:320-2. https://doi.org/10.13192/j.issn.1000-1719.2016.02.039
- [Google Scholar]
- Clinical observation of combination of Bruceolic Oil Emulsion, Aidi Injection and chemotherapy in treating advanced non-small cell lung cancer. Chinese Medicine. 2014;46:152-4. https://doi.org/10.13457/j.cnki.jncm.2014.02.091
- [Google Scholar]
- Effect of FOLFOX4 combined with brucea javanica emulsion on VEGF in patients with gastric cancer. Oncology Letters. 2018;15:1079-1083. https://doi.org/10.3892/ol.2017.7429
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Effect of postoperative application of Brucea Javanica Oil Injection adjuvant chemotherapy in treating gastric cancer. Chinese Journal of Experimental Traditional Medical Formulae. 2014;20:178-180. https://doi.org/10.13422/j.cnki.syfjx.2014180178
- [Google Scholar]
- Brucea javanica oil emulsion significantly improved the effect of anti–programmed cell death protein-1 immunotherapy. Phytomedicine. 2022;107:154446. https://doi.org/10.1016/j.phymed.2022.154446
- [CrossRef] [PubMed] [Google Scholar]
- Bruceae fructus oil inhibits triple-negative breast cancer by restraining autophagy: Dependence on the gut microbiota-mediated amino acid regulation. Frontiers in Pharmacology. 2021;12:727082. https://doi.org/10.3389/fphar.2021.727082
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Seed oil of brucea javanica induces apoptosis through the PI3K/Akt signaling pathway in acute lymphocytic leukemia jurkat cells. Chinese Journal of Natural Medicines. 2021;19:608-620. https://doi.org/10.1016/S1875-5364(21)60060-2
- [CrossRef] [PubMed] [Google Scholar]
- Involvement of autophagy inhibition in brucea javanica oil emulsion-induced colon cancer cell death. Oncology Letters. 2015;9:1425-1431. https://doi.org/10.3892/ol.2015.2875
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Optimization of biodiesel production from brucea javanica seeds oil as novel non-edible feedstock using response surface methodology. Energy Conversion and Management. 2017;149:392-400. https://doi.org/10.1016/j.enconman.2017.07.037
- [CrossRef] [Google Scholar]
- Determination of brusatol in Brucea javanica of different materials and parts. Asia-Pacific Traditional Medicine. 2016;12:54-6. https://doi.org/10.11954/ytctyy.201615022
- [Google Scholar]
- Brusatol enhances the efficacy of chemotherapy by inhibiting the Nrf2-mediated defense mechanism. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:1433-8. https://doi.org/10.1073/pnas.1014275108
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Brusatol suppresses the tumor growth and metastasis of colorectal cancer via upregulating ARRDC4 expression through modulating PI3K/YAP1/TAZ pathway. Phytomedicine. 2023;109:154567. https://doi.org/10.1016/j.phymed.2022.154567
- [CrossRef] [PubMed] [Google Scholar]
- Brusatol has therapeutic efficacy in non-small cell lung cancer by targeting Skp1 to inhibit cancer growth and metastasis. Pharmacological Research. 2022;176:106059. https://doi.org/10.1016/j.phrs.2022.106059
- [CrossRef] [PubMed] [Google Scholar]
- The flavonoid brusatol induces apoptosis in aggressive lymphoma cells and exhibits synergistic effect with venetoclax. Blood. 2022;140:4945-6. https://doi.org/10.1182/blood-2022-166678
- [CrossRef] [Google Scholar]
- In vitro antimalarial and cytotoxic activities of semisynthetic derivatives of brusatol. European Journal of Medicinal Chemistry. 1993;28:265-9. https://doi.org/10.1016/0223-5234(93)90143-3
- [CrossRef] [Google Scholar]
- Antimalarial agents, 4 Synthesis of a brusatol analog and biological activity of brusatol-related compounds. Journal of Natural Products. 1987;50:847-851. https://doi.org/10.1021/np50053a012
- [CrossRef] [PubMed] [Google Scholar]
- Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians. 2021;71:209-249. https://doi.org/10.3322/caac.21660
- [CrossRef] [PubMed] [Google Scholar]
- Brucea javanica: A review on anticancer of its pharmacological properties and clinical researches. Phytomedicine. 2021;86:153560. https://doi.org/10.1016/j.phymed.2021.153560
- [CrossRef] [PubMed] [Google Scholar]
- Ethanol extract of brucea javanica seed inhibit triple-negative breast cancer by restraining autophagy via PI3K/Akt/mTOR pathway. Frontiers in Pharmacology. 2020;11:606. https://doi.org/10.3389/fphar.2020.00606
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- RASL-seq for massively parallel and quantitative analysis of gene expression. CP Molecular Biology. 2012;98 https://doi.org/10.1002/0471142727.mb0413s98
- [Google Scholar]
- High-throughput strategies for the discovery of anticancer drugs by targeting transcriptional reprogramming. Frontiers in Oncology. 2021;11:762023. https://doi.org/10.3389/fonc.2021.762023
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- A large-scale transcriptional study reveals inhibition of COVID-19 related cytokine storm by traditional chinese medicines. Science Bulletin. 2021;66:884-8. https://doi.org/10.1016/j.scib.2021.01.005
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Network pharmacology: Towards the artificial intelligence-based precision traditional chinese medicine. Briefings in Bioinformatics. 2023;25:bbad518. https://doi.org/10.1093/bib/bbad518
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Multi-technology integrated network pharmacology-based study on phytochemicals, active metabolites, and molecular mechanism of psoraleae fructus to promote melanogenesis. Journal of Ethnopharmacology. 2024;325:117755. https://doi.org/10.1016/j.jep.2024.117755
- [CrossRef] [PubMed] [Google Scholar]
- Integrated network pharmacology and experimental validation-based approach to reveal the underlying mechanisms and key material basis of jinhua qinggan granules against acute lung injury. Journal of Ethnopharmacology. 2024;326:117920. https://doi.org/10.1016/j.jep.2024.117920
- [CrossRef] [PubMed] [Google Scholar]
- Yupingfeng san exhibits anticancer effect in hepatocellular carcinoma cells via the MAPK pathway revealed by HTS2 technology. Journal of Ethnopharmacology. 2023;306:116134. https://doi.org/10.1016/j.jep.2023.116134
- [CrossRef] [PubMed] [Google Scholar]
- Fiji: An open-source platform for biological-image analysis. Nature Methods. 2012;9:676-682. https://doi.org/10.1038/nmeth.2019
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Guizhi fuling decoction inhibiting the PI3K and MAPK pathways in breast cancer cells revealed by HTS2 technology and systems pharmacology. Computational and Structural Biotechnology Journal. 2020;18:1121-1136. https://doi.org/10.1016/j.csbj.2020.05.004
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology. 2014;15:550. https://doi.org/10.1186/s13059-014-0550-8
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell. 2017;171:1437-1452. https://doi.org/10.1016/j.cell.2017.10.049
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2023.
- clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb). 2021;2:100141. https://doi.org/10.1016/j.xinn.2021.100141
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Welcome to the tidyverse. Journal of Open Source Software. 2019;4:1686. https://doi.org/10.21105/joss.01686
- [CrossRef] [Google Scholar]
- SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Research. 2019;47:W357-W364. https://doi.org/10.1093/nar/gkz382
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Mendelian Inheritance in Man: A catalog of human genes and genetic disorders (12th ed). Baltimore: Johns Hopkins University Press; 1998.
- TTD: Therapeutic target database describing target druggability information. Nucleic Acids Research. 2024;52:D1465-D1477. https://doi.org/10.1093/nar/gkad751
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- An evidence-Based framework for evaluating pharmacogenomics knowledge for personalized medicine. Clinical Pharmacology & Therapeutics. 2021;110:563-572. https://doi.org/10.1002/cpt.2350
- [PubMed] [Google Scholar]
- DrugBank 6.0: The drugBank knowledgebase for 2024. Nucleic Acids Research. 2024;52:D1265-D1275. https://doi.org/10.1093/nar/gkad976
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- STRING v10: Protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Research. 2015;43:D447-D452. https://doi.org/10.1093/nar/gku1003
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- The protein data bank. Nucleic Acids Research. 2000;28:235-242. https://doi.org/10.1093/nar/28.1.235
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein–Ligand complexes. Journal of Medicinal Chemistry. 2006;49:6177-6196. https://doi.org/10.1021/jm051256o
- [CrossRef] [PubMed] [Google Scholar]
- Skp2 in the ubiquitin-proteasome system: A comprehensive review. Medicinal Research Reviews. 2020;40:1920-1949. https://doi.org/10.1002/med.21675
- [CrossRef] [PubMed] [Google Scholar]
- Cell cycle regulation: P53-p21-RB signaling. Cell Death and Differentiation. 2022;29:946-960. https://doi.org/10.1038/s41418-022-00988-z
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- DNA damage and p53-mediated cell cycle arrest: A reevaluation. Proceedings of the National Academy of Sciences of the United States of America. 1996;93:15209-15214. https://doi.org/10.1073/pnas.93.26.15209
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Caspase-7 activates ASM to repair gasdermin and perforin pores. Nature. 2022;606:960-7. https://doi.org/10.1038/s41586-022-04825-8
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- TXNIP: A key protein in the cellular stress response pathway and a potential therapeutic target. Experimental & Molecular Medicine. 2023;55:1348-1356. https://doi.org/10.1038/s12276-023-01019-8
- [CrossRef] [PubMed] [Google Scholar]
- Identification of genomic binding sites and direct target genes for the transcription factor DDIT3/CHOP. Experimental Cell Research. 2023;422:113418. https://doi.org/10.1016/j.yexcr.2022.113418
- [CrossRef] [PubMed] [Google Scholar]
- GADD45A inhibits autophagy by regulating the interaction between BECN1 and PIK3C3. Autophagy. 2015;11:2247-2258. https://doi.org/10.1080/15548627.2015.1112484
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer. Molecular Cancer. 2023;22:138. https://doi.org/10.1186/s12943-023-01827-6
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Targeting PI3K/Akt signal transduction for cancer therapy. Signal Transduction and Targeted Therapy. 2021;6:425. https://doi.org/10.1038/s41392-021-00828-5
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Trichosanthis fructus: Botany, traditional uses, phytochemistry and pharmacology. Journal of Ethnopharmacology. 2018;224:177-194. https://doi.org/10.1016/j.jep.2018.05.034
- [CrossRef] [PubMed] [Google Scholar]
- Treatment and bioresources utilization of traditional chinese medicinal herb residues: Recent technological advances and industrial prospect. Journal of Environmental Management. 2021;299:113607. https://doi.org/10.1016/j.jenvman.2021.113607
- [CrossRef] [PubMed] [Google Scholar]
- Comprehensive utilization of chinese medicine residues for industry and environment protection: Turning waste into treasure. Journal of Cleaner Production. 2021;279:123856. https://doi.org/10.1016/j.jclepro.2020.123856
- [CrossRef] [Google Scholar]
- Screening of plant extracts for antiprotozoal and cytotoxic activities. Journal of Ethnopharmacology. 2003;89:185-191. https://doi.org/10.1016/s0378-8741(03)00269-1
- [CrossRef] [PubMed] [Google Scholar]
- Screening of Indonesian medicinal plant extracts for antibabesial activity and isolation of new quassinoids from brucea javanica. Journal of Natural Products. 2007;70:1654-7. https://doi.org/10.1021/np070236h
- [CrossRef] [PubMed] [Google Scholar]
- Screening of selected indigenous plants of Cambodia for antiplasmodial activity. Journal of Ethnopharmacology. 2006;107:12-8. https://doi.org/10.1016/j.jep.2006.01.028
- [CrossRef] [PubMed] [Google Scholar]
- Potency of Indonesian medicinal plants as tyrosinase inhibitor and antioxidant agent. Journal of Biological Sciences. 2010;10:138-144. https://doi.org/10.3923/jbs.2010.138.144
- [CrossRef] [Google Scholar]
- Brusatol ameliorates 2, 4, 6-trinitrobenzenesulfonic acid-induced experimental colitis in rats: Involvement of NF-κB pathway and NLRP3 inflammasome. International Immunopharmacology. 2018;64:264-274. https://doi.org/10.1016/j.intimp.2018.09.008
- [CrossRef] [PubMed] [Google Scholar]
- A phase II study of bruceantin (NSC-165, 563) in advanced malignant melanoma. Investigational New Drugs. 1983;1:239-242. https://doi.org/10.1007/BF00208896
- [CrossRef] [PubMed] [Google Scholar]
- Multiple myeloma regression mediated by bruceantin. Clinical Cancer Research. 2004;10:1170-9. https://doi.org/10.1158/1078-0432.ccr-0362-3
- [CrossRef] [PubMed] [Google Scholar]
- Phase II trial of bruceantin in metastatic breast carcinoma. American Journal of Clinical Oncology. 1982;5:389-391. https://doi.org/10.1097/00000421-198208000-00007
- [CrossRef] [PubMed] [Google Scholar]
- Recent advances in targeted strategies for triple-negative breast cancer. Journal of Hematology & Oncology. 2023;16:100. https://doi.org/10.1186/s13045-023-01497-3
- [CrossRef] [PubMed] [Google Scholar]
- Natural products targeting the PI3K-akt-mTOR signaling pathway in cancer: A novel therapeutic strategy. Seminars in Cancer Biology. 2022;80:1-17. https://doi.org/10.1016/j.semcancer.2019.12.008
- [CrossRef] [PubMed] [Google Scholar]
- Therapeutic potential of quercetin on human breast cancer in different dimensions. Inflammopharmacology. 2020;28:39-62. https://doi.org/10.1007/s10787-019-00660-y
- [CrossRef] [PubMed] [Google Scholar]
- Treatment with kaempferol suppresses breast cancer cell growth caused by estrogen and triclosan in cellular and xenograft breast cancer models. The Journal of Nutritional Biochemistry. 2016;28:70-82. https://doi.org/10.1016/j.jnutbio.2015.09.027
- [CrossRef] [PubMed] [Google Scholar]
- Kaempferol induces mitochondrial dysfunction and mitophagy by activating the LKB1/AMPK/MFF pathway in breast precancerous lesions. Phytotherapy Research. 2023;37:3602-3616. https://doi.org/10.1002/ptr.7838
- [CrossRef] [PubMed] [Google Scholar]
- Luteolin suppresses androgen receptor-positive triple-negative breast cancer cell proliferation and metastasis by epigenetic regulation of MMP9 expression via the AKT/mTOR signaling pathway. Phytomedicine. 2021;81:153437. https://doi.org/10.1016/j.phymed.2020.153437
- [CrossRef] [PubMed] [Google Scholar]
- Sophoraflavanone G suppresses the progression of triple-negative breast cancer via the inactivation of EGFR–PI3K–AKT signaling. Drug Development Research. 2022;83:1138-1151. https://doi.org/10.1002/ddr.21938
- [CrossRef] [PubMed] [Google Scholar]
- Fisetin inhibited growth and metastasis of triple-negative breast cancer by reversing epithelial-to-mesenchymal transition via PTEN/Akt/GSK3β signal pathway. Frontiers in Pharmacology. 2018;9:772. https://doi.org/10.3389/fphar.2018.00772
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Chrysin inhibits metastatic potential of human triple-negative breast cancer cells by modulating matrix metalloproteinase-10, epithelial to mesenchymal transition, and PI3K/Akt signaling pathway. Journal of Applied Toxicology. 2014;34:105-112. https://doi.org/10.1002/jat.2941
- [CrossRef] [PubMed] [Google Scholar]
- Therapeutic potentials and targeting strategies of quercetin on cancer cells: Challenges and future prospects. Phytomedicine. 2024;133:155902. https://doi.org/10.1016/j.phymed.2024.155902
- [CrossRef] [PubMed] [Google Scholar]
- Novel extraction techniques and pharmaceutical activities of luteolin and its derivatives. Journal of Food Biochemistry. 2019;43:e12974. https://doi.org/10.1111/jfbc.12974
- [CrossRef] [PubMed] [Google Scholar]
- Kaempferol efficacy in metabolic diseases: Molecular mechanisms of action in diabetes mellitus, obesity, non-alcoholic fatty liver disease, steatohepatitis, and atherosclerosis. Biomedicine & Pharmacotherapy. 2024;175:116694. https://doi.org/10.1016/j.biopha.2024.116694
- [CrossRef] [PubMed] [Google Scholar]
- HPLC–MS/MS method for bioavailability study of bruceines D & E in rat plasma. Journal of Chromatography B. 2017;1063:183-8. https://doi.org/10.1016/j.jchromb.2017.08.037
- [Google Scholar]
- Pharmacokinetic, metabolic profiling and elimination of brusatol in rats. Biomedical Chromatography. 2018;32:e4358. https://doi.org/10.1002/bmc.4358
- [CrossRef] [PubMed] [Google Scholar]
- Tumor immunological phenotype signature-based high-throughput screening for the discovery of combination immunotherapy compounds. Science Advances. 2021;7:eabd7851. https://doi.org/10.1126/sciadv.abd7851
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- TRIB3 reduces CD8+ T cell infiltration and induces immune evasion by repressing the STAT1-CXCL10 axis in colorectal cancer. Science Translational Medicine. 2022;14:eabf0992. https://doi.org/10.1126/scitranslmed.abf0992
- [CrossRef] [PubMed] [Google Scholar]
- PI3Kβ controls immune evasion in PTEN-deficient breast tumours. Nature. 2023;617:139-146. https://doi.org/10.1038/s41586-023-05940-w
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- IL-17E, a proinflammatory cytokine, has antitumor efficacy against several tumor types in vivo. Cancer Immunology, Immunotherapy. 2010;59:805-817. https://doi.org/10.1007/s00262-009-0802-8
- [CrossRef] [Google Scholar]
- The potential role and status of IL-17 family cytokines in breast cancer. International Immunopharmacology. 2021;95:107544. https://doi.org/10.1016/j.intimp.2021.107544
- [CrossRef] [PubMed] [Google Scholar]
- The interleukin-17 family of cytokines in breast cancer. International Journal of Molecular Sciences. 2018;19:3880. https://doi.org/10.3390/ijms19123880
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]







