Translate this page into:
Synthesis, α-glucosidase inhibition and molecular docking studies of natural product 2-(2-phenyethyl)chromone analogues
⁎Corresponding author. pengzhiyun1986@163.com (Zhiyun Peng), mailofhy@126.com (Yong Huang), wanggch123@163.com (Guangcheng Wang)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
A series of natural product (2-phenyethyl)chromone analogues (3–34) were designed, synthesized, and screened for their α-glucosidase inhibitory activity. The results indicated that some of the synthesized derivatives displayed inhibitory activities against α-glucosidase with IC50 values ranging from 11.72 ± 0.08 to 85.58 ± 2.30 μM when compared to the standard drug acarbose (IC50 = 832.22 ± 2.00 μM). Among them, compound 4 with a hydroxyl group at the 7-position of chromone and a chloro group at the 4-position of the benzene ring, displayed the most significant inhibitory activity with the IC50 value of 11.72 ± 0.08 μM. The inhibitory mechanism of compound 4 against α-glucosidase was studied by enzyme kinetic, circular dichroism spectra, fluorescence quenching, and molecular docking. Sucrose loading test in vivo further demonstrated that it could decrease blood glucose levels after sucrose administration in normal Kunming mice. In vitro cytotoxicity showed that 4 exhibited low cytotoxicity against normal human cell lines. The ADME study suggested that all compounds are likely to be orally active as they obeyed Lipinski’s rule of five. In summary, our studies showed that these derivatives are a new class of α-glucosidase inhibitors.
Keywords
2-(2-Phenylethyl)chromone
α-Glucosidase inhibitor
Synthesis
1 Introduction
Diabetes mellitus (DM) is one of the most common chronic metabolic diseases in the world, which is characterized by hyperglycemia with plenty of serious complications, including cardiomyopathy, nephropathy, retinopathy, and neuropathy (Huang et al., 2020a, 2020b; Javid et al., 2018; Kasturi et al., 2017; Ogurtsova et al., 2017). Controlling postprandial hyperglycemia is necessary for the early therapy for diabetes that reduces the risk of developing long-term, micro- and macrovascular complications. α-Glucosidase exists on the small intestine brush border, which hydrolyzes the glycosidic bonds of polysaccharides to monosaccharides like glucose, this is mainly responsible to cause hyperglycemia (Fallah et al., 2022; Shin et al., 2019). Therefore, α-glucosidase inhibitors were recognized as available therapeutic drugs for treating type 2 diabetes by the maintenance of postprandial blood glucose at a normal or near-normal level (Choi et al., 2018; Perera et al., 2016). Three α-glucosidase inhibitors (acarbose, miglitol, and voglibose) are being clinically used for the treatment of type 2 diabetes (DiNicolantonio et al., 2015). However, α-glucosidase inhibitors are associated with gastrointestinal side effects including bloating pain, flatulence, diarrhea, abdominal discomfort, and enteritis for long-term use (Li et al., 2020). Therefore, the design and development of more potent and specific α-glucosidase inhibitors to treat DM are essential.
2-(2-Phenylethyl)chromones (PECs) are a kind of naturally occurring products, which are mainly isolated from Aquilaria sinensis (Lour.) Gilg (Fan et al., 2022). They are characterized as phenylethyl substituent group at the C2 position of the benzopyran ring, which displays a wide spectrum of pharmacological activity such as anticancer (Yang et al., 2018), anti-inflammatory (Wang et al., 2018), antioxidant (Ibrahim & Mohamed, 2015), and antibacterial activities (Lei et al., 2018). As an important heterocyclic skeleton, chromone also appeared in many α-glucosidase inhibitors (Dhameja & Gupta, 2019). In the past few years, some 2-(2-phenylethyl)chromones isolated from agarwood had been found to act as α-glucosidase inhibitors (Liao et al., 2016; Mi et al., 2021). Moreover, researchers have reported chromonyl enaminones as α-glucosidase inhibitor showed good inhibition of the α-glucosidase enzyme (Mendieta-Moctezuma et al., 2019).
Meanwhile, it is generally acknowledged that modifying the structural skeletons of natural products is an important approach to finding biologically active ingredients (Guo, 2017). Based on the multiple pharmacological activities of PEC, and some PECs have been found to exhibit α-glucosidase inhibitory activity. So that PECs core is a privileged scaffold for the design and development of new α-glucosidase inhibitors. In this present study, we reported the design, synthesis, and α-glucosidase inhibitory activity studies of new (2-phenyethyl)chromone derivatives, based on the compound skeleton of the active ingredient of the natural source (Fig. 1). Furthermore, the interaction between the optimization compounds and α-glucosidase was investigated by kinetic analysis, circular dichroism (CD) spectroscopy, fluorescence quenching, and molecular docking. In vivo inhibition of sucrase in mice was researched by oral loading tests with sucrose. The ADME (Absorption, distribution, metabolism and excretion) properties have been measured for all target compounds by using the Molinspiration online property calculation toolkit.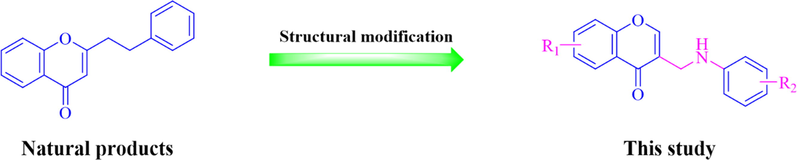
Rationale design of the target compounds of this study.
2 Results and discussion
2.1 Chemistry
The synthetic pathway of the title compounds 3–34 was shown in Scheme 1. The starting materials 1 was treated with POCl3 in the presence of dimethylformamide solution to obtain intermediates 2 via Vilsmeier-Haack reaction (Xiang et al., 2012). Then, intermediates 2 condensed with various anilines in dichloromethane in the presence of Na(AcO)3BH. After the completion of the reaction, the mixture was washed with sodium bicarbonate and purified by chromatography on silica gel to afford the corresponding (2-phenyethyl)chromones derivatives 3–34 (McGonagle et al., 2013). The chemical structures of all the target products 3–34 were characterized by spectroscopic techniques such as 1H NMR, 13C NMR, and HRMS (Supplementary material, Fig. S1-S96). For instance, the 1H NMR spectrum of 3 showed a doublet peak at δH 4.09 ppm with a coupling constant of 5.6 Hz representing the methylene. The proton of the –NH appeared at δH 6.04 ppm with a coupling constant of 5.6 Hz as a triplet peak. Two doublet peaks at δH 6.64 ppm and δH 7.24 ppm with a coupling constant of 8.8 Hz were attributed to the benzene ring protons of C2′,6′-H and C3′,5′-H, respectively. The protons of the C2-H of the chromone ring were observed as a singlet at δH 8.15 ppm. Two doublet signals at δH 7.97 ppm (J = 8.8 Hz) and δH 6.87 ppm (J = 2.0 Hz) and a doublet–doublet signal at δH 6.98 ppm (J = 8.8 Hz, 2.0 Hz) were attributed to C5-H, C8-H, and C6-H of chromone ring, respectively. The 13C NMR spectrum of 3 showed the methylene carbon appeared at δC 38.12 ppm. The characteristic peak of carbonyl carbon appeared at δC 175.51 ppm and the signal at δC 162.48 ppm was attributed to C7. The correlation signal of benzene ring carbons were observed to assign the C-2′,6′ and C-3′,5′ signal to δC 131.24 and 114.23 ppm, respectively. The remaining aromatic and benzopyran ring carbons resonate at δC 102.08–157.66 ppm. Furthermore, the mass spectrum of compound 3 showed a molecular ion peak at m/z 343.9934 as [M + H]- which also supported the proposed chemical structure of the product.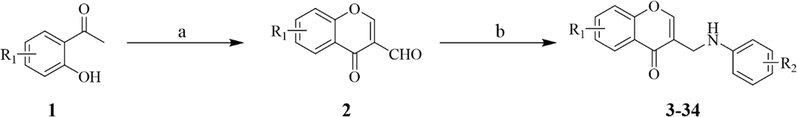
Reagents and conditions: (a) POCl3, dimethyl formamide, 50℃, 2 h; (b) different anilines (R2ArNH2), Na(AcO)3BH, CH2Cl2, rt, 0.5 h.
2.2 α-Glucosidase inhibitory activity and structure–activity relationships (SARs)
All the synthesized compounds 3–34 were screened for their α-glucosidase inhibitory activity by in vitro enzyme assay (Table 1). Some of the synthesized derivatives displayed inhibitory activity against α-glucosidase with IC50 values ranging from 11.72 ± 0.08 to 85.58 ± 2.30 μM. Acarbose as positive control (IC50 = 832.22 ± 2.00 μM), the value of IC50 is similar to prevenient literature report (Khan et al., 2014; Rahim et al., 2015; Taha et al., 2015). Among them, compound 4 showed the most active inhibitory activity against α-glucosidase with an IC50 of 11.72 ± 0.08 μM. From the SAR point of view (Fig. 2), compounds 3–18 showed higher inhibition compared with 19–34, which demonstrated the importance of OH at the C-7 position for the inhibitory activity. The introduction of electron-withdrawing groups such as –NO2, -Br, -Cl, -CF3, and -F on the phenyl ring enhances the inhibitory activity against α-glucosidase. Comparing 20 (IC50>100 μM), 25 (IC50>100 μM), and 26 (IC50 = 30.63 ± 0.19 μM), shows that substitution at para-position displayed the more potent inhibitory effect than other positions. Moreover, the introduction of the electron-donating group (–CH3) on the phenyl ring could enhance the inhibitory activity against α-glucosidase, while the oxygen-containing electron donor group (–OCH3 and –OCH2CH3) resulted in the reduction of the activity. In conclusion, these results show that there are differences in the biological activities of this type of compound due to the pattern of substitution between aniline and chromone. .
.
Compound
R1
R2
IC50 (μM)a,b
3
OH
4-Br
31.88 ± 0.85
4
OH
4-Cl
11.72 ± 0.08
5
OH
2-OCH2CH3
>100
6
OH
4-OCH2CH3
>100
7
OH
4-CF3
25.90 ± 1.05
8
OH
4-OCH3
85.58 ± 2.30
9
OH
H
85.37 ± 2.21
10
OH
4-OC6H5
75.33 ± 0.19
11
OH
4-CH3
22.62 ± 0.58
12
OH
3-CH3
60.47 ± 2.27
13
OH
2-CH3
65.06 ± 1.24
14
OH
3-OCH3
48.78 ± 1.71
15
OH
3-CF3
31.50 ± 3.64
16
OH
3, 4, 6-(OCH3)3
>100
17
OH
4-F
23.90 ± 1.00
18
OH
2, 4-(CH3)2
72.81 ± 2.54
19
H
4-CH3
>100
20
H
2-CF3
>100
21
H
2,4-(CH3)2
>100
22
H
4-F
>100
23
H
2-F
>100
24
H
H
>100
25
H
3-CF3
>100
26
H
4-CF3
30.63 ± 0.19
27
H
4-OCH2CH3
>100
28
H
2, 5-Cl2
59.93 ± 4.23
29
H
2-OCH2CH3
>100
30
H
4-NO2
44.45 ± 1.38
31
H
2-OCH3
>100
32
H
4-OH
>100
33
H
3-OH, 4-OCH3
30.63 ± 0.19
34
H
1-methyl-1H-indole
>100
Acarbose
–
–
832.22 ± 2.00
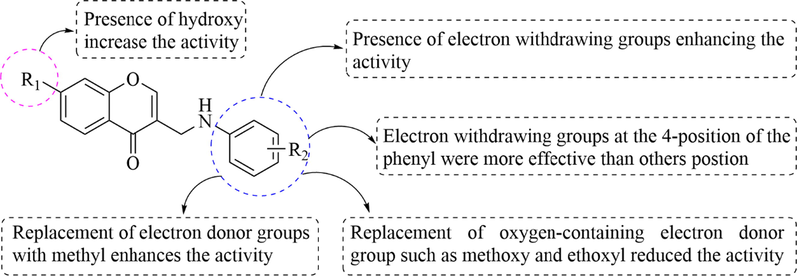
SARs analysis of this class of compounds.
2.3 Enzyme kinetic study
To further understand the inhibition mechanism, the most active compound 4 on α-glucosidase for the hydrolyzation of 4-nitrophenyl-β-d-galactopyranoside (pNPG) was studied. As shown in Fig. 3A, the lines of Lineweaver-Burk plot had a fixed intercept on the Y-intercept and X-slopes which increase with the concentration of the inhibitor. The values of Vmax remained the same and the values of Km increased with increasing concentration of the inhibitor, which suggested that compound 4 was a competitive inhibitor of α-glucosidase (Fig. 3B). Besides, the Ki value was determined as 1.88 × 10-6 M by using Dixon plots analysis of enzymatic reactions (El Ashry et al., 2022). The inhibitory mechanism is similar to previous article (Sun et al., 2015). The result revealed that compound 4 could only bind with the free enzyme.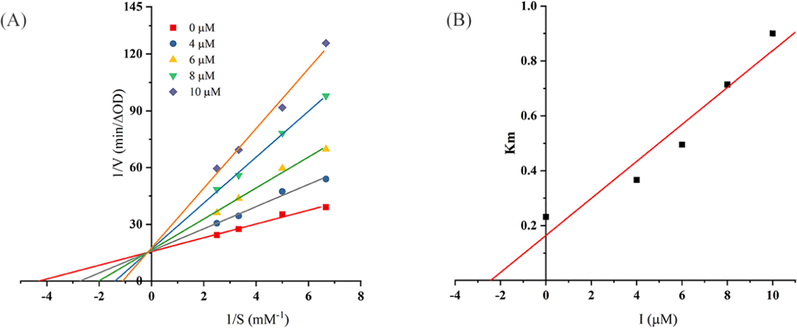
(A) Lineweaver-Burk plot for compound 4 (the concentrations of 4 were 0, 4, 6, 8, and 10 μM, and pNPG were 0.15, 0.2, 0.3, and 0.4 mM). (B) Secondary replot of slopes (Lineweaver-Burk plot) vs various concentrations of the compound.
2.4 Circular dichroism spectra
Circular dichroism (CD) is an excellent tool for researching the interactions between proteins and small molecules by analyzing the changes in the protein secondary structure (such as α-helix, β-sheet, β-turn, and random coil) and microenvironment (Zheng et al., 2020). The CD spectra of α-glucosidase with and without compound 4 were shown in Fig. 4. The CD spectrum of the α-glucosidase sample showed two negative peaks at 209 nm and 220 nm, which were the characteristic peaks of α-helix. Furthermore, compound 4 changed the intensity of two peaks, especially at peak of 209 nm by the change of concentration. Results of CD spectra indicated that the binding of 4 and α-glucosidase impelled a modification in the secondary structure of the enzyme. Besides, the calculation result shows that the α-helix decreased from 41.2 to 39.0 %. The parameter of β-sheet (%) showed slight changes increased from 15.5 % to 15.8 %, and β-turn (%) showed irregular changes (Table 2). These results suggested that the structure of α-glucosidase became looser and unstable (Wang et al., 2020). In addition, in molecular docking, it is shown that compound 4 linked α-glucosidase enzyme to form strong hydrophobic binding, which emphasized that α-helix is the appropriate site for 4 binding in α-glucosidase (Jiang et al., 2014). In previous articles, malvidin-3-O-galactoside and diacylated anthocyanins also show a similar mechanism of action (Xue et al., 2022; Yang et al., 2021).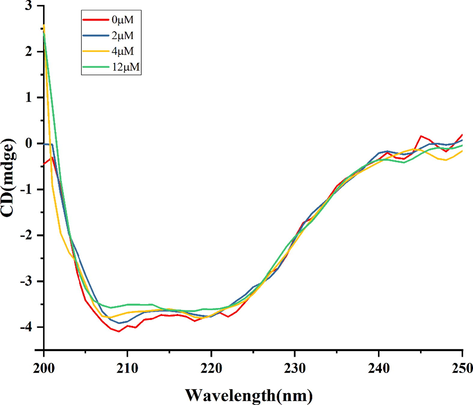
The circular dichroism spectra of 4. The concentration of α-glucosidase was 4 μM and its mixtures with different concentrations of 4 (0, 2, 4, 12 μM).
4 (μM)
α-Helix (%)
β-Sheet (%)
β-Turn (%)
Random coil (%)
0
41.2
15.5
14.9
28.0
2
39.4
15.8
14.2
29.1
4
39.3
15.8
13.7
29.3
12
39.0
15.8
14.3
29.4
2.5 Fluorescence quenching
The fluorescence spectrum can be used to research the interaction between biological macromolecules protein and small molecules (Nan et al., 2022). In order to study the interaction between the 4 and α-glucosidase, the fluorescence quenching experiment of α-glucosidase with 4 at various concentrations was conducted. According to Fig. 5A, the fluorescence intensity of α-glucosidase decreased with the increased concentrations of compound 4, indicating it was able to change the polarity and conformation of α-glucosidase. Regarding the addition of compound 4, the blue shift toward a lower wavelength is attributed to descending microenvironment polarity and increasing hydrophobicity of the tryptophan residue (Nan et al., 2022). To further understand the quenching mechanism of compound 4 on α-glucosidase, the quenching rate constant (Ksv) was calculated using the Stern-Volmer equation (Han et al., 2022).
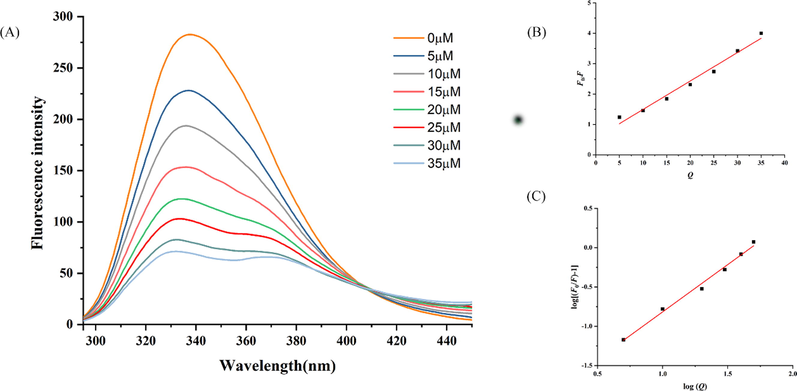
(A) Fluorescence emission spectra of protein α-glucosidase at different concentrations of 4, T = 298 K, α-glucosidase = 3.5 U/mL. (B) The Stern-Volmer plots of 4 and α-glucosidase. (C) The double logarithm regression plots of 4 and α-glucosidase.
F0 stands for the fluorescence intensity without a quencher. F is the fluorescence intensity with the different concentrations of quencher. Kq denotes the quenching rate constant of the biomolecule. τ0(10-8 s) is the lifetime of the fluorophore in the absence of the quencher. [Q] is the concentration of quencher. Accordingly, a good linear relationship is observed in Fig. 5B which demonstrates the interactions that existed in a single quenching mechanism, static or dynamic. Moreover, according to Table 3, the value of Kq (92.3 × 1011 Lmol−1) is much>2.0 × 1010 Lmol−1. This result demonstrated that the ligand quenching type of compound 4 was a static quenching mechanism. Using the formula (II), the value of binding sites (n) and binding constant (Ka) with α-glucosidase were obtained (Fig. 5C).
Ksv (×103 Lmol−1)
Kq (×1011 Lmol−1)
Ka (×104 Lmol−1)
n
92.3
92.3
2.50
1.33
log(F-F)/F = nlog[Q] + log Ka (II).
The binding constant (Ka) and the numerical value of binding sites (n) of 4 were 2.500 × 104 L/mol and 1.33, respectively. The value of n was approximately equal to 1, which suggested just one binding site on the enzyme for 4 (Han et al., 2022).
2.6 Molecular docking
To further explain the interaction between compound 4 and α-glucosidase. The theoretical binding modes of compound 4 with α-glucosidase are shown in Fig. 6. Because the 3D structure of a-glucosidase is still unclear, the homology model was constructed using the known crystal structures of the Saccharomyces cerevisiae isomaltase (PDB ID: 3AJ7), according to our previous article (Wang et al., 2017). The binding energy of 4 with the enzyme was evaluated to be −7.7 kcal•mol−1. Compound 4 adopted a Y-shaped conformation in the pocket of the α-glucosidase. It was located at the hydrophobic site, surrounded by the residues Phe-157, Phe-177, Leu-218, Pro-240, and Phe-300, forming a strong hydrophobic binding. Detailed analysis showed that the 4-chlorophenyl group of 4 formed hydrogen bond acceptor with the residue Lys-155. In addition, the chromone scaffold of compound 4 and the residues Glu-276 and Asp-349 formed hydrogen bond donor, and formed π-π stacking interaction with the residue Phe-157 (El Ashry et al., 2022). In conclusion, the above molecular docking explained the interactions between 4 and α-glucosidase, which afforded more information for further development of α-glucosidase inhibitors.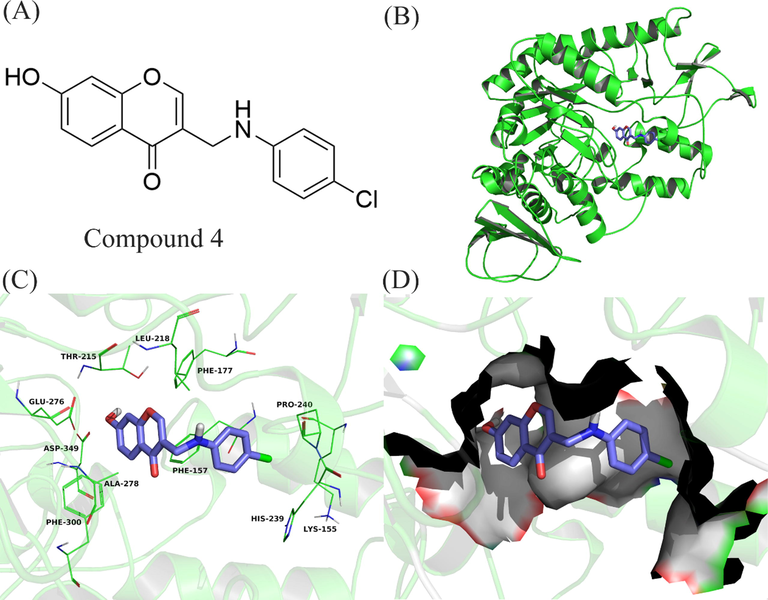
Compound 4 was docked to the binding pocket of the α-glucosidase. (A) The chemical structure of compound 4. (B) The overall structure of α-glucosidase with 4. (C) Binding pose of 4 at the binding site. (D) Binding pose of 4 in the surface of the binding pocket.
2.7 Effects of compound 4 on postprandial blood glucose in mice
The reduction of postprandial blood glucose by 4 in vivo was measured using the sucrose loading test, and the level of glucose was tested at 0, 15, 30, 60, 90, and 120 min, respectively. The results were presented in Fig. 7, acarbose served as the positive control. In the experiment, the blood glucose levels of the mice in all groups rose to a peak value at 15 min and returned to the pretreatment level at 120 min. It was displayed that the level of blood glucose of the blank group was no remarkable changes throughout the whole process. Furthermore, the groups of compound 4 and acarbose were significantly lower than the control group, which could be concluded that with oral administration at a 20 mg/kg dose, acarbose and 4 were able to reduce glucose uptake in normal mice fed with saccharose.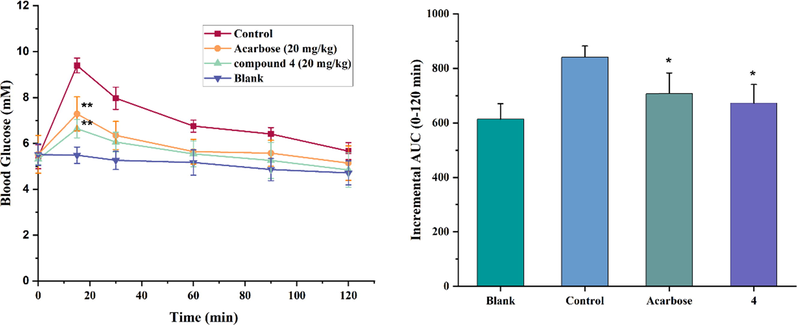
(A) Effect of 4 on blood glucose in Kunming mice. The animal assays were conducted with the administration of sucrose at a dose of 2.5 g/kg. The mice of each assay were orally administrated along with water (control group), 4 (20 mg/mL), and acarbose (20 mg/mL), respectively. Data are presented as the mean ± SEM (n = 7, **p < 0.01); (B) The inhibition of postprandial sucrose response by 4 in normal Kunming mic. The incremental AUC0-120 min in normal mice after sucrose administration. Asterisks indicate a significant difference (*p < 0.05,) compared with the control group.
2.8 In vitro cytotoxicity
In vitro cytotoxicity testing is one of the important methods to measure the toxicity of the drug. The toxicity of compound 4 (0, 25, 75, 125, and 675 μM) toward the growth of the human embryonic kidney (HEK 293) cell line was evaluated by the CCK-8 method (Wen et al., 2016). The result was shown that 4 exhibited low cytotoxicity against normal human cell lines with an IC50 value of 556.12 ± 6.42 μM, which was much higher than that of glucosidase inhibition (11.72 ± 0.08 μM). It was demonstrated that 4 exhibited an inhibitory effect on α-glucosidase at non-cytotoxic concentrations. Therefore, it could be concluded that 4 had a good safety drug for the application as an anti-diabetic agent.
2.9 ADME predictions
Optimization of absorption, distribution, metabolism, and excretion (ADME) profiles of biotherapeutics is critical in modern drug discovery. To measure the ADME properties of compounds 3–34, the pharmacokinetic parameters of all synthesized compounds were calculated by using the Molinspiration online property calculation toolkit (https://www.molinspiration.com/) and the results are presented in Table 4. According to the result, all the synthesized compounds 3–34 showed an excellent absorption range from 78.62 % to 94.43 %. The ADME properties of all the compounds have shown satisfactory results. The Lipinski rule of five states that an orally active drug has no more than one violation of the following criteria: molecular weight (MV) < 500, values of the octanol–water partition coefficient (log P) < 5, the number of hydrogen bond donor (NHD) < 5 and number of hydrogen bond acceptor (NHA) < 10 (Caron et al., 2018). From all these parameters, it can be predicted that among synthesized compounds, all compounds are likely to be orally active as they obeyed Lipinski’s rule of five.
Compd
%ABSa
TPSAb(A0)
n-rotbc
MWd
< 500MVe
miLogPf
< 5
n-OHNHg
< 10
n-ONh
< 1Lipinski’s violation
3
94.43
42.24
3
265.31
245.76
3.45
1
3
0
4
94.43
42.24
4
319.28
260.50
3.85
1
3
0
5
94.43
42.24
3
279.34
262.32
3.83
1
3
0
6
94.43
42.24
3
269.27
234.13
3.17
1
3
0
7
94.43
42.24
3
269.27
234.14
3.12
1
3
0
8
94.43
42.24
3
251.28
229.20
3.00
1
3
0
9
94.43
42.24
4
319.28
260.50
3.87
1
3
0
10
94.43
42.24
4
319.28
260.50
3.90
1
3
0
11
91.24
51.47
5
295.34
271.55
3.39
1
4
0
12
94.43
42.24
3
320.18
256.27
4.29
1
3
0
13
91.24
51.47
5
295.34
271.55
3.39
1
4
0
14
78.62
88.6
4
296.28
252.53
2.96
1
6
0
15
87.45
62.47
3
267.28
237.22
2.52
2
4
0
16
91.24
51.47
4
281.31
254.75
3.06
1
4
0
17
84.26
71.70
4
297.31
262.76
2.34
2
5
0
18
92.73
47.17
3
304.35
275.12
3.27
1
4
0
19
87.45
62.47
3
301.73
250.75
3.18
2
4
0
20
84.26
71.70
5
311.34
279.56
2.88
2
5
0
21
87.45
62.47
3
346.18
255.10
3.31
2
4
0
22
84.26
71.70
5
311.34
279.56
2.93
2
5
0
23
87.45
62.47
4
335.28
268.51
3.39
2
4
0
24
84.26
71.70
4
297.31
262.76
2.56
2
5
0
25
87.45
62.47
3
267.28
237.22
2.50
2
4
0
26
84.26
71.70
5
359.38
317.61
4.25
2
5
0
27
87.45
62.47
3
281.31
253.78
2.95
2
4
0
28
87.45
62.47
3
281.31
253.78
2.92
2
4
0
29
87.45
62.47
3
281.31
253.78
2.90
2
4
0
30
84.26
71.70
4
297.31
262.76
2.53
2
5
0
31
87.45
62.47
4
335.28
268.51
3.37
2
4
0
32
77.89
90.17
6
357.36
313.85
2.13
2
7
0
33
87.45
62.47
3
285.27
242.15
2.66
2
4
0
34
87.45
62.47
3
295.34
270.34
3.32
2
4
0
3 Conclusion
In this study, a series of (2-phenyethyl)chromone derivatives 3–34 were designed, synthesized, and screened for their α-glucosidase inhibitory activity. Among them, compound 4 (IC50 = 11.72 ± 0.08 μM) displayed the potent inhibitory activity against α-glucosidase. SAR studies indicate that the hydroxyl on the chromone ring would increase the α-glucosidase inhibitory activity. The kinetics of α-glucosidase inhibition were carried out for the most potent compound 4, and revealed competitive inhibitory mechanisms with Ki value of 1.88 × 10-6 M. However, the binding between α-glucosidase compound 4 changed the secondary structure of α-glucosidase, reflect on α-helix at 209 nm and 220 nm, based on the CD results. Fluorescence quenching revealed the binding constant (Ka) was 2.500 × 104 L/mol and suggested just one binding site on the enzyme for compound 4. Results from the molecular docking showed that Phe-157, Phe-177, Leu-218, Pro-240, Phe-300, Lys-155, Glu-276, and Asp-349 were the main residues of α-glucosidase, and their were the critical residues for the interaction between α-glucosidase and compound 4. The inhibitory effect of 4 on α-glucosidase in vivo was proved in the postprandial blood glucose experiment. The result of ADME predictions showed that all compounds obeyed Lipinski’s rule of five to become a “drug-like” molecule. Hence, this study indicated that compound 4 is a promising α-glucosidase inhibitor and can be used as a lead compound for the development of new antidiabetic drugs.
4 Experimental section.
4.1 Chemistry
All needful starting materials and reagents were supplied from commercial suppliers. TLC was performed on 0.20 mm Silica Gel 60 F254 plates (Qingdao Ocean Chemical Factory, Shandong, China). Nuclear magnetic resonance spectra (NMR) were recorded on a JEOL spectrometer (400 MHz) with TMS as an external reference and reported in parts per million. The Supplemental Materials contain sample 1H NMR, 13C NMR, and HRMS spectrum for target products 3–34.
4.1.1 General methods for the synthesis of 2
POCl3 (4.5 mL) was added to dimethylformamide (13 mL) at 50 ℃, the mixture was stirred for 2 h. This mixture was added to a solution of substituted acetophenone (10 mmol) in dimethylformamide (3.6 mL) and stirred at 58 °C for 2 h. The mixture was poured into ice water (300 g) and stirred at room temperature for 6 h. Finally, the white solid intermediate 2 was obtained by filtration and recrystallization from ethanol.
4.1.2 General methods for the synthesis of 3–34
To a solution of intermediate 2 (1.4 mmol) in CH2Cl2 (5 mL) was added Na(AcO)3BH (2.0 mmol) and substituted anilines 3 (1.4 mmol) and the mixture was stirred at room temperature for 0.5 h. After the completion of the reaction, the mixture was poured into 200 mL of dichloromethane and then washed with saturated sodium bicarbonate solution (100 mL × 2). The organic layers were dried over Na2SO4 and then concentrated under vacuum. The residue was filtered, dried, and purified by silica gel column (petroleum ether/ethyl acetate) to obtain the target compounds 3–34, respectively.
4.1.2.1 3-(((4-Bromophenyl)amino)methyl)-7-hydroxy-4H-chromen-4-one (3)
Yellow solid; Yield 69.73 %; mp: 221.3–218.5 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.74 (s, 1H), 8.15 (s, 1H), 7.97 (d, J = 8.8 Hz, 1H), 7.24 (d, J = 8.8 Hz, 2H), 6.98 (dd, J = 8.8, 2.4 Hz, 1H), 6.87 (d, J = 2.0 Hz, 1H), 6.64 (d, J = 8.8 Hz, 2H), 6.04 (t, J = 5.6 Hz, 1H), 4.09 (d, J = 5.6 Hz, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.51, 162.48, 157.66, 153.33, 147.44, 131.24, 126.56, 120.14, 116.03, 115.00, 114.23, 106.61, 102.08, 38.12; HRMS calcd for C16H11BrNO3- [M−H]-: m/z = 343.9928, found 343.9934.
4.1.2.2 3-(((4-Chlorophenyl)amino)methyl)-7-hydroxy-4H-chromen-4-one (4)
Yellow solid; Yield 12.4 %; mp: 205.8–211.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.82 (s, 1H), 8.13 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 7.08 (d, J = 8.0 Hz, 2H), 6.93 (d, J = 8.8 Hz, 1H), 6.83 (s, 1H), 6.62 (d, J = 8.0 Hz, 2H), 4.02 (s, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.54, 162.49, 157.67, 153.34, 147.08, 128.43, 126.57, 120.19, 119.30, 116.04, 115.01, 113.68, 102.09, 38.22; HRMS calcd for C16H11ClNO3- [M−H]-: m/z = 300.0433, found 300.0430.
4.1.2.3 3-(((2-Ethoxyphenyl)amino)methyl)-7-hydroxy-4H-chromen-4-one (5)
Yellow solid; Yield 12.6 %; mp: 178.8–191.5 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.83 (s, 1H), 8.13 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 6.92 (dd, J = 8.8, 2.0 Hz, 1H), 6.78–6.81 (m, 2H), 6.74 (t, J = 7.6 Hz, 1H), 6.60 (dd, J = 7.6, 1.2 Hz, 1H), 6.54 (td, J = 7.6, 1.2 Hz, 1H), 5.20 (t, J = 6.4 Hz, 1H), 4.12 (d, J = 6.4 Hz, 2H), 3.99 (q, J = 7.2 Hz, 2H), 1.33 (t, J = 6.8 Hz, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.50, 163.16, 158.34, 153.91, 146.53, 138.29, 127.23, 121.63, 121.06, 116.81, 115.68, 111.95, 110.68, 102.77, 64.20, 39.24, 15.30; HRMS calcd for C18H16NO4- [M−H]-: m/z = 310.1085, found 310.1082.
4.1.2.4 3-(((4-Ethoxyphenyl)amino)methyl)-7-hydroxy-4H-chromen-4-one (6)
Yellow solid; Yield 12.3 %; mp: 184.9–200.1 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.76 (s, 1H), 8.08 (s, 1H), 7.91 (dd, J = 8.8, 2.8 Hz, 1H), 6.91 (dt, J = 8.8, 2.4 Hz, 1H), 6.81 (t, J = 2.4 Hz, 1H), 6.69 (dd, J = 9.2, 2.4 Hz, 2H), 6.55 (dd, J = 8.8, 2.4 Hz, 2H), 5.38 (s, 1H), 3.99 (d, J = 4.4 Hz, 2H), 3.86 (qd, J = 6.8, 2.4 Hz, 2H), 1.25 (td, J = 6.8, 2.4 Hz, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.66, 162.43, 157.64, 153.18, 150.15, 142.27, 126.55, 120.69, 116.04, 115.37, 114.95, 113.53, 102.06, 63.27, 14.70; HRMS calcd for C18H16NO4- [M−H]-: m/z = 310.1085, found 310.1085.
4.1.2.5 7-Hydroxy-3-(((4-(trifluoromethyl)phenyl)amino)methyl)-4H-chromen-4-one (7)
Yellow solid; Yield 16.0 %; mp: 195.9–206.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.70 (s, 1H), 8.14 (s, 1H), 7.93 (dd, J = 8.8, 2.0 Hz, 1H), 7.37 (d, J = 7.6 Hz, 2H), 6.93 (dt, J = 10.4, 2.0 Hz, 1H), 6.83 (s, 1H), 6.74 (d, J = 8.0 Hz, 2H), 6.52 (d, J = 5.2 Hz, 1H), 4.11 (d, J = 5.2 Hz, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.67, 162.70, 157.85, 153.74, 151.30, 126.78, 126.28, 126.25, 124.04, 119.98, 116.16, 115.70, 115.39, 115.23, 111.65, 102.27, 37.86; HRMS calcd for C17H11F3NO3- [M−H]-: m/z = 334.0697, found 334.0692.
4.1.2.6 7-Hydroxy-3-(((4-methoxyphenyl)amino)methyl)-4H-chromen-4-one (8)
Yellow solid; Yield 16.3 %; mp: 171.8–176.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.62 (s, 1H), 8.06 (s, 1H), 7.91 (d, J = 8.4 Hz, 1H), 6.91 (dd, J = 8.4, 2.0 Hz, 1H), 6.80 (d, J = 1.2 Hz, 1H), 6.70 (d, J = 8.8 Hz, 2H), 6.58 (d, J = 9.2 Hz, 2H), 5.26 (s, 1H), 4.01 (d, J = 2.4 Hz, 2H), 3.63 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.67, 162.44, 157.65, 153.19, 151.01, 142.29, 126.56, 120.68, 116.05, 114.96, 114.58, 113.56, 102.07, 55.23; HRMS calcd for C17H14NO4- [M−H]-: m/z = 296.0928, found 296.0923.
4.1.2.7 7-Hydroxy-3-((phenylamino)methyl)-4H-chromen-4-one (9)
Yellow solid; Yield 7.8 %; mp: 210.1–218.1 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.67 (s, 1H), 8.09 (s, 1H), 7.92 (dd, J = 8.8, 2.0 Hz, 1H), 7.04–7.08 (m, 2H), 6.92 (dt, J = 8.8, 2.8 Hz, 1H), 6.81 (t, J = 2.4 Hz, 1H), 6.61 (d, J = 6.8 Hz, 2H), 6.54 (t, J = 7.6 Hz, 1H), 5.71 (s, 1H), 4.01 (s, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.81, 162.62, 157.81, 153.51, 148.27, 128.95, 126.77, 120.55, 116.13, 115.16, 112.41, 102.23, 38.29; HRMS calcd for C16H12NO3- [M−H]-: m/z = 266.0823, found 266.0820.
4.1.2.8 7-Hydroxy-3-(((4-phenoxyphenyl)amino)methyl)-4H-chromen-4-one (10)
Yellow solid; Yield 10.6 %; mp: 181.8–188.0 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.82 (s, 1H), 8.16 (s, 1H), 7.93 (d, J = 8.8 Hz, 1H), 7.27–7.31 (m, 2H), 7.00 (t, J = 7.2 Hz, 1H), 6.93 (dd, J = 8.8, 2.4 Hz, 1H), 6.82–6.86 (m, 5H), 6.66 (d, J = 8.8, Hz, 2H), 5.86 (s, 1H), 4.04 (d, J = 4.8 Hz, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.60, 162.46, 158.64, 157.67, 153.28, 146.02, 145.03, 129.51, 126.57, 121.74, 120.69, 120.50, 116.53, 116.07, 114.99, 113.30, 102.09, 38.67; HRMS calcd for C22H16NO4- [M−H]-: m/z = 358.1085, found 358.1080.
4.1.2.9 7-Hydroxy-3-((p-tolylamino)methyl)-4H-chromen-4-one (11)
Yellow solid; Yield 14.1 %; mp: 129.9–133.5 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.69 (s, 1H), 8.08 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 6.91–6.94 (m, 3H), 6.81 (d, J = 2.0 Hz, 1H), 6.60 (d, J = 8.4 Hz, 2H), 4.05 (s, 2H), 2.15 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.59, 162.56, 157.67, 153.56, 144.64, 129.93, 129.30, 126.55, 125.86, 121.65, 120.04, 115.98, 115.05, 113.53, 102.11, 38.89, 19.92; HRMS calcd for C17H14NO3- [M−H]-: m/z = 280.0979, found 280.0975.
4.1.2.10 7-Hydroxy-3-((m-tolylamino)methyl)-4H-chromen-4-one (12)
Yellow solid; Yield 17.1 %; mp: 191.9–203.0 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.67 (s, 1H), 8.07 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 6.90–6.96 (m, 2H), 6.81 (d, J = 2.0 Hz, 1H), 6.36–6.44 (m, 3H), 5.61 (s, 1H), 4.04 (d, J = 5.2 Hz, 2H), 2.17 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.61, 162.44, 157.65, 153.15, 148.14, 137.74, 128.61, 126.55, 120.56, 117.02, 116.04, 114.96, 113.03, 109.59, 102.06, 38.26, 21.14; HRMS calcd for C17H14NO3- [M−H]-: m/z = 280.0979, found 280.0975.
4.1.2.11 7-Hydroxy-3-((o-tolylamino)methyl)-4H-chromen-4-one (13)
Yellow solid; Yield 14.2 %; mp: 189.9–197.1 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.67 (s, 1H), 8.04 (s, 1H), 7.93 (d, J = 8.8 Hz, 1H), 6.91–6.97 (m, 3H), 6.80 (d, J = 2.4 Hz, 1H), 6.49–6.54 (m, 2H), 5.10 (t, J = 5.6 Hz, 1H), 4.15 (d, J = 5.2 Hz, 2H), 2.10 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.50, 163.14, 158.33, 153.92, 146.31, 130.36, 127.28, 127.24, 122.72, 121.02, 116.84, 116.68, 115.67, 110.26, 102.74, 39.19, 17.97; HRMS calcd for C17H14NO3- [M−H]-:280.0979, found 280.0976.
4.1.2.12 7-Hydroxy-3-(((3-methoxyphenyl)amino)methyl)-4H-chromen-4-one (14)
Yellow solid;Yield12.0 %; mp: 183.1–185.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.67 (s, 1H), 8.09 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 6.90–6.97 (m, 2H), 6.81 (d, J = 2.0 Hz, 1H), 6.12–6.23 (m, 3H), 5.74 (t, J = 6.0 Hz, 1H), 4.04 (d, J = 5.6 Hz, 2H), 3.65 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.56, 162.45, 160.26, 157.65, 153.29, 149.52, 129.45, 126.56, 120.54, 116.04, 114.98, 105.35, 102.08, 101.72, 98.19, 54.53, 38.16; HRMS calcd for C17H14NO4- [M−H]-: m/z = 296.0928, found 296.0924.
4.1.2.13 7-Hydroxy-3-(((3-(trifluoromethyl)phenyl)amino)methyl)-4H-chromen-4-one (15)
Yellow solid; Yield 10.9 %; mp: 188.6–191.4 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 8.17 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 7.27 (t, J = 7.6 Hz, 1H), 6.82–6.95 (m, 7H), 4.10 (s, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.47, 162.64, 157.68, 153.60, 148.43, 126.54, 126.54, 119.94, 115.99, 115.69, 115.11, 112.08, 112.04, 108.52, 102.13, 38.11; HRMS calcd for C17H11F3NO3- [M−H]-:334.0697, found 334.0694.
4.1.2.14 7-Hydroxy-3-(((3,4,5-trimethoxyphenyl)amino)methyl)-4H-chromen-4-one (16)
Yellow solid; Yield 6.7 %; mp: 184.4–191.7 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.64 (s, 1H), 8.11 (s, 1H), 7.88 (d, J = 8.8 Hz, 1H), 6.88 (dd, J = 8.8, 2.0 Hz, 1H), 6.78 (d, J = 2.4 Hz, 1H), 5.91 (s, 2H), 5.49 (s, 1H), 4.00 (d, J = 4.4 Hz, 2H), 3.65 (s, 6H), 3.49 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.59, 162.46, 157.64, 153.42, 153.36, 144.82, 129.04, 126.57, 120.79, 116.07, 115.00, 102.10, 90.53, 59.99, 55.53, 38.39; HRMS calcd for C19H18NO6- [M−H]-: m/z = 356.1140 found 356.1137.
4.1.2.15 3-(((4-Fluorophenyl)amino)methyl)-7-hydroxy-4H-chromen-4-one (17)
Yellow solid; Yield 5.1 %; mp: 205.1–216.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.62 (s, 1H), 8.08 (s, 1H), 7.91 (d, J = 8.8 Hz, 1H), 6.87–6.93 (m, 3H), 6.81 (d, J = 2.0 Hz, 1H), 6.59–6.63 (m, 2H), 5.62 (t, J = 6.0 Hz, 1H), 4.03 (d, J = 5.6 Hz, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.61, 155.95, 154.23, 148.73, 140.97, 134.17, 125.44, 124.97, 123.22, 121.48, 118.49, 115.71, 113.92, 38.90; HRMS calcd for C16H11FNO3- [M−H]-: m/z = 284.0728, found 284.0725.
4.1.2.16 3-(((2,4-Dimethylphenyl)amino)methyl)-7-hydroxy-4H-chromen-4-one (18)
Yellow solid; Yield 18.7 %; mp: 180.5–183.7 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.67 (s, 1H), 8.02 (s, 1H), 7.92 (d, J = 8.8 Hz, 1H), 6.92 (dd, J = 8.8, 2.4 Hz, 1H), 6.77–6.80 (m, 3H), 6.44 (d, J = 8.0 Hz, 1H), 4.91 (s, 1H), 4.11 (d, J = 2.4 Hz, 2H), 2.12 (s, 3H), 2.07 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.86, 162.45, 157.63, 153.20, 143.33, 130.53, 126.82, 126.54, 124.56, 122.18, 120.43, 116.00, 114.97, 109.91, 102.05, 19.84, 17.21; HRMS calcd for C18H16NO3- [M−H]-: m/z = 294.1136, found 294.1133.
4.1.2.17 3-((p-Tolylamino)methyl)-4H-chromen-4-one (19)
Yellow solid; Yield 48.44 %; mp: 157.6–160.1 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 8.24 (s, 1H), 8.09 (dd, J = 8.0, 1.2 Hz, 1H), 7.80 (td, J = 7.2, 1.6 Hz, 1H), 7.63 (d, J = 8.8 Hz, 1H), 7.49 (td, J = 8.0, 0.8 Hz, 1H), 6.88 (d, J = 8.0 Hz, 2H), 6.53 (d, J = 8.0 Hz, 2H), 5.66 (t, J = 6.0 Hz, 1H), 4.06 (d, J = 6.0 Hz, 2H), 2.12 (s, 3H); 13C NMR (100 MHz; DMSO‑d6) δ: 176.57, 155.95, 154.27, 145.94, 134.20, 129.42, 125.46, 124.97, 124.61, 123.21, 121.25, 118.49, 112.61, 38.62, 20.11; HRMS calcd for C17H14NO2- [M−H]-: m/z = 264.1030, found 264.1031.
4.1.2.18 3-(((2-(Trifluoromethyl)phenyl)amino)methyl)-4H-chromen-4-one (20)
Yellow solid; Yield 38.8 %; mp: 116.8–119.1 °C; 1H NMR (400 MHz, CDCl3) δ: 8.26 (dd, J = 8.0, 1.2 Hz, 1H), 7.87 (s, 1H), 7.68 (td, J = 7.2, 2.0 Hz, 1H), 7.41–7.48 (m, 3H), 7.35 (t, J = 7.2 Hz, 1H), 6.74–6.79 (m, 2H), 4.40 (s, 2H); 13C NMR (100 MHz, CDCl3) δ: 178.17, 157.56, 156.66, 155.22, 153.34, 143.69, 133.95, 125.78, 125.38, 123.91, 121.08, 118.35, 116.04, 115.82,114.78, 114.70, 41.55; HRMS calcd for C17H11F3NO2- [M−H]-: m/z = 318.0747, found 318.0745.
4.1.2.19 3-(((2,4-Dimethylphenyl)amino)methyl)-4H-chromen-4-one (21)
Yellow solid; Yield 27.4 %; mp: 108.3–116.5 °C; 1H NMR (400 MHz, CDCl3) δ: 8.23 (dd, J = 8.0, 1.6 Hz, 1H), 7.91 (s, 1H), 7.66 (td, J = 6.8, 1.6 Hz, 1H), 7.39–7.44 (m, 2H), 6.90 (d, J = 5.6 Hz, 2H), 6.57 (d, J = 8.8 Hz, 1H), 4.31 (s, 2H), 2.22 (s, 3H), 2.18 (s, 3H); 13C NMR (100 MHz, CDCl3) δ: 178.24, 156.66, 153.42, 142.89, 133.82, 131.38, 127.45, 127.28, 125.79, 125.27, 123.96, 123.37, 121.25, 118.32, 110.97, 40.99, 20.49, 17.63; HRMS calcd for C18H17NNaO2+ [M + Na]+: m/z = 302.1151, found 302.1151.
4.1.2.20 3-(((4-fluorophenyl)amino)methyl)-4H-chromen-4-one (22)
Brown solid; Yield 29.5 %; mp: 167.0–170.7 °C; 1H NMR (400 MHz, CDCl3) δ: 8.23 (dd, J = 7.6, 1.2 Hz, 1H), 7.92 (s, 1H), 7.67 (td, J = 7.2, 1.6 Hz, 1H), 7.40–7.45 (m, 2H), 6.86–6.91 (m, 2H), 6.60–6.63 (m, 2H), 4.23 (s, 2H); 13C NMR (100 MHz, CDCl3) δ: 177.87, 156.66, 153.28, 144.52, 133.99, 133.37, 126.96, 126.91, 125.85, 125.43, 123.79, 120.45, 118.34, 117.31, 112.70, 40.05; HRMS calcd for C16H11FNO2- [M−H]-: m/z = 268.0779, found 268.0779.
4.1.2.21 3-(((2-Fluorophenyl)amino)methyl)-4H-chromen-4-one (23)
Brown solid; Yield 19.7 %; mp: 110.2–111.8 °C; 1H NMR (400 MHz, CDCl3) δ: 8.25 (dd, J = 8.4, 1.6 Hz, 1H), 7.93 (s, 1H), 7.67 (td, J = 7.2, 1.6 Hz, 1H), 7.40–7.45 (m, 2H), 7.96–7.01 (m, 2H), 6.67–6.75 (m, 2H), 4.33 (s, 2H); 13C NMR (100 MHz, CDCl3) δ: 177.90, 156.63, 153.32, 135.64, 135.54, 133.89, 125.81, 125.34, 124.74, 124.71, 123.84, 120.92, 118.30, 117.97, 117.90, 114.96, 114.78, 113.10, 40.25; HRMS calcd for C16H11FNO2- [M−H]-: m/z = 268.0779, found 268.0780.
4.1.2.22 3-((Phenylamino)methyl)-4H-chromen-4-one (24)
Yellow solid; Yield 46.8 %; mp: 125.5–127.6 °C; 1H NMR (400 MHz, CDCl3) δ: 8.23 (dd, J = 8.0, 1.6 Hz, 1H), 7.93 (s, 1H), 7.67 (td, J = 6.8, 1.6 Hz, 1H), 7.48–7.37 (m, 2H), 7.24–7.14 (m, 2H), 6.77 (t, J = 7.2 Hz, 1H), 6.71 (d, J = 7.2 Hz, 2H), 4.29 (d, J = 0.8 Hz, 2H); 13C NMR (100 MHz, CDCl3) δ: 178.18, 156.65, 153.52, 146.85, 133.91, 129.51, 125.76, 125.34, 123.89, 120.94, 118.85, 118.33, 114.06, 41.06; HRMS calcd for C16H14NO2+ [M + H]+: m/z = 252.1019, found 252.1029.
4.1.2.23 3-(((3-(Trifluoromethyl)phenyl)amino)methyl)-4H-chromen-4-one (25)
Yellow solid; Yield 58.4 %; mp: 159.8–162.1 °C; 1H NMR (400 MHz, CDCl3) δ: 8.25 (dd, J = 8.0, 1.6 Hz, 1H), 7.93 (s, 1H), 7.68 (td, J = 6.8, 1.6 Hz, 1H), 7.41–7.46 (m, 2H), 7.26 (t, J = 6.8 Hz, 1H), 6.97 (d, J = 7.6 Hz, 1H), 6.89 (s, 1H), 6.82 (dd, J = 8.4, 2.4 Hz, 1H), 4.29 (s, 2H); 13C NMR (100 MHz, CDCl3) δ: 178.12, 156.66, 153.24, 147.77, 134.04, 131.88, 131.57, 129.89, 125.78, 125.45, 123.91, 123.00, 120.77, 118.38, 116.50, 114.64, 114.60, 109.68, 109.64, 40.53; HRMS calcd for C17H11F3NO2- [M−H]-: m/z = 318.0747, found 318.0747.
4.1.2.24 3-(((4-(Trifluoromethyl)phenyl)amino)methyl)-4H-chromen-4-one (26)
Brown solid; Yield 29.5 %; mp: 190.6–201.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 8.29 (s, 1H), 8.06 (dd, J = 8.0, 1.6 Hz, 1H), 7.75–7.80 (m, 1H), 7.61 (d, J = 8.4 Hz, 1H), 7.47 (t, J = 7.6 Hz, 1H), 7.33 (d, J = 8.4 Hz, 2H), 6.70 (d, J = 8.4 Hz, 2H), 6.64 (t, J = 5.4 Hz, 1H), 4.10 (d, J = 5.2 Hz, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.43, 155.99, 154.53, 151.27, 134.30, 126.29, 126.25, 125.55, 124.98, 123.24, 120.55, 118.51, 115.77, 115.46, 111.67, 37.92; HRMS calcd for C17H11F3NO2- [M−H]-: m/z = 318.0747, found 318.0746.
4.1.2.25 3-(((4-Ethoxyphenyl)amino)methyl)-4H-chromen-4-one (27)
Yellow solid; Yield 24.0 %; mp: 141.5–144.1 °C; 1H NMR (400 MHz, CDCl3) δ: 8.23 (dd, J = 8.0, 1.6 Hz, 1H), 7.90 (s, 1H), 7.66 (td, J = 6.8, 1.6 Hz, 1H), 7.39–7.44 (m, 2H), 6.76 (d, J = 8.8 Hz, 2H), 6.61 (d, J = 9.2 Hz, 2H), 4.21 (d, J = 0.8 Hz, 2H), 3.94 (q, J = 6.8 Hz, 2H), 1.36 (t, J = 7.2 Hz, 3H); 13C NMR (100 MHz, CDCl3) δ: 178.06, 156.51, 153.18, 151.86, 141.39, 133.68, 125.64, 125.12, 123.82, 121.36, 118.18, 115.65, 114.98, 63.96, 41.60, 14.99; HRMS calcd for C18H16NO3- [M−H]-: m/z = 294.1136, found 294.1137.
4.1.2.26 3-(((2,5-Dichlorophenyl)amino)methyl)-4H-chromen-4-one (28)
Yellow solid; Yield 80.1 %; mp: 134.9–138.9 °C; 1H NMR (400 MHz, CDCl3) δ: 8.25 (dd, J = 8.0, 1.6 Hz, 1H), 7.87 (s, 1H), 7.68 (td, J = 7.2, 1.6 Hz, 1H), 7.40–7.46 (m, 2H), 7.21–7.16 (d, J = 9.2, Hz, 1H), 6.60–6.63(m, 2H), 4.31 (d, J = 1.2 Hz, 2H); 13C NMR (100 MHz, CDCl3) δ: 177.70, 156.54, 152.94, 144.17, 133.90, 133.62, 129.98, 125.74, 125.33, 123.72, 120.28, 118.22, 117.86, 117.64, 111.30, 39.90; HRMS calcd for C16H12Cl2NO2+ [M + H]+: m/z = 320.0240, found 320.0240.
4.1.2.27 3-(((2-Ethoxyphenyl)amino)methyl)-4H-chromen-4-one (29)
Yellow solid; Yield 56.8 %; mp: 94.5–96.6 °C; 1H NMR (400 MHz, CDCl3) δ: 8.25 (dd, J = 7.6, 1.6 Hz, 1H), 7.88 (s, 1H), 7.66 (td, J = 7.2, 1.6 Hz, 1H), 7.39–7.44 (m, 2H), 6.77–6.84 (m, 2H), 6.67 (td, J = 7.6, 1.2 Hz, 1H), 6.58 (dd, J = 8.0, 1.6 Hz, 1H), 4.34 (d, J = 0.4 Hz, 2H), 4.08 (q, J = 6.8 Hz, 2H), 1.45 (t, J = 6.8 Hz, 3H); 13C NMR (100 MHz, CDCl3) δ: 177.84, 156.55, 153.18, 146.41, 137.31, 133.56, 125.67, 125.03, 123.77, 121.35, 121.11, 118.16, 117.18, 110.54, 110.34, 63.81, 39.85, 15.00; HRMS calcd for C18H18NO3+ [M + H]+: m/z = 296.1281, found 296.1294.
4.1.2.28 3-(((4-Nitrophenyl)amino)methyl)-4H-chromen-4-one (30)
Brown solid; Yield 27.9 %; mp: 182.5–198.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 10.45 (s, 1H), 8.78 (s, 1H), 8.09 (dd, J = 8.0, 1.6 Hz, 1H), 7.93 (s, 1H), 7.79 (td, J = 7.2, 1.6 Hz, 1H), 7.67 (d, J = 8.4 Hz, 1H), 7.49 (td, J = 8.0, 1.6 Hz, 1H), 7.17 (t, J = 8.0 Hz, 2H), 7.02 (d, J = 7.6 Hz, 2H), 6.71 (t, J = 7.2 Hz, 1H); 13C NMR (100 MHz, DMSO‑d6) δ: 175.54, 156.28, 152.82, 145.53, 134.84, 129.63, 128.38, 126.24, 125.73, 123.76, 120.10, 119.47, 119.20, 112.55, 31.68; HRMS calcd for C16H12KN2O4+ [M + K]+: m/z = 335.0429, found 335.0483.
4.1.2.29 3-(((4-Hydroxyphenyl)amino)methyl)-4H-chromen-4-one (31)
Yellow solid; Yield 31.0 %; mp: 168.5–174.9 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 8.43 (s, 1H), 8.19 (s, 1H), 8.04 (dd, J = 8.0, 1.6 Hz, 1H), 7.76 (td, J = 7.2, 1.6 Hz, 1H), 7.59 (d, J = 8.4 Hz, 1H), 7.45 (td, J = 8.0, 0.8 Hz,1H), 6.50 (d, J = 8.8 Hz, 2H), 6.44 (d, J = 9.2 Hz, 2H), 5.21 (t, J = 6.0 Hz, 1H) 3.97 (d, J = 5.6 Hz, 2H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.60, 155.94, 154.21, 153.65, 148.74, 140.96, 134.14, 125.42, 124.96, 124.10, 123.21, 121.48, 118.48, 115.71, 113.92, 55.36; HRMS calcd for C16H12NO3- [M−H]-: m/z = 266.0823, found 266.0818.
4.1.2.30 3-(((4-Methoxyphenyl)amino)methyl)-4H-chromen-4-one (32)
Yellow solid; Yield 35.6 %; mp: 151.5–155.3 °C; 1H NMR (400 MHz, DMSO‑d6) δ 8.26 (s, 1H), 8.09 (dd, J = 7.6, 1.6 Hz, 1H), 7.81 (td, J = 7.2, 1.6 Hz, 1H), 7.64 (d, J = 8.8 Hz, 1H), 7.50 (t, J = 7.2, 0.8 Hz, 1H), 6.71 (d, J = 8.8 Hz, 2H), 6.57 (d, J = 9.2 Hz, 2H), 5.49 (t, J = 6.4 Hz, 1H), 4.05 (d, J = 5.6 Hz, 2H), 3.62 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.60, 155.96, 154.27, 151.05, 142.34, 134.21, 125.47, 124.98, 123.21, 121.34, 118.49, 114.63, 113.62, 55.27; HRMS calcd for C17H16NO3+ [M + H]+: m/z = 282.1125, found 282.1135.
4.1.2.31 3-(((3-Hydroxy-4-methoxyphenyl)amino)methyl)-4H-chromen-4-one (33)
Brown solid; Yield 64.4 %; mp: 151.4–156.1 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 8.26 (s, 1H), 8.10 (dd, J = 8.0, 1.6 Hz, 1H), 7.80 (td, J = 7.2, 1.6 Hz, 1H), 7.62 (d, J = 8.4 Hz, 1H), 7.50 (t, J = 7.6 Hz, 1H), 7.17 (d, J = 8.8 Hz, 1H), 7.11 (d, J = 2.8 Hz, 1H), 6.70–6.62 (m, 2H), 6.15 (s, 1H), 5.38 (t, J = 5.2 Hz, 1H), 4.12 (d, J = 0.8 Hz, 2H), 3.67 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.34, 155.96, 147.50, 134.57, 125.83, 125.01, 123.08, 118.62, 113.82, 56.40; HRMS calcd for C17H16NO4+ [M + H]+: m/z = found 298.1087.
4.1.2.32 3-(((1-Methyl-1H-indol-5-yl)amino)methyl)-4H-chromen-4-one (34)
Yellow solid; Yield 69.73 %; mp: 149.5–166.8 °C; 1H NMR (400 MHz, DMSO‑d6) δ: 8.26 (s, 1H), 8.10 (dd, J = 8.0, 1.6 Hz, 1H), 7.80 (ddd, J = 7.2, 1.6 Hz, 1H), 7.62 (d, J = 8.4 Hz, 1H), 7.50 (t, J = 7.6 Hz, 1H), 7.17 (d, J = 8.8 Hz, 1H), 7.11 (d, J = 2.8 Hz, 1H), 6.32–6.67 (m, 2H), 6.15 (s, 1H), 5.38 (d, J = 5.2 Hz, 1H), 4.11 (d, J = 4.8 Hz, 2H), 3.67 (s, 3H); 13C NMR (100 MHz, DMSO‑d6) δ: 176.70, 155.94, 154.19, 141.69, 134.14, 130.61, 129.05, 128.87, 125.42, 124.97, 123.22, 121.49, 118.47, 111.37, 110.06, 101.25, 99.11, 32.44; HRMS calcd for C19H17N2O2+ [M + H]+: m/z = 305.1285, found 305.1295.
4.2 α-Glucosidase inhibitory activities
The α-glucosidase inhibitory activities of these compounds were screened according to the previous methods with minor modifications, acarbose was used as a positive control (Ji et al., 2021; Wang et al., 2017a, 2017b; Xu et al., 2020). The α-glucosidase (750 U) was purchased by Sigma Aldrich (St. Louis, MO, USA). Different concentrations of the tested compounds or acarbose (10 μL) and 150 μL of α-glucosidase solutions (0.2 U/mL) were added to a 96-well plate and the mixture was incubated at 37.5 °C for 15 min. Then, 40 μL of pNPG (1.25 mM) was added to the mixture. After incubation at 37 °C for 30 min, the absorbance of released p-nitrophenyl was measured at 405 nm using a microplate reader (Varioskan LUX, Thermo, USA); The inhibition percentage was obtained by the formula: Inhibition (%) = (1-ΔA sample/ΔA control) / *100 %. Where ΔA sample and ΔA control represent the absorbance with and without the test compounds, respectively. The IC50 value was calculated by the linear with inhibition percentage and the concentrations of test compounds.
4.3 Enzyme kinetic study
The enzyme kinetics assays for α-glucosidase were performed using the method the above-mentioned. The substrate pNPG (0.15, 0.2, 0.3, and 0.4 mM) and compound 4 (0, 4, 6, 8, and 10 μM) were used in varying concentrations. The Michaelis constant (Km) and maximum velocity of the reaction (Vmax) were calculated by using the Lineweaver-Burk plot.
4.4 Circular dichroism spectra
The circular dichroism (CD) spectrum was measured from 190 to 260 nm by using Chirascan Plus (Applied Photophysics, Leatherhead, UK) at 25 °C. The α-glucosidase solution (4 μM) and its mixtures with different concentrations of 4 (0, 2, 4, and 12 μM) were measured. The CD spectra were recorded in the far UV region between 190 and 260 nm with a bandwidth of 1 nm on a Chirascan qCD. All collected CD spectra were subtracted from the PBS buffer signal. The CDNN 2.1 software was used to investigate the α-helix content change of α-glucosidase (Zhang et al., 2018).
4.5 Fluorescence quenching
Fluorescence spectra of samples were recorded on Cary Eclipse (Agilent Technologies) at 25 °C. Briefly, 1.5 uL of 4 stock solution (10 mM) was added seven times to the α-glucosidase solution (3.5 U/mL, 3 mL). The concentrations of 4 varied from 0 to 40 μM. In the following step, the spectra were recorded with the excitation wavelength of 280 nm and the emission wavelength ranging from 300 to 450 nm. Noteworthy, the fluorescence spectrum of PBS (pH = 6.8) was tested using the same method.
4.6 Molecular docking
Autodock Vina 1.1.2 was used for molecular docking to study the binding mode between compound 4 and α-glucosidase (Trott & Olson, 2010). The 3D structure of compound 4 was drawn by ChemBioDraw Ultra 14.0 and ChemBio3D Ultra 14.0. The AutoDockTools 1.5.6 package is used for generating the docking input files (David S. Goodsell, 1996; Morris et al., 2009; Sanner, 1999). The search grid of α-glucosidase was identified as center_x: −19.676, center_y: −7.243, and center_z: −21.469 with dimensions size_x: 15, size_y: 15, and size_z: 15. The value of exhaustiveness was set to 16. For Vina docking, the default parameters were used if it was not mentioned. The best-scoring pose as judged by the Vina docking score was chosen and visually analyzed using PyMOL 1.7.6 software (https://www.pymol.org);
4.7 Effects of 4 on postprandial blood glucose
To further study the potency of compound 4 to the reduction of postprandial blood glucose levels in vivo, the saccharose loading test in normal Kunming mice was performed by using the modifying previous reported method (Huang et al., 2020a, 2020b; Ji et al., 2021; Yu et al., 2020). The Kunming (25–30 g and eight-week-old) mice were purchased from Guizhou Medical University (Guiyang, China). All mice for experiments were fasted for 12 h but had free access to water, before treatment. In this experiment, all mice were randomly divided into four groups with 7 animals in each group. Gave different drugs or blank solutions to the four groups: the control group (0.5 % CMC-Na solution), acarbose group (20 mg/kg, dissolved in 0.5 % CMC-Na solution), compound 4 treated group (20 mg/kg, dissolved in 0.5 % CMC-Na solution), and blank group (0.5 % CMC-Na solution). Forty minutes after intragastric administration, the all animals were orally administrated with saccharose (2.5 g/kg, dissolved in 0.5 % CMC-Na solution), Their postprandial blood glucose was tested from the tail tip of mice by using Roche Accu-Chek Instant at 0 h, 0.25 h, 0.5 h, 1 h, 1.5 h, and 2 h. Kunming mice (25–30 g) were purchased from Laboratory Animal Center of Guizhou Medical University with the license number SCXK (Qian) 2018–0001. All animal procedures were performed in accordance with the Guidelines for Care and Use of Laboratory Animals and the experiments were approved by the animal ethics committee of Guizhou Medical University.
4.8 In vitro cytotoxicity
The cytotoxicity in vitro was evaluated by the CCK-8 method. First, HEK 293 cells were seeded in 96-well plates at a density of 1 × 105 cells per well and then incubated for 24 h. The cells were incubated with 4 at various concentrations (0, 25, 75, 125, and 675 µM) for 48 h (37 °C and 5 % CO2), respectively. 10 µL of CCK-8 reagent was added to the cell culture medium and then incubated at 37 °C for 1 h. Finally, absorbance was recorded at 450 nm using a multifunctional microplate reader (Varioskan LUX, Thermo Fisher Scientific, Vantaa, Finland). The relative cell viability (%) was calculated as the formula: (Asample - Ablank)/(Acontrol - Ablank)*100.
4.9 ADME predictions
Calculations of ADME properties were done by using the molinspiration online property toolkit (https://www.molinspiration.com/). Pharmaceutically important properties of new drugs including mi log P, H-bond donors, H-bond acceptors, number of rotatable bonds, molecular volume, and molecular weight were analyzed in accordance with Lipinski’s rule of five. The rule of five indicates that poor absorption or permeation is expected when MW > 500, NHD > 5, NHA > 10 or log P > 5 (Choy & Prausnitz, 2011). The percentage of absorption (% ABS) was calculated using the expression: % ABS = 109–0.345 × TPSA.
Acknowledgments
This work was supported by Guizhou Provincial Natural Science Foundation (ZK[2021]027), China ; Guizhou Science and Technology Department (GCC[2022]031-1), China.
References
- Log P as a tool in intramolecular hydrogen bond considerations. Drug Discov. Today Technol.. 2018;27:65-70.
- [CrossRef] [Google Scholar]
- A green chemical oligomerization of phloroglucinol induced by plasma as novel alpha-glucosidase inhibitors. Biosci. Biotechnol. Biochem.. 2018;82(12):2059-2063.
- [CrossRef] [Google Scholar]
- The rule of five for non-oral routes of drug delivery: ophthalmic, inhalation and transdermal. Pharm. Res.. 2011;28(5):943-948.
- [CrossRef] [Google Scholar]
- Synthetic heterocyclic candidates as promising alpha-glucosidase inhibitors: An overview. Eur. J. Med. Chem.. 2019;176:343-377.
- [CrossRef] [Google Scholar]
- Acarbose: safe and effective for lowering postprandial hyperglycaemia and improving cardiovascular outcomes. Openhrt.. 2015;2(1):e000327
- [CrossRef] [Google Scholar]
- New 4-(arylidene)amino-1,2,4-traizole-5-thiol derivatives and their acyclo thioglycosides as α-glucosidase and α-amylase inhibitors: Design, synthesis, and molecular modelling studies. J. Mol. Struct.. 2022;1259
- [CrossRef] [Google Scholar]
- A review on synthesis, mechanism of action, and structure-activity relationships of 1,2,3-triazole-based α-glucosidase inhibitors as promising anti-diabetic agents. J. Mol. Struct.. 2022;1255:132469
- [CrossRef] [Google Scholar]
- Occurrence, synthesis and biological activity of 2-(2-phenyethyl)chromones. Eur. J. Med. Chem.. 2022;237:114397
- [CrossRef] [Google Scholar]
- Automated docking of flexible ligands: Applications of autodock. J. Mol. Recognit.. 1996;9:1-5.
- [CrossRef] [Google Scholar]
- The modification of natural products for medical use. Acta Pharm. Sin. B. 2017;7(2):119-136.
- [CrossRef] [Google Scholar]
- Inhibitory activity and mechanism of calycosin and calycosin-7-O-β-D-glucoside on α-glucosidase: Spectroscopic and molecular docking analyses. Process Biochem.. 2022;118:227-235.
- [CrossRef] [Google Scholar]
- A review on the potential of Resveratrol in prevention and therapy of diabetes and diabetic complications. Biomed. Pharmacother.. 2020;125:109767
- [CrossRef] [Google Scholar]
- Identification of the new covalent allosteric binding site of fructose-1,6-bisphosphatase with disulfiram derivatives toward glucose reduction. J. Med. Chem.. 2020;63(11):6238-6247.
- [CrossRef] [Google Scholar]
- Cucumin S, a new phenylethyl chromone from Cucumis melo var. reticulatus seeds. Rev. Bras. Farmacogn.. 2015;25(5):462-464.
- [CrossRef] [Google Scholar]
- Synthesis, in vitro alpha-glucosidase inhibitory potential and molecular docking study of thiadiazole analogs. Bioorg. Chem.. 2018;78:201-209.
- [CrossRef] [Google Scholar]
- In vitro and in vivo inhibitory effect of anthocyanin-rich bilberry extract on α-glucosidase and α-amylase. Lwt. 2021;145
- [CrossRef] [Google Scholar]
- An ESIPT fluorescent probe sensitive to protein α-helix structures. Org. Biomol. Chem.. 2014;12(28):5243-5249.
- [CrossRef] [Google Scholar]
- Synthesis and alpha-glucosidase inhibition activity of dihydroxy pyrrolidines. Bioorg. Med. Chem. Lett.. 2017;27(12):2818-2823.
- [CrossRef] [Google Scholar]
- Synthesis and molecular docking studies of potent alpha-glucosidase inhibitors based on biscoumarin skeleton. Eur. J. Med. Chem.. 2014;81:245-252.
- [CrossRef] [Google Scholar]
- A New 2-(2-Phenylethyl)Chromone From Aquilaria sinensis. Chem. Nat. Compd.. 2018;54(1):30-33.
- [CrossRef] [Google Scholar]
- Altering the inhibitory kinetics and molecular conformation of maltase by Tangzhiqing (TZQ), a natural alpha-glucosidase inhibitor. BMC Complement Med. Ther.. 2020;20(1):350.
- [CrossRef] [Google Scholar]
- 2-(2-Phenylethyl)chromone derivatives in artificial agarwood from Aquilaria sinensis. Fitoterapia. 2016;110:38-43.
- [CrossRef] [Google Scholar]
- Development of a solvent selection guide for aldehyde-based direct reductive amination processes. Green Chem.. 2013;15(5):1159-1165.
- [CrossRef] [Google Scholar]
- Synthesis and biological evaluation of novel chromonyl David as α-glucosidase inhibitors. Med. Chem. Res.. 2019;28(6):831-848.
- [CrossRef] [Google Scholar]
- 2-(2-Phenylethyl)chromone derivatives: Promising alpha-glucosidase inhibitors in agarwood from Aquilaria filaria. Phytochemistry. 2021;181:112578
- [CrossRef] [Google Scholar]
- AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem.. 2009;30(16):2785-2791.
- [CrossRef] [Google Scholar]
- An on-line detection system for screening small molecule inhibitors of alpha-Amylase and alpha-Glucosidase in Prunus mume. J. Chromatogr. A. 2022;1663:462754
- [CrossRef] [Google Scholar]
- IDF Diabetes Atlas: Global estimates for the prevalence of diabetes for 2015 and 2040. Diabetes Res. Clin. Pract.. 2017;128:40-50.
- [CrossRef] [Google Scholar]
- alpha-glucosidase and glycation inhibitory effects of costus speciosus leaves. BMC Complement Altern. Med.. 2016;16:2.
- [CrossRef] [Google Scholar]
- Isatin based Schiff bases as inhibitors of alpha-glucosidase: Synthesis, characterization, in vitro evaluation and molecular docking studies. Bioorg. Chem.. 2015;60:42-48.
- [CrossRef] [Google Scholar]
- Python: a programming language for software integration and development. J. Mol. Graph. Model.. 1999;17(1):57-61.
- [Google Scholar]
- Optimization of in vitro carbohydrate digestion by mammalian mucosal α-glucosidases and its applications to hydrolyze the various sources of starches. Food Hydrocoll.. 2019;87:470-476.
- [CrossRef] [Google Scholar]
- Synthesis, α-glucosidase inhibitory and molecular docking studies of prenylated and geranylated flavones, isoflavones and chalcones. Bioorg. Med. Chem. Lett.. 2015;25(20):4567-4571.
- [CrossRef] [Google Scholar]
- Synthesis of novel inhibitors of alpha-glucosidase based on the benzothiazole skeleton containing benzohydrazide moiety and their molecular docking studies. Eur. J. Med. Chem.. 2015;92:387-400.
- [CrossRef] [Google Scholar]
- AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem.. 2010;31(2):455-461.
- [CrossRef] [Google Scholar]
- In vitro inhibitory effects of Chinese bayberry (Myrica rubra Sieb. et Zucc.) leaves proanthocyanidins on pancreatic alpha-amylase and their interaction. Bioorg. Chem.. 2020;101:104029
- [CrossRef] [Google Scholar]
- Synthesis, in vitro evaluation and molecular docking studies of novel triazine-triazole derivatives as potential alpha-glucosidase inhibitors. Eur. J. Med. Chem.. 2017;125:423-429.
- [CrossRef] [Google Scholar]
- Synthesis, in vitro evaluation and molecular docking studies of novel triazine-triazole derivatives as potential α-glucosidase inhibitors. Eur. J. Med. Chem.. 2017;125:423-429.
- [CrossRef] [Google Scholar]
- 2-(2-Phenylethyl)-4H-chromen-4-one Derivatives from the Resinous Wood of Aquilaria sinensis with Anti-Inflammatory Effects in LPS-Induced Macrophages. Molecules. 2018;23(2):12.
- [CrossRef] [Google Scholar]
- Cholecystokinin-8 inhibits methamphetamine-induced neurotoxicity via an anti-oxidative stress pathway. Neurotoxicology. 2016;57:31-38.
- [CrossRef] [Google Scholar]
- One-pot syntheses of novel pyrazole-containing bisphosphonate esters at room temperature. Org. Biomol. Chem.. 2012;10:7730.
- [CrossRef] [Google Scholar]
- Synthesis and biological evaluation of coumarin derivatives as alpha-glucosidase inhibitors. Eur. J. Med. Chem.. 2020;189:112013
- [CrossRef] [Google Scholar]
- Mechanism underlying the interaction of malvidin-3-O-galactoside with protein tyrosine phosphatase-1B and α-glucosidase. J. Mol. Struct.. 2022;1253
- [CrossRef] [Google Scholar]
- Dimeric sesquiterpenoid-4&ITH&IT-chromone derivatives from agarwood of & ITAquilaria crassna&IT and their cytotoxicity. Phytochemistry. 2018;145:207-213.
- [CrossRef] [Google Scholar]
- Inhibition mechanism of diacylated anthocyanins from purple sweet potato (Ipomoea batatas L.) against alpha-amylase and alpha-glucosidase. Food Chem.. 2021;359:129934
- [CrossRef] [Google Scholar]
- Exploring efficacy of natural-derived acetylphenol scaffold inhibitors for alpha-glucosidase: Synthesis, in vitro and in vivo biochemical studies. Bioorg. Med. Chem. Lett.. 2020;30(23):127528
- [CrossRef] [Google Scholar]
- Investigation of the interaction of 2,4-dimethoxy-6,7-dihydroxyphenanthrene with α-glucosidase using inhibition kinetics, CD, FT-IR and molecular docking methods. Spectrochim. Acta A Mol. Biomol. Spectrosc.. 2018;203:13-18.
- [CrossRef] [Google Scholar]
- Inhibition mechanism of ferulic acid against α-amylase and α-glucosidase. Food Chem.. 2020;317
- [CrossRef] [Google Scholar]
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2022.104301.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







