Translate this page into:
A comprehensive assessment of phytochemicals from Phyla nodiflora (L.) Greene as a potential enzyme inhibitor, and their biological potential: An in-silico, in-vivo, and in-vitro approach
⁎Corresponding author. hammad.saleem@uvas.edu.pk (Hammad Saleem)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
This work explored Phyla nodiflora (L.) Greene as a potential source of the bioactive medicinal agent. In this aspect, methanol (PN-M) and dichloromethane (PN-D) extracts were prepared from the whole plant and evaluated for phytochemical composition (total bioactive contents, UHPLC-MS analysis, and HPLC-PDA polyphenolic quantification), biological (antioxidant and enzyme inhibition) potential and in-vivo toxicity. The PN-M was found to contain higher phenolic (26.08 mg GAE/g extract) and flavonoid (50.25 mg QE/g extract) contents which might correlate to the higher radical scavenging (DPPH: 52.94 mg TE/g extract; ABTS: 72.11 mg TE/g extract) and reducing power (FRAP: 71.96 mg TE/g extract; CUPRAC: 142.65 mg TE/g extract) antioxidant potential, as well as AChE (4.33 mg GALAE/g extract), tyrosinase (125.36 mg KAE/g extract), and amylase (1.86 mmol ACAE/g extract) inhibition activity of this extract. In contrast, the PN-D extract was found to be most active for phosphomolybdenum (1.30 mg TE/ g extract) and metal chelation (54.84 mg EDTAE/g extract) assays in addition to BChE (4.70 mg GALAE/g extract) and glucosidase (0.62 mmol ACAE/g extract) enzyme inhibition activity. The PN-M extract on UHPLC-MS analysis revealed the tentative identification of 24 different secondary metabolites, most of which belonged to the flavonoid, glycoside, and terpenoid classes of phytochemicals. The polyphenolic composition of the extracts was appraised by HPLC-PDA. Seven phenolic compounds were identified in the extracts. PN-M was found to be rich in catechin (0.25 µg/extract) and 3-OH benzoic acid (0.64 µg/extract), while PN-D contained epicatechin (0.30 µg/extract), 3-OH-4-MeO benzaldehyde (0.21 µg/extract), and 2,3-Di-Meo benzoic acid (0.97 µg/extract) in higher amounts. The methanol extract was found to be non-toxic even at higher doses. Furthermore, the relationship between the phytochemicals and the tested enzymes was highlighted by molecular docking studies. In sum, this research showed that the studied extracts were effective as enzyme inhibitors and antioxidants, suggesting it would be worth investigating in more depth for further advanced studies to explore its pharmacological properties.
Keywords
Phyla nodiflora
Phytochemicals
Antioxidant
Enzyme inhibition
Toxicity
HPLC
1 Introduction
Plant species have been used for medicine for as long as humans have existed. Different civilizations have provided documentary evidence of the medicinal use of plants (Phillipson 2001, Suroowan, Llorent-Martínez et al. 2023). The therapeutic effects of plants are due to the presence of different organic and inorganic salts (Kurmukov 2013, Acquaviva, Di Simone et al. 2023). After years of struggle, the use of plants was established as medicine against various ailments (Petrovska 2012). Primary and secondary metabolites produced by the plants are the sources for different drugs to be used medicinally (Balick and Cox 1996). Humans have been using folk medicines for thousands of years extracted from plants, which participated in the conception of modern medicines (Hassan 2015, Acquaviva, Di Simone et al. 2023). Drugs are obtained from plants by employing different botanical, biological, phytochemical, and molecular techniques (Balunas and Kinghorn 2005).
The medicinal plant “P. nodiflora” is a predominant member of the Verbenaceae family. Its common habitat is in Central and South America, Pakistan, Sri Lanka, and India. P. nodiflora grows best on moist soil. It is present in soil rich in moisture (Sharma and Singh 2013, Al-Snai 2019, Rahman, Javaid et al. 2021). Two flavones, luteolin-7-O-glucoside and 6-hydroxyluteolin-7-apioside, are mainly present in the flowers of P. nodiflora and the leaves of P. nodiflora were also found to be contain different flavone glycosides (Barnabas, Gunasingh et al. 1980, Jabeen, Jillani et al. 2016). Acetoside and demethoxycentaureidin are the two compounds yielded from the alcoholic extract of P. nodiflora (Khalil, Lahloub et al. 1995). The isolation of stigmasterol, beta-sitosterol, and triterpenoid lippacin was reported from the methanolic extract of P. nodiflora (Sharma and Singh 2013). The aerial parts also reported numerous steroidal constituents (Siddiqui, Ahmed et al. 2009). Methanol and dichloromethane extract of P. nodiflora leaves are reported to contain the compounds halleridone and halleron (Ravikanth, Ramesh et al. 2000). A skin whitening agent, eupafolin, was yielded from dried aerial parts of the P. nodiflora (Yen, Wang et al. 2012). The methanol extract of P. nodiflora yields hispidulin, nodifloretin, and eupafolin (Ko, Chiang et al. 2014). The ethanolic extract of P. nodiflora contains a diverse group of chemical compounds, including betatifolin, nodifloridin, jaceosidin, lippiflorin, and nepetin (Regupathi and Chitra 2015). The ethanolic extract of P. nodiflora has also been reported as the main source of the ecteoside (Khalil, Lahloub et al. 1995). Likewise, stigmasterol, eugenol, α-copaene, β-bisabolene, and γ-sitosterol are among the various constituents isolated from the plant’s methanolic extract (Ko, Chiang et al. 2014). Similarly, steam distillation of P. nodiflora has reported the presence of linalool, methyl salicylate, cymen-8-ol, and β carboline (Elakovich and Stevens 1985). P. nodiflora have been previously reported for a number of biological effects (Al-Snai 2019, Paua 2022), including antimicrobial (Gopal, Balkrishna et al. 1996, Patel Janki, Shah Kinjal et al. 2005, Pirzada, Iqbal et al. 2005, Durairaj, Vaiyapuri et al. 2007, Malathi, Cholarajan et al. 2011, Sharma and Singh 2013), antitumor (Vanajothi, Sudha et al. 2012, Cheong and Teoh 2014), hepatoprotective (Durairaj, Vaiyapuri et al. 2008, Arumanayagam and Arunmani 2015), antioxidant (Ashokkumar, Thamilselvan et al. 2008, Durairaj, Vaiyapuri et al. 2008), diuretic (Shukla, Patel et al. 2009, Balamurugan and Ignacimuthu 2011), antidiabetic, hypolipidemic (Balamurugan, Duraipandiyan et al. 2011, Balamurugan and Ignacimuthu 2011), neuropharmacological (Thirupathy, Tulshkar et al. 2011, AO and SL 2012), anti-urolithiasis (Dodoala, Diviti et al. 2010), anti-inflammatory (Ahmed, Selim et al. 2004, Balakrishnan, Janakarajan et al. 2010, Al-Snafi and Faris 2013), melanogenesis inhibition (Yen, Wang et al. 2012, Ko, Chiang et al. 2014), antihypertensive (Gadhvi, Mishra et al. 2012), skin whitening effect (Yen, Wang et al. 2012), central inhibitory (AO and SL 2012), apoptotic (Cheong and Teoh 2014), and antidandruff (Regupathi and Chitra 2015) activities.
Hence, the estimation of phytochemical contents both quantitatively and qualitatively, along with the assessment of antioxidant and enzyme inhibition potential, as well as the in-vivo toxicological studies of methanol and dichloromethane extracts of P. nodiflora, were the main objectives of this paper. The phytochemical profile was evaluated by ascertaining preliminary phytochemical testing, total phenolic and flavonoid contents, UHPLC-MS analysis, and HPLC-PDA polyphenolic quantification. The antioxidant potential was assessed via different assays, including radical scavenging (DPPH and ABTS), reducing power (FRAP and CUPRAC), total antioxidant activity (phosphomolybdenum), and metal chelation assays. Likewise, the inhibitory potential of both extracts was tested against the five therapeutically relevant enzymes, including AChE and BChE (involved in neurological problems), amylase, glucosidase (involved in diabetes), and tyrosinase (skin problems). Furthermore, in-silico studies against the tested enzymes were also conducted. The in-vivo toxicity was tested on the chicks for 21 days. This work will provide evidence for the pharmacological effectiveness of these species by identifying the best methods for isolating their bioactive components.
2 Material and methods
2.1 Plant collection and extraction
The whole plant material was collected from the peripheries of Bahawalpur City, Pakistan (GPS co-ordinates: 29°23′19.7″N 71°42′07.0″E). The plant was identified and authenticated as P. nodiflora by Mr. Hafiz Waris, Taxonomist at Cholistan Institute of Desert Studies (CIDS), The Islamia University of Bahawalpur. For future reference, a voucher specimen of the plant material (PN-WP-01–14-131) was also deposited at the herbarium of the Islamia University of Bahawalpur's Faculty of Pharmacy and Alternative Medicines. The plant material was subjected to shade drying for 15 days. The dried plant material was grounded into a fine powder with the help of a grinding mill. The powdered plant material was extracted with dichloromethane and methanol successively. The filtrate collected was concentrated on a Rotary evaporator at 35 °C under reduced pressure.
2.2 Phytochemical composition
2.2.1 Total phenolic and total flavonoid contents
The total bioactive contents, including total phenolic and flavonoid contents, were estimated using well-established Folin-Ciocalteu and aluminium chloride assays, respectively. (Saleem, Zengin et al. 2019). The standard for total phenolic content was gallic acid, with results expressed as mg GAE/g (gallic acid equivalents), while quercetin was used as a standard for total flavonoid content, and the results were expressed as mg QE/g (quercetin equivalent). The detailed protocols for these assays are presented in the supplementary material section.
2.2.2 UHPLC-MS analysis
UHPLC-MS analysis is a technique that is mostly used for the tentative identification of polar compounds in plant extracts. Therefore, the methanol extract (being polar one) was subjected to the UHPLC-MS analysis (negative ionization mode) for the tentative identification of possible secondary metabolites. The identification of the phytochemicals was made by using METLIN database library (Saleem, Zengin et al. 2019). The detailed instrumentation for this analysis is presented in the supplementary material section.
2.2.3 HPLC-PDA quantification of different phytochemicals
HPLC-PDA analysis (of both the methanol and dichloromethane extracts of P. nodiflora) for 22 different phytochemicals (including 4-hydroxybenzoic acid, 3-dimethoxybenzoic acid, quercetin, gallic acid, benzoic acid, catechin, epicatechin, harpagoside, naringin, vanillic acid, carvacrol, 3-hydroxybenzoic acid, naringenin, 3-hydroxy-4-methoxy benzaldehyde, t-cinnamic acid, p-coumaric acid, chlorogenic acid, rutin, syringic acid, sinapinic acid, t-ferulic acid, and o-coumaric acid) was done and quantified using previously reported method (Locatelli, Zengin et al. 2017). The gradient elution program used for HPLC analyses is given in Table S1 of the supplementary material section. Likewise, the standards used, retention times, and maximum wavelengths used for quantitative analyses are shown in Table S2 of the supplementary material section. The chemical standards chromatogram for the 22 standards used is depicted in Figure S1 of the supplementary material section. The detailed instrumentation for this analysis is presented in the supplementary material section.
2.3 Biological assays
2.3.1 Antioxidant assays
The metal chelating, phosphomolybdenum, FRAP, CUPRAC, ABTS, and DPPH activities were made as per previously described methods (Saleem, Zengin et al. 2019). The antioxidant activities were reported as Trolox equivalents, whereas EDTA was used for the metal-chelating assay. The detailed protocols for these assays are presented in the supplementary material section.
2.3.2 Enzyme inhibition activities
The possible inhibitory effects of the extracts against cholinesterases, tyrosinase, α-amylase, and α-glucosidase were assessed using standard bio-assays (Saleem, Zengin et al. 2019). Galantamine was used as a standard for AChE, and BChE inhibition activity was expressed as (mg GALAE/g extract). In contrast, the inhibition activities for α-glucosidase and α-amylase were reported as millimoles of acarbose equivalent (mmol ACAE/g extract), whereases tyrosinase inhibition was noted as milligrams of kojic acid equivalent (mg KAE/g extract). The detailed protocols for these assays are presented in the supplementary material section.
2.4 Computation methods
2.4.1 Structures preparation and docking studies
Sybyl-X1.3/SKETCH (Jain 2003) was used to create a three-dimensional conformation of inhibitors 6-hydroxyluteolin 5-rhamnoside (HLR), luteolin 7-rhamnosyl (1->6) galactoside (LRG), and torosaflavone D (TFD) (Fig. 3). To get biologically active conformation of compounds under study, all three inhibitors were subjected to energy optimization using the Tripos force field with Gasteiger Hückel atomic charges (Powell 1978). The co-crystal structures of acetylcholinesterase (ACHE), butyrylcholinesterase (BCHE), and tyrosinase (TYR) were obtained from the RCSB Protein Data Bank under the following PDB entries; (PDB ID’s: 4EY7 (Cheung, Rudolph et al. 2012), 6QAE (Meden, Knez et al. 2019) and 2Y9X (Ismaya, Rozeboom et al. 2011), respectively). Before docking investigations, all protein structures were further processed using the structure preparation tools contained in the SYBYL-X 1.3 biopolymer module (Ghersi and Sanchez 2011). The energy was minimized using the Powell algorithm with a convergence gradient of 0.5 kcal (mol)-1 for 1000 cycles, with missing hydrogens added, charges applied, and atom types assigned according to the AMBER 7 FF99 force field (Onufriev, Bashford et al. 2004). Finally, the energy-optimized bioactive conformation of selected compounds was docked into the active site of ACHE, BCHE, and TYR enzymes using the Surflex-Dock module of the SYBYL-X 1.3 software package, following the same protocol and parameters as those published in our prior work.(Chohan, Chen et al. 2016, Chohan, Qian et al. 2016). The experimentally determined active conformation of donepezil in 4EY7, N1, N2-dimethylpropane-1,2-diamine in 6QAE, and tropolone in 2Y9X was used as initial conformation to define the potential binding pocket for protomol (an idealized active site) generation (Jain 2007). The top twenty docked conformations were saved and investigated for their binding modes into the active site of their respective target. The Hammerhead scoring method was used to rate these potential ligand poses (Jain 1996, Jain 2003).
2.5 In-vivo toxicological studies
The in-vivo toxicological parameters were studied on the broiler chicks. The animal's selection, grouping, toxicity parameters, hematological and biochemical analysis, and relative and absolute weight of body organs were calculated as reported in our previously published data (Saleem, Zengin et al. 2019).
2.6 Statistical analysis
All the biological experiments were done in triplet, and the estimation of results was made as mean value ± standard deviation (SD). One-way ANOVA test was used to find the mean difference, followed by data analysis using SPSS v.17.0 software.
3 Results and discussion
3.1 Antioxidant potential
The involvement of oxidative stress in disease development and progression is becoming more evident. Increasing endogenous antioxidant defenses or supplementing the body with exogenous antioxidants is a viable approach to combating ROS-induced oxidative damage (Pizzino, Irrera et al. 2017, Lan, Liu et al. 2021). The plant kingdom provides a rich supply of beneficial substances to one's health, particularly natural antioxidants. (Zengin, Mahomoodally et al. 2022). In the current study, PN-M and PN-D extracts of the P. nodiflora plant were tested for a panoply of antioxidant assays, including radical scavenging (DPPH and ABTS), reducing power (FRAP and CUPRAC), total antioxidant activity (phosphomolybdenum) and metal chelation activities, and the results are presented in Table 1. A similar pattern to total bioactive contents was noted, and the methanol extract was found to have a higher antioxidant potential for radical scavenging (DPPH: 52.94 mg TE/g extract; ABTS: 72.11 mg TE/g extract) and reducing power (FRAP: 71.96 mg TE/g extract; CUPRAC: 142.65 mg TE/g extract) assays. This higher antioxidant activity of methanol extract might be linked to the higher amount of phenolic and flavonoid contents for this extract. These findings agree with some of the previous research data, which have provided a positive relationship between bioactive contents and antioxidant potential (Chang, Ye et al. 2022, Chen, Ran et al. 2022, Li, Shi et al. 2022, Zengin, Fernández-Ochoa et al. 2023). However, in the case of phosphomolybdenum and MCA assay, different results were obtained, and the dichloromethane extract was found to contain higher activity. This higher activity could be linked to the non-phenolic compounds present in this extract. These findings correlate with the previous studies, which also presented the same trend, and the methanolic extract of P. nodiflora has higher antioxidant potential for tested reducing power, scavenging free radical, and scavenging superoxide anion radical activities and other antioxidant assays (Ashokkumar, Thamilselvan et al. 2008, Durairaj, Vaiyapuri et al. 2008). Meticulous correlation values between these assays in plant extracts have been published by various researchers, consistent with our findings. However, we noticed a low correlation value for the PBD assay, which could be explained by additional reducing chemicals in the extracts (peptides, sugars, etc.). The poor correlation value obtained between total phenolics and the metal chelating assay can be explained by the activity of non-phenolic chelators present in the examined extracts, as demonstrated in the previous study (References) (Zengin, Ak et al. 2022). PN-M: P. nodiflora methanol extract; PN-D: P. nodiflora dichloromethane extract. GAE: gallic acid equivalent; QE: quercetin equivalent. TE: trolox equivalent; EDTAE: EDTA equivalent. Values expressed are means ± S.D. of three parallel measurements.
Extracts
Total bioactive contents
Antioxidant activities
Radical Scavenging activity
Reducing power
Total antioxidant capacity (TAC)
Ferrous chelating
TPC (mg GAE/g)
TFC (mg QE/g)
DPPH (mgTE/g extract)
ABTS (mgTE/g extract)
FRAP (mgTE/g extract))
CUPRAC (mgTE/g extract)
PBD (mgTE/g extract)
Metal Chelating (mgEDTA/g)
PN-M
26.08 ± 1.32
50.25 ± 0.15
52.94 ± 0.47
72.11 ± 2.49
71.96 ± 1.67
142.65 ± 1.41
1.17 ± 0.03
32.18 ± 0.53
PN-D
18.85 ± 0.38
31.80 ± 0.84
3.44 ± 0.58
12.86 ± 0.86
35.66 ± 0.67
86.61 ± 1.01
1.30 ± 0.09
54.84 ± 0.43
3.2 Enzyme inhibition
Drug discovery relies heavily on the use of enzyme inhibitors. The malfunction, overexpression, or hyperactivation of enzymes is the root cause of many diseases, thanks to advances in molecular biology. Enzyme inhibitors can be used to treat such hyperactivation or overexpression of enzymes. These efforts have resulted in the clinical use of various enzyme inhibitors, some of which are of natural origin. (Zengin, Mahomoodally et al. 2022).
Several clinically relevant enzymes involved in major pathologies, including neurodegenerative disorders (AChE and BChE), skin diseases (tyrosinase), and diabetes (amylase and glucosidase), were tested for enzyme inhibition potential using P. nodiflora methanol and dichloromethane extracts. The outcomes of these tests are summarised in Table 2. The PN-M extract was most active against AChE and tyrosinase enzymes with an inhibition potential of 4.33 mg GALAE/g extract and 125.36 mg KAE/g extract, respectively. Likewise, the dichloromethane extract was noted to show higher inhibition values against BChE (4.70 mg GALAE/g extract) and amylase (1.97 mmol ACAE/g extracts) enzymes. However, both extracts presented weak inhibition potential against the glucosidase enzyme. PN-M: P. nodiflora methanol extract; PN-D: P. nodiflora dichloromethane extract. GALAE: galantamine equivalent; KAE: kojic acid equivalent; ACAE: acarbose equivalent. All values expressed are means ± S.D. of three parallel measurements.
Extracts
Neurodegenerative diseases
Skin problems
Diabetes
AChE inhibition (mg GALAE/g extract)
BChE inhibition (mg GALAE/g extract)
Tyrosinase (mg KAE/g extract)
Glucosidase (mmol ACAE/g extract)
Amylase (mmol ACAE/g extract)
PN-M
4.33 ± 0.05
2.11 ± 0.06
125.36 ± 1.65
0.60 ± 0.02
1.86 ± 0.03
PN-D
3.48 ± 0.53
4.70 ± 0.14
116.04 ± 1.12
0.62 ± 0.01
1.97 ± 0.01
Mice have been shown to benefit from extracts of P. nodiflora in chloroform and ethanol as anxiolytics and anticonvulsants (Thirupathy, Tulshkar et al. 2011). P. nodiflora methanol extract has previously shown anti-diabetic action in rats induced with Streptozotocin. A significant increase in insulin levels and body weight resulted in decreased blood glucose and glycosylated hemoglobin levels (Balamurugan and Ignacimuthu 2011). Eupafolin, an active flavonoid derived from P. nodiflora, exhibited skin-whitening effects in B16F10 mice via a melanin suppression (Ko, Chiang et al. 2014). Likewise, another study has reported the tyrosinase inhibitory potential of P. nodiflora in the dose-dependent matter (Yen, Wang et al. 2012).
3.3 Phytochemical composition
Natural products play a key role in drug development; therefore, screening plants to identify and isolate active ingredients is a necessary step to developing effective drugs against a wide range of diseases (Bibi Sadeer, Sinan et al. 2022). Therefore, total flavonoid and phenolic contents were estimated, with results depicted in Table 1. The tested extracts contained considerable quantities of total phenolic and flavonoid contents. However, the methanol extract was higher in polyphenolic contents (TPC: 26.08 mg GAE/g extract; TFC: 50.25 mg QE/g extract) than dichloromethane extracts. The higher amount of phenolic and flavonoids in methanol extract might be attributed to the polar nature of the methanol solvent. Our findings are in line with the previous studies that had reported higher bioactive contents for polar extracts (Zekri, Zerkani et al. 2021, Wairata, Fadlan et al. 2022). Current studies made use of colorimetric methods to quantify phenolic compounds. Although, the effectiveness of these methods is quite controversial in producing accurate results (Zengin, Ak et al. 2022). Thus, a punctual UHPLC-MS and HPLC-PDA analysis was used to reveal the quantitative and qualitative composition of the phenolic compounds.
As mentioned above, to have further insight into the composition of the secondary metabolites, the methanol extract was subjected to UHPLC-MS analysis in negative ionization mode which resulted in the tentative identification of 24 different phytochemicals. The list of these identified compounds is presented in Table 3, while the total ion chromatogram (TIC) is shown in Fig. 1. It can be noted from Table 2 that the maximum number of phytochemicals identified belong to the flavonoid group of secondary metabolites. The identified flavonoids were 6-methoxytaxifolin, luteolin 7-rhamnosyl (1->6) galactoside, 6-hydroxyluteolin 5-rhamnoside, luteolin 7-rhamnosyl (1->6) galactoside, maysin, acacetin 7-(6′'-methylglucuronide), demethyltorosaflavone D, grossamide, and torosaflavone D. Likewise, periandrin V, betavulgaroside X, spinacoside D, spinacoside C, betavulgaroside VI, and medicoside H were belonging to terpene derivatives. Similarly, three glycosides, including sarmentosin epoxide, citrusin F, and calendulaglycoside E, were also identified. One phenolic (formononetin 7-O-glucoside-6′'-O-malonate), and saponin (durupcoside B) derivative was also identified. The presence of these phytochemical classes in the PN-M extract agrees with some previous studies that reported the isolation/identification of different flavonoids, glucosides, and triterpenoid derivatives (Barnabas, Gunasingh et al. 1980, Khalil, Lahloub et al. 1995, Ko, Chiang et al. 2014, Al-Snai 2019). RT: retention time; B.P: base peak; Mol. Formula; AUC: area under curve.
S.no
RT (min)
BP (m/z)
Compounds tentatively identified
Mol. Formula
Comp class
Mol. mass
AUC
1
0.63
333.06
6-Methoxytaxifolin
C16H14O8
Flavonoid
334.06
180,964
2
0.762
290.09
Sarmentosin epoxide
C11H17NO8
C. glycoside
291.09
1,010,800
3
8.462
593.15
Luteolin 7-rhamnosyl(1->6)galactoside
C27H30O15
Flavonoid
594.15
665,330
4
8.504
225.12
12-hydroxyjasmonic acid
C12H18O4
Acid
226.12
559,385
5
8.512
447.10
6-Hydroxyluteolin 5-rhamnoside
C21H20O11
Flavonoid
448.10
1,264,764
6
8.66
519.18
Citrusin F
C22H32O14
Phenolic Glycoside
520.18
257,059
7
8.979
577.16
Isovitexin 7-O-rhamnoside
C27H30O14
Flavonoid
578.16
1,665,617
8
8.987
593.15
Luteolin 7-rhamnosyl(1->6)galactoside
C27H30O15
Flavonoid
594.15
665,330
10
9.504
575.14
Maysin
C27H28O14
Flavonoid
576.14
379,636
11
9.505
577.16
Isovitexin 7-O-rhamnoside
C27H30O14
Flavonoid
578.16
561,198
12
9.604
515.12
Formononetin 7-O-glucoside-6′'-O-malonate
C25H24O12
Phenol
516.12
225,692
13
9.939
473.11
Acacetin 7-(6′'-methylglucuronide)
C23H22O11
Flavonoid
474.11
536,694
14
10.77
355.05
Demethyltorosaflavone D
C18H12O8
Flavonoid
356.05
536,952
15
11.37
777.41
Periandrin V
C41H62O14
Triterpene
778.41
335,538
16
11.39
909.45
Betavulgaroside X
C46H70O18
Diterpenoid
910.45
296,898
17
11.85
793.44
Calendulaglycoside E
C42H66O14
Glycoside
794.44
599,057
18
11.99
623.24
Grossamide
C36H36N2O8
Flavonoid
624.24
793,947
19
12.03
763.39
Spinacoside D
C40H60O14
Oleane triterpenoid
764.39
321,379
20
12.08
925.45
Spinacoside C
C46H70O19
Oleane triterpenoid
926.45
773,840
21
12.081
971.45
Betavulgaroside VI
C47H72O21
Diterpenoid
972.45
421,626
22
12.36
369.06
Torosaflavone D
C19H14O8
Flavonoid
370.06
311,323
23
12.43
941.48
Medicoside H
C47H74O19
Triterpene glycoside
942.48
356,757
24
13.37
925.48
Durupcoside B
C47H74O18
Saponin
926.48
3,474,724
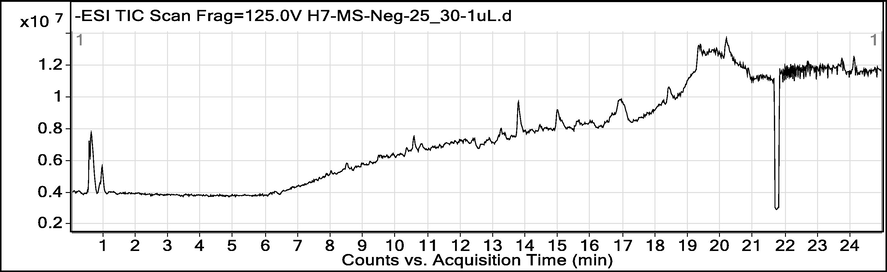
Total ion chromatogram (TIC) of UHPLC-MS analysis P. nodiflora methanol extract.
Likewise, the polyphenolic composition of the extracts was appraised by HPLC-PDA quantification for the 22 different phenolic standards, and the results of the quantified compounds are presented in Table 4, while their HPLC chromatograms are depicted in Fig. 2. It was noted that seven phenolic compounds were quantified in both extracts. PN-M was found to be rich in catechin (0.25 µg/extract) and 3-OH benzoic acid (0.64 µg/extract). In comparison, PN-D contained epicatechin (0.30 µg/extract), 3-OH-4-MeO benzaldehyde (0.21 µg/extract), and 2,3-Di-Meo benzoic acid (0.97 µg/extract) in higher amounts. PN-M: P. nodiflora methanol extract; PN-D: P. nodiflora dichloromethane extract. nd: not detected; BLD: below limit of detection (<0.1 μg/mL); Chlorogenic acid, p-coumaric acid, rutin, sinapinic acid, t-ferullic acid, o-coumaric acid, quercetin, harpagoside, t-cinnamic, carvacrol, benzoic acid, naringin, syringic acid, vanillic acid, and epicatechin were not detected in any of the tested extracts.
Tested extracts
Polyphenolics quantified
Catechin
p-OH benzoic acid
Vanillic acid
Epicatechin
3-OH benzoic acid
3-OH-4-MeO benzaldehyde
2,3-di-MeO benzoic acid
PN-M
0.25 ± 0.02
BLD
BLD
nd
0.64 ± 0.05
nd
nd
PN-D
nd
nd
nd
0.30 ± 0.02
nd
0.21 ± 0.01
0.97 ± 0.08
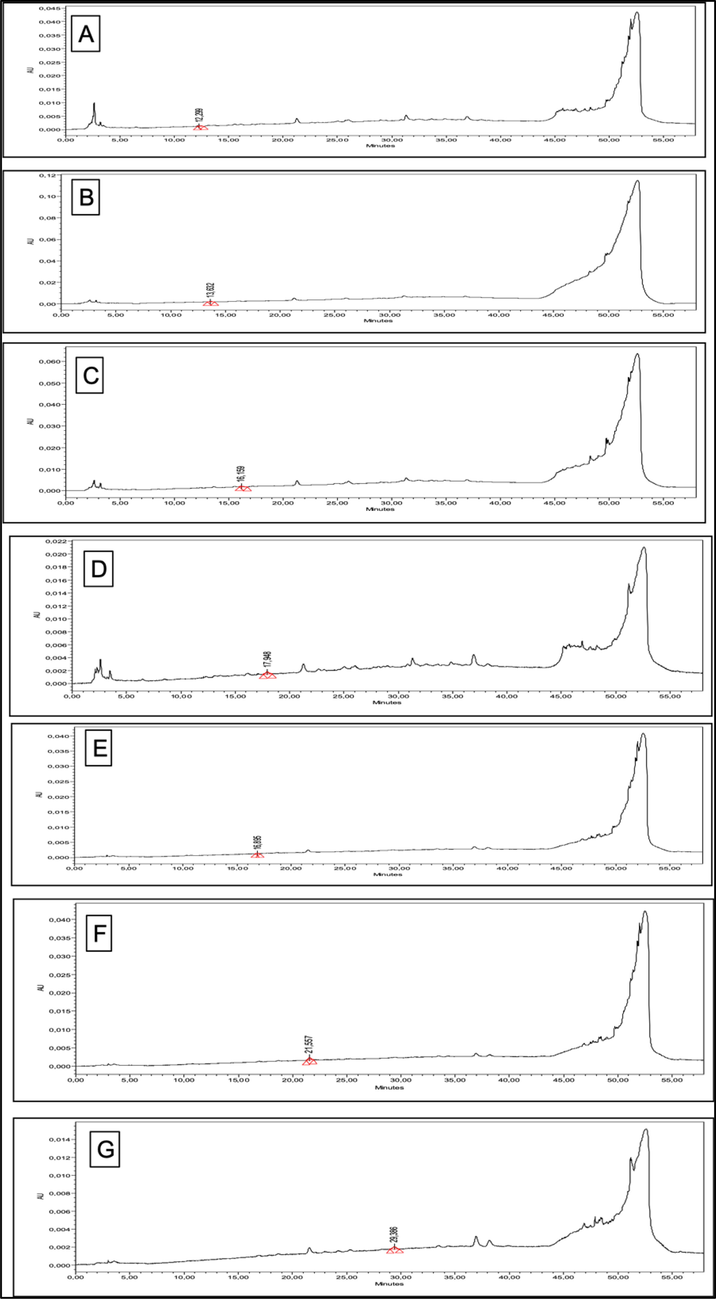
HPLC chromatograms of the quantified compounds from P. nodiflora extracts. (A: catechin 278 nm; B: p-OH benzoic acid 256 nm; C: vanillic acid 260 nm; D: 3-OH benzoic acid 295 nm; E: epicatechin 278 nm; F: 3-OH 4-MeO benzaldheyde 275 nm; G: 2,3-di-MeO benzoic acid 299 nm).
3.4 Molecular docking studies
Molecular docking studies in the ligand-binding cavity of enzymes AChE, BChE, and TYR were done to explain the binding behavior of three phytoconstituents, HLR, LRG, and TFD (Fig. 3). A greater understanding of the structures of AChE, BChE, and TYR may aid in identifying important molecular interactions involved in forming ligand–protein complexes. In order to investigate the changes in interaction patterns that contribute to different docking scores, optimal docking conformations for each compound are saved and displayed graphically. As shown in Fig. 2, the examined ligands occupy the same binding site of AChE, BChE, and TYR as observed in their co-crystallized complexes (4EY7, 6QAE, and 2Y9X) with their respective inhibitors.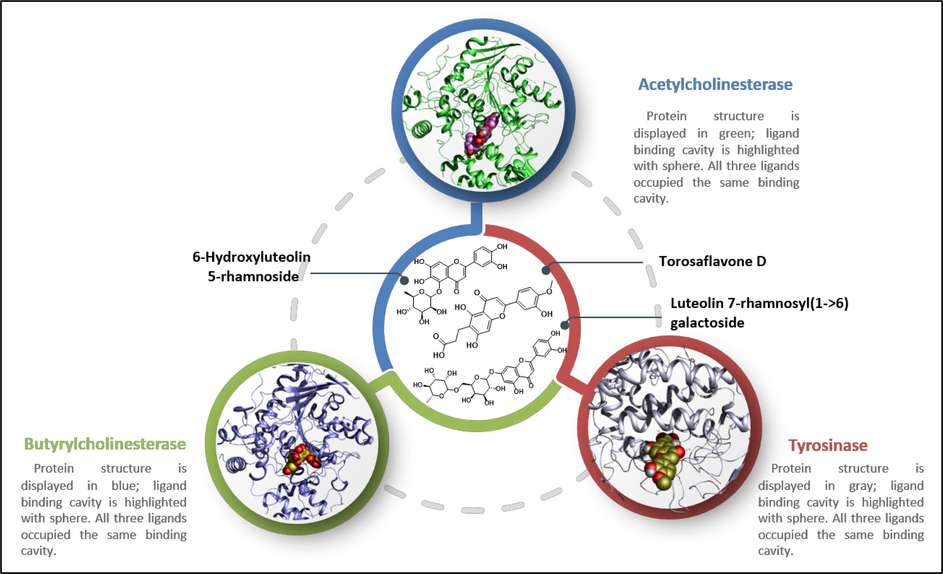
Chemical structures of selected ligands HLR, LRG, and TFD for molecular docking analysis. 3D-conformation of selected hits in their corresponding molecular targets.
A series of molecular docking simulations were performed to study the pattern of molecular interactions contributing to the variance in the conformations of the analyzed ligands to the target protein. Based on the cumulative score (cScore) (Table 5), the best probable poses of ligands in the active site of proteins were chosen. The docking scores (cScore) of selected inhibitors bonded to the AChE complex are 8.78, 7.94, and 7.20 for ligands HLR, LRG, and TFD, respectively. These results indicate that the compound HLR forms the strongest complex with the AChE enzyme among the studied ligands. Other compounds, such as LRG and TFD have also shown a comparable binding affinity towards the AChE. In the case of BChE-ligand bonded systems, compounds HLR and TFD demonstrated almost similar docking scores (cScore: 5.01 and 5.12, respectively), while the compound HLR demonstrated the highest binding affinity towards BChE (cScore: 6.52). Among TYR-ligand complexes, TFD showed substantial binding potential towards TYR (cScore: 7.80, whereas compounds HLR and LRG demonstrated weak to moderate binding affinity towards the TYR enzyme. To get detailed insight into the docking results, the docking scores and the list of residues involved in H-bond interactions are summarized in Table 5. Furthermore, the simplified 2D ligand interaction diagrams for all docking poses are depicted in Fig. 4 to get better insight into the interaction pattern of ligand-receptor complexes.
Protein
Ligand
CScorea
Crash scoreb
Polar scorec
D scored
PMF scoree
G scoref
Chem scoreg
Amino acid interaction
AChE
HLR
8.78
−3.20
3.71
−133.143
−86.23
−187.93
−35.13
D72, Y121, S286, Y334, H440
LRG
7.94
−2.17
5.37
−170.15
−97.31
−206.44
−25.41
S81, G118, Y121, E199, S200, Y334,
TFD
7.20
−3.70
1.74
−165.67
−83.16
−241.91
−18.35
S286, F288
BChE
HLR
5.09
−2.38
4.83
−92.17
−24.4
−151.4
−24.7
D70, W82, E197, S198, P285, L286, Y332, W430,
LRG
6.52
−2.11
4.91
−102.12
−28.1
−161.6
−19.6
W82, N83, T120, G116, G117, S287, W231, P285
TFD
4.12
−2.81
5.12
−166.7
−44.4
−212.1
−14.0
D70, S72, G116, S198, A199, P285
TYR
HLR
4.41
−1.49
1.23
−98.21
33.14
−121.10
−34.45
N260, R268
LRG
5.98
−4.73
7.10
−170.3
62.053
−319.8
−6.22
H85, N260, E322
TFD
7.80
−2.62
5.56
−148.0
32.933
−191.8
−4.97
H85, H244, H259, H263, R268
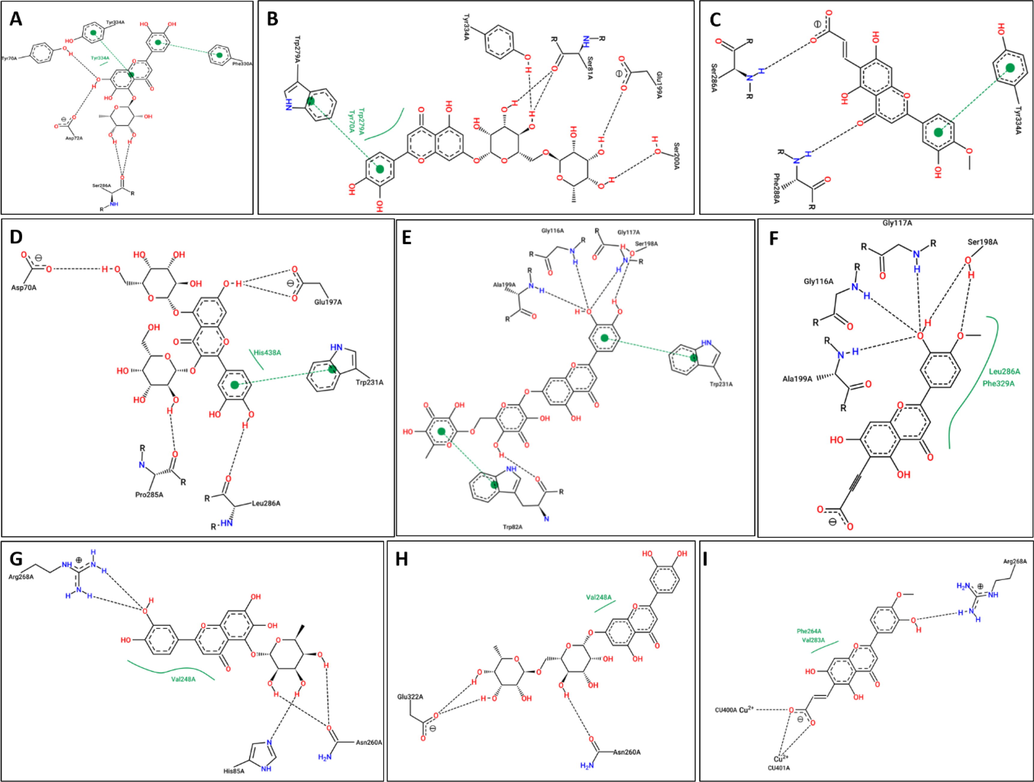
2D-interaction diagram of studied ligands in complex with AChE, BChE and TYR enzymes. (A) AChE-HLR, (B) AChE-LRG, (C) AChE-TFD, (D) BChE-HLR, (E) BChE-LRG, (F) BChE-TFD, (G) TYR-HLR, (H) TYR-LRG, (I) TYR-TFD.
The best docking conformations were saved for each ligand-receptor complex and graphically inspected to determine the ligand–protein interactions responsible for the variation in binding affinity. Moreover, only the top-ranking docking complexes for each ligand–protein system are presented in Fig. 5 (A-I) to reveal the important molecular interactions responsible for the superior binding affinities of top-scored ligands for their corresponding molecular target. Graphical analysis of all docking complexes (Figs. 3 and 4) demonstrates that all ligands occupy the same binding cleft to adopt identical binding orientations in their respective molecular target. However, the differences in binding affinities may originate from variations in the interaction pattern of the ligand-bound system. As presented in Fig. 5A-C, all three ligands (HLR, LRG, and TFD) penetrate deeply into the two main binding subsites (CAS and PAS) of AChE. In the active site of AChE, both ligands HLR, LRG, and TFD acquire such a binding conformation to establish several interactions with surrounding residues Y70, D72, Y121, W279, S286, I287, R289, F290, F330, F331, G335, Y334, and H440 (Fig. 5A and 2B). In the HLR-bonded system, HLR covers a major area of AChE’s active site to establish at least six H-bond interactions with D72, Y121, S286, Y334, and H440. In addition to the conserved H-bond contacts with Y121 and Y334, compound LRG also establishes four H-bond interactions with S81, G118, E199, and S200. Compound TFD was found to establish only three H-bond contacts with residues S286 and I287. Despite fewer H-bonds, considering the binding affinity of TFD towards AChE might be attributed to van der Waals's contact of TFD with nearby residues D72, W84, L282, F331, Y334. In addition, the benzyl moiety faces the aromatic indole moiety of W279 to establish pi-pi contact. Hence, these nonbonding interactions may play a crucial role in improved binding affinities of HLR, LRG, and TFD towards AChE. In BChE-ligand bonded systems, compound LRG demonstrated the highest binding affinities among studied ligands, which a network of nine H-bonds may explain between LRG and residues W82, N83, T120, G116, G117, S287, W231, P285of BChE. Compound LFD displays the least binding affinity towards BChE by establishing fewer H-bond contacts. Interestingly, LFD demonstrated the highest binding affinity in TYR-ligand complexes by acquiring six H-bonds with residues H85, H244, H259, H263, and R268 of the TYR enzyme. Moreover, the aromatic moiety in LFD is sandwiched between V283 and H263 to contribute to sigma-pi and pi-pi contacts, respectively. Compounds of HLR and LRG with a moderate affinity towards AChE develop three to four H-bonds with the nearby residues in the active site of the TYR enzyme (Fig. 5G and H). Overall, the docking analysis is consistent with the results and effectively explains the influence of interaction changes on the binding affinities of chosen inhibitors.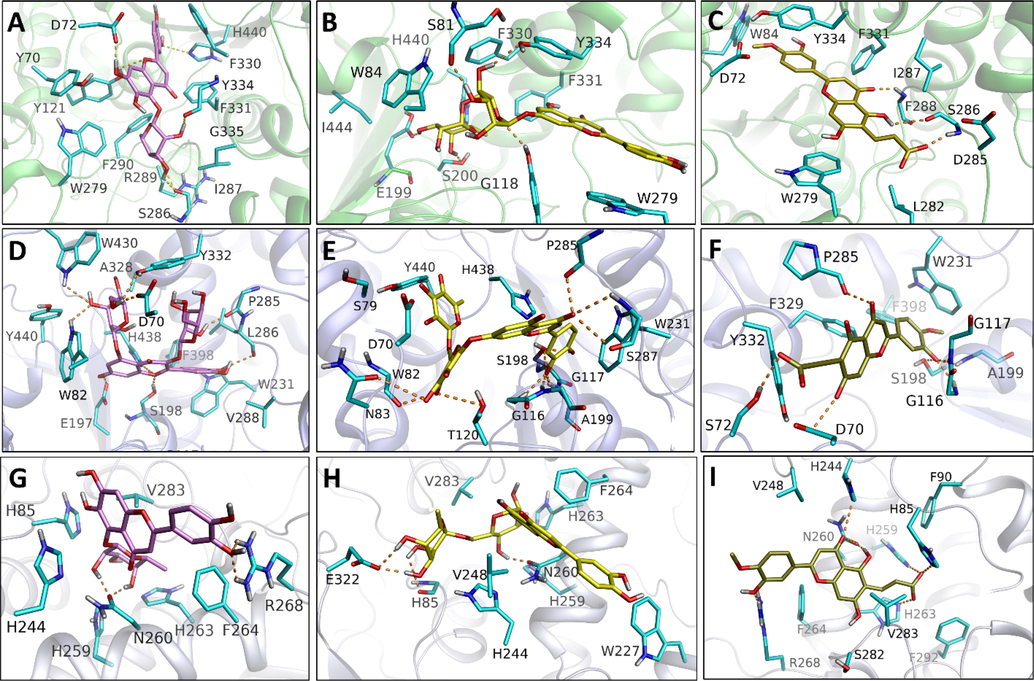
Docking generated complexes of proteins bonded to their tested compounds: (A) AChE-HLR, (B) AChE-LRG, (C) AChE-TFD, (D) BChE-HLR, (E) BChE-LRG, (F) BChE-TFD, (G) TYR-HLR, (H) TYR-LRG, (I) TYR-TFD.
3.5 Toxicological studies
The PN-M extract was subjected to toxicological studies to assess the plant's toxic effects, including different hematological, biochemical, and physical parameters. The effect of oral administration of methanol extract of P. nodiflora on the haematological parameters of chicks is shown in Table 6, while the effect of oral administration of methanol extract of P. nodiflora on the biochemical parameters of chicks is shown in Table 7; likewise, the effect of oral administration of methanol extract of P. nodiflora on body organs of chicks is shown in Table 8. Broiler chicks were selected and distributed in four experimental groups. They were orally fed with the extract for 21 days. The chicks were physically evaluated, and the number of chicks remained the same with no abnormality. After comparing the weights of different groups, no significant variation was found. On day 21, absolute and relative weights of other body organs, including kidney, heart, spleen, trachea, liver, intestine, thymus, and lungs, were also done with no significant variation. Hematological and biochemical parameters were evaluated after collecting blood samples on days 7, 14, and 21. On account of statistical analysis, P. nodiflora was non-toxic, as there was no significant difference in experimental values to that of the control group. Overall, we have used the methanolic extract (at different concentrations of 150 mg/kg, 200 mg/kg, 250 mg/kg, and 300 mg/kg) of P. nodiflora plant in order to check the in-vivo toxicity. The results have indicated that the tested extract was found safer even at the higher doses of 300 mg/kg.
Parameters
Group (ZP)
Control
150 mg/kg
200 mg/kg
250 mg/kg
300 mg/kg
Erythrocyte counts
Day 7
2.5 ± 0.12
2.63 ± 0.08
2.43 ± 0.08
2.48 ± 0.08
2.74 ± 0.07
Day 14
2.60 ± 0.13
2.46 ± 0.07
2.70 ± 0.02
2.43 ± 0.04
2.48 ± 0.05
Day 21
2.40 ± 0.07
2.54 ± 0.07
2.74 ± 0.07
2.73 ± 0.07
2.56 ± 0.05
Leucocyte count (103/ul)
Day 7
197.38 ± 3.62
193.9 ± 2.45
190.8 ± 5.05
195.3 ± 4.31
222.2 ± 5.30
Day 14
202.8 ± 6.11
196.1 ± 3.73
193.5 ± 4.25
213.0 ± 10.9
233.5 ± 3.75
Day 21
213.3 ± 16.3
221.3 ± 3.82
198.1 ± 4.90
224.7 ± 2.81
229.4 ± 6.31
Lymphocyte count (103 / µ L)
Day 7
168.98 ± 2.85
168.90 ± 2.60
214.10 ± 3.64
174.90 ± 11.89
167.50 ± 3.48
Day 14
167.6 ± 3.28
170.4 ± 1.67
211.0 ± 2.23
215.0 ± 0.89
216.2 ± 0.71
Day 21
213.3 ± 16.3
209.8 ± 3.79
210.8 ± 2.67
162.1 ± 5.18
159.7 ± 2.58
Haemoglobin (gm/dl)
Day 7
10.14 ± 0.37
10.50 ± 0.28
11.36 ± 0.49
11.40 ± 0.18
10.14 ± 0.47
Day 14
10.28 ± 0.63
10.34 ± 0.37
9.46 ± 0.26
11.28 ± 0.28
10.8 ± 0.29
Day 21
10.34 ± 0.37
10.54 ± 0.34
9.58 ± 0.22
11.43 ± 0.43
11.10 ± 0.27
MCV (FL)
Day 7
125.10 ± 0.96
123.90 ± 1.33
137.24 ± 1.07
129.20 ± 2.90
124.80 ± 1.89
Day 14
124.10 ± 1.18
116.80 ± 0.77
134.20 ± 1.08
136.70 ± 1.78
136.90 ± 1.78
Day 21
136.30 ± 1.91
137.3 ± 2.73
132.7 ± 1.57
135.0 ± 1.49
137.0 ± 1.15
MCHC
Day 7
31.60 ± 0.29
31.20 ± 0.14
38.20 ± 2.85
40.80 ± 0.57
42.20 ± 1.06
Day 14
29.86 ± 1.90
31.50 ± 0.40
37.40 ± 2.72
38.30 ± 1.90
41.10 ± 0.73
Day 21
30.10 ± 0.28
31.70 ± 0.18
40.40 ± 0.62
41.20 ± 0.53
41.60 ± 0.57
HCT (%)
Day 7
31.48 ± 0.23
31.66 ± 0.65
30.98 ± 1.10
33.58 ± 2.41
36.74 ± 1.23
Day 14
31.90 ± 1.49
30.74 ± 0.87
29.92 ± 0.33
29.64 ± 1.53
37.60 ± 1.12
Day 21
32.10 ± 0.91
32.12 ± 0.68
30.0 ± 0.49
31.90 ± 0.77
32.80 ± 2.03
Biochemical parameters
Control
Plant extract
150 mg/kg
200 mg/kg
250 mg/kg
300 mg/kg
Alkaline Phosphatase
2650.0 ± 111.10
2637.2 ± 144.50
3066.5 ± 153.60
2345.0 ± 141.90
2012.5 ± 141.90
Aspartate Aminotransferase
6.0 ± 0.47
8.50 ± 1.44
10.50 ± 0.64
8.75 ± 0.85
7.0 ± 0.85
Alanine Aminotransferase
149.4 ± 13.44
174.50 ± 7.90
174.25 ± 7.43
183.75 ± 6.20
160.0 ± 7.43
Urea level
14.0 ± 1.08
21.0 ± 2.08
17.0 ± 1.88
18.0 ± 0.85
14.0 ± 0.85
Creatinine
0.25 ± 0.01
0.18 ± 0.03
0.34 ± 0.03
0.39 ± 0.02
0.40 ± 0.04
Parameters/Day
Group (PN)
Control
150 mg/kg
200 mg/kg
250 mg/kg
300 mg/kg
Liver weight
Absolute
53.11 ± 1.70
52.62 ± 1.83
44.73 ± 0.85
37.09 ± 1.32
35.30 ± 1.37
Relative
2.10 ± 0.08
2.33 ± 0.13
1.88 ± 0.02
1.67 ± 0.04
1.68 ± 0.05
Kidney weight
Absolute
4.03 ± 0.01
12.4 ± 0.94
3.73 ± 0.43
4.16 ± 0.26
3.5 ± 0.20
Relative
0.48 ± 0.03
0.55 ± 0.05
0.15 ± 0.01
0.18 ± 0.009
0.16 ± 0.003
Lung’s weight
Absolute
4.03 ± 0.01
11.36 ± 0.82
4.43 ± 0.08
3.73 ± 0.26
3.73 ± 0.31
Relative
0.41 ± 0.03
0.17 ± 0.008
0.16 ± 0.01
0.18 ± 0.002
0.50 ± 0.02
Thymus weight
Absolute
4.59 ± 0.54
5.36 ± 0.31
3.66 ± 0.17
4.4 ± 0.36
4.1 ± 0.11
Relative
0.18 ± 0.02
0.19 ± 0.005
0.19 ± 0.01
0.15 ± 0.006
0.23 ± 0.007
Intestine weight
Absolute
143.47 ± 10.7
141.7 ± 18.50
146.6 ± 5.89
149.0 ± 13.01
132.2 ± 13.11
Relative
5.62 ± 0.28
6.85 ± 1.51
6.64 ± 0.29
6.30 ± 0.58
5.89 ± 0.82
Heart weight
Absolute
10.44 ± 0.76
10.60 ± 0.85
8.53 ± 0.54
10.96 ± 0.74
8.56 ± 0.17
Relative
0.41 ± 0.03
0.40 ± 0.01
0.49 ± 0.03
0.36 ± 0.02
0.47 ± 0.05
Spleen weight
Absolute
3.27 ± 0.03
1.97 ± 0.16
2.35 ± 0.02
1.64 ± 0.14
1.36 ± 0.15
Relative
0.13 ± 0.002
0.08 ± 0.005
0.09 ± 0.005
0.07 ± 0.007
0.06 ± 0.004
Trachea weight
Absolute
3.43 ± 0.31
3.65 ± 0.43
2.51 ± 0.08
2.59 ± 0.17
2.46 ± 0.26
Relative
0.13 ± 0.010
0.16 ± 0.020
0.10 ± 0.004
0.11 ± 0.008
0.11 ± 0.006
Bursa weight
Absolute
1.17 ± 0.01
1.19 ± 0.005
1.77 ± 0.18
1.07 ± 0.08
1.04 ± 0.07
Relative
0.04 ± 0.002
0.05 ± 0.001
0.07 ± 0.007
0.04 ± 0.004
0.05 ± 0.005
4 Conclusion
Based on the results of current study, methanolic and dichloromethane extracts of P. nodiflora exhibit varying inhibitory effects on enzymes and antioxidant properties. More precisely, the methanol extract showed better antioxidant activity than the dichloromethane extract. In addition, weak inhibiting propensities were exhibited by the studied extracts against the screened enzymes. Significant quantities of TPC and TFC were yielded from the extracts. UHPLC-MS and HPLC-PDA analysis conform to the presence of several important phenolics, flavonoids, terpenes, and glycosides. The molecular docking highlighted the interaction between the identified phytochemicals and tested enzymes. In acute toxicity studies, oral doses of methanolic extract did not result in death or adverse effects on general behavior. Almost no significant changes were observed in any of the hematological and biochemical parameters of the animals. This medicinal plant was found to be effective as an enzyme inhibitor and antioxidant, making it a possible option for further advanced research into its pharmacological capabilities.
Acknowledgment
This research was funded by Scientific Research Deanship at University of Hail- Saudi Arabia through project number RG-23076.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Acquaviva, A., S. C. Di Simone, Nilofar, A. Bouyahya, G. Zengin, L. Recinella, S. Leone, L. Brunetti, A. I. Uba and O. Guler (2023). “Screening for Chemical Characterization and Pharmacological Properties of Different Extracts from Nepeta italica.” Plants 12(15): 2785.
- Anti-inflammatory and antinociceptive activities of Lippia nodiflora Linn. Die Pharm.-Int. J. Pharma. Sci.. 2004;59(4):329-330.
- [Google Scholar]
- Anti-inflammatory and antibacterial activities of Lippia nodiflora and its effect on blood clotting time. J. Thi-Qar Sci.. 2013;4(1)
- [Google Scholar]
- Pharmacological and therapeutic effects of Lippia nodiflora (Phyla nodiflora) IOSR J. Pharm.. 2019;9(8):15-25.
- [Google Scholar]
- AO, T. and B. SL (2012). “EFFECTS OF LIPPIA NODIFLORA EXTRACTS ON MOTOR COORDINATION, EXPLORATORY BEHAVIOR PATTERN, LOCOMOTOR ACTIVITY, ANXIETY AND CONVULSIONS ON ALBINO MICE.” Asian Journal of Pharmaceutical and Clinical Research 4(3): 133-138.
- Hepatoprotective and antibacterial activity of Lippia nodiflora Linn. against lipopolysaccharides on HepG2 cells. Pharmacogn. Mag.. 2015;11(41):24.
- [Google Scholar]
- Antioxidant and free radical scavenging effects of Lippia nodiflora. Pharm. Biol.. 2008;46(10–11):762-771.
- [Google Scholar]
- Molecular basis of the anti-inflammatory property exhibited by cyclo-pentano phenanthrenol isolated from Lippia nodiflora. Immunol. Invest.. 2010;39(7):713-739.
- [Google Scholar]
- Antidiabetic activity of sitosterol isolated from Lippia nodiflora L. in streptozotocin induced diabetic rats. Eur. J. Pharmacol.. 2011;667(1):410-418.
- [Google Scholar]
- Antidiabetic and hypolipidemic effect of methanol extract of Lippia nodiflora L. in streptozotocin induced diabetic rats. Asian Pac. J. Trop. Biomed.. 2011;1(1):S30-S36.
- [Google Scholar]
- Plants, people, and culture: the science of ethnobotany. Scientific American Library; 1996.
- Barnabas, C., G. Gunasingh and S. Nagarajan (1980). FLAVONOIDS FROM THE FLOWERS OF PHYLA-NODIFLORA LINN, COUNCIL SCIENTIFIC INDUSTRIAL RESEARCH PUBL & INFO DIRECTORATE, NEW DELHI 110012, INDIA. 19: 822-822.
- Towards the pharmacological validation and phytochemical profiling of the decoction and maceration of Bruguiera gymnorhiza (L.) Lam.—A traditionally used medicinal halophyte. Molecules. 2022;27(6):2000.
- [Google Scholar]
- Comparative analysis of phytochemical profiles and antioxidant activities between sweet and sour wampee (Clausena lansium) fruits. Foods. 2022;11(9):1230.
- [Google Scholar]
- Antioxidant activity of phenolic extraction from different sweetpotato (Ipomoea batatas (L.) Lam.) blades and comparative transcriptome analysis reveals differentially expressed genes of phenolic metabolism in two genotypes. Genes. 2022;13(6):1078.
- [Google Scholar]
- Antiproliferative and apoptotic effects of Phyla nodiflora extracts on human breast cancer cell line. Asian Pac. J. Trop. Disease. 2014;4(3):238.
- [Google Scholar]
- Structures of human acetylcholinesterase in complex with pharmacologically important ligands. J. Med. Chem.. 2012;55(22):10282-10286.
- [Google Scholar]
- Molecular modeling studies to characterize N-phenylpyrimidin-2-amine selectivity for CDK2 and CDK4 through 3D-QSAR and molecular dynamics simulations. Mol. Biosyst.. 2016;12(4):1250-1268.
- [Google Scholar]
- Molecular simulation studies on the binding selectivity of 2-anilino-4-(thiazol-5-yl)-pyrimidines in complexes with CDK2 and CDK7. Mol. Biosyst.. 2016;12(1):145-161.
- [Google Scholar]
- Effect of ethanolic extract of Phyla nodiflora (Linn.) Greene against calculi producing diet induced urolithiasis. Indian J. Nat. Prod. Resour.. 2010;1(3):314-321.
- [Google Scholar]
- Antimicrobial and lipid peroxide scavenging activity of Lippia nodiflora (Verbenaceae) Pharmacologyonline. 2007;3:177-189.
- [Google Scholar]
- Protective activity and antioxidant potential of Lippia nodiflora extract in paracetamol induced hepatotoxicity in rats. Iran. J. Pharmacol. Ther.. 2008;7(1):83-89.
- [Google Scholar]
- Antihypertensive efficacy of Lippia nodiflora whole plant on uninephrectomized doca salt hypertensive rats. IOSR J. Pharm.. 2012;2(6):24-28.
- [Google Scholar]
- Beyond structural genomics: computational approaches for the identification of ligand binding sites in protein structures. J. Struct. Funct. Genomics. 2011;12(2):109-117.
- [Google Scholar]
- Activity of Lippia nodiflora essential oil on bacteria. New delhi: Semin Research in Ayurveda and Siddha; 1996.
- A short history of the use of plants as medicines from ancient times. CHIMIA Int. J. Chem.. 2015;69(10):622-623.
- [Google Scholar]
- Crystal structure of Agaricus bisporus mushroom tyrosinase: identity of the tetramer subunits and interaction with tropolone. Biochemistry. 2011;50(24):5477-5486.
- [Google Scholar]
- Phytochemical and pharmacological studies of Phyla Nodiflora (Verbenaceae): A review. Pak. J. Pharm. Res.. 2016;2:49-54.
- [Google Scholar]
- Scoring noncovalent protein-ligand interactions: a continuous differentiable function tuned to compute binding affinities. J. Comput. Aided Mol. Des.. 1996;10(5):427-440.
- [Google Scholar]
- Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J. Med. Chem.. 2003;46(4):499-511.
- [Google Scholar]
- Surflex-Dock 2.1: robust performance from ligand energetic modeling, ring flexibility, and knowledge-based search. J. Comput. Aided Mol. Des.. 2007;21(5):281-306.
- [Google Scholar]
- Eupafolin, a skin whitening flavonoid isolated from Phyla nodiflora, downregulated melanogenesis: Role of MAPK and Akt pathways. J. Ethnopharmacol.. 2014;151(1):386-393.
- [Google Scholar]
- Phytochemistry of Medicinal Plants. Uzbekistan and Kyrgyzstan, Springer: Medicinal Plants of Central Asia; 2013. p. :13-14.
- Tumor-specific carrier-free nanodrugs with GSH depletion and enhanced ROS generation for endogenous synergistic anti-tumor by a chemotherapy-photodynamic therapy. Chem. Eng. J.. 2021;407:127212
- [Google Scholar]
- Li, C., B. Shi, X. Li, W. e. Zhang and X. Pan (2022). “Changes of phenolics contents, antioxidant activities, and enzyme activities in pellicles of Juglans sigillata Dode during fruits development.” International Journal of Food Properties 25(1): 2133-2145.
- Multicomponent pattern and biological activities of seven Asphodeline taxa: potential sources of natural-functional ingredients for bioactive formulations. J. Enzyme Inhib. Med. Chem.. 2017;32(1):60-67.
- [Google Scholar]
- Antimicrobial studies on selected medicinal plants (Coleus amboinicus, Phyla nodiflora and Vitex negundo) Asian J. Pharma. Technol.. 2011;1(2):53-55.
- [Google Scholar]
- Tryptophan-derived butyrylcholinesterase inhibitors as promising leads against Alzheimer's disease. Chem. Commun.. 2019;55(26):3765-3768.
- [Google Scholar]
- Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins Struct. Funct. Bioinf.. 2004;55(2):383-394.
- [Google Scholar]
- Evaluation of antibacterial activity of methanolic extract of seeds of PHYLA NODIFLORA LINN. Microbiology. 2005;43(2):295-300.
- [Google Scholar]
- “Toxicity and Teratogenicity Effects of Aqueous Leaf Extract of Phyla nodiflora in Zebrafish (Danio rerio) Embryos”. Egypt. Acad. J. Biol. Sci. B Zool.. 2022;14(2):439-447.
- [Google Scholar]
- Studies on the elemental Composition and antifungal activity of medicinal plant Lippia nodiflora L. against skin fungi. J. Pak. Assoc. Dermatal.. 2005;15:113-118.
- [Google Scholar]
- Pizzino, G., N. Irrera, M. Cucinotta, G. Pallio, F. Mannino, V. Arcoraci, F. Squadrito, D. Altavilla and A. Bitto (2017). “Oxidative stress: harms and benefits for human health.” Oxidative medicine and cellular longevity 2017.
- Powell, M. J. (1978). A fast algorithm for nonlinearly constrained optimization calculations. Numerical analysis, Springer: 144-157.
- Neuropharmacological investigation, ultra-high performance liquid chromatography analysis, and in silico studies of Phyla nodiflora. J. Physiol. Pharmacol.. 2021;72(4)
- [Google Scholar]
- Halleridone and Hallerone from Phyla nodiflora as taxonomic markers. Biochem. Syst. Ecol.. 2000;28(9):905-906.
- [Google Scholar]
- “Antidandruff activity of Eclipta alba (l.) Hassk. and Lippia nodiflora linn”. Int. J. Pharma. Phytopharmacol. Res. 2015
- [Google Scholar]
- Pharmacological, phytochemical and in-vivo toxicological perspectives of a xero-halophyte medicinal plant: Zaleya pentandra (L.) Jeffrey. Food Chem. Toxicol.. 2019;131:110535
- [Google Scholar]
- A review on Phyla nodiflora Linn.: a wild wetland medicinal herb. Int. J. Pharmaceut. Sci. Rev. Res.. 2013;20(1):57-63.
- [Google Scholar]
- Study of phytochemical and diuretic potential of methanol and aqueous extracts of aerial parts of Phyla nodiflora Linn. Int. J. Pharm. Pharm. Sci.. 2009;1(1):85-91.
- [Google Scholar]
- Steroidal constituents from the aerial parts of Lippia nodiflora Linn. Nat. Prod. Res.. 2009;23(5):436-441.
- [Google Scholar]
- Unveiling the phytochemical and pharmacological potential of Mimusops maxima (Poiret) Vaughan-an endemic plant with potential therapeutic effects. Process Biochem.. 2023;132:157-165.
- [Google Scholar]
- Neuropharmacological activity of Lippia nodiflora Linn. Pharmacogn. Res.. 2011;3(3):194.
- [Google Scholar]
- Luffa acutangula and Lippia nodiflora leaf extract induces growth inhibitory effect through induction of apoptosis on human lung cancer cell line. Biomed. Prev. Nutr.. 2012;2(4):287-293.
- [Google Scholar]
- Total phenolic and flavonoid contents, antioxidant, antidiabetic and antiplasmodial activities of Garcinia forbesii King: A correlation study. Arab. J. Chem.. 2022;15(2):103541
- [Google Scholar]
- Yen, F.-L., M.-C. Wang, C.-J. Liang, H.-H. Ko and C.-W. Lee (2012). “Melanogenesis inhibitor (s) from Phyla nodiflora extract.” Evidence-Based Complementary and Alternative Medicine 2012.
- Extracts of M. pulegium (L.) and M. spicata (L.): Effect of extraction conditions on phenolics and flavonoids contents and their antioxidant power. Egypt. J. Chem.. 2021;64(3):1447-1459.
- [Google Scholar]
- Novel perceptions on chemical profile and biopharmaceutical properties of Mentha spicata extracts: Adding missing pieces to the scientific puzzle. Plants. 2022;11(2):233.
- [Google Scholar]
- Zengin, G., M. F. Mahomoodally, E. Yıldıztugay, S. Jugreet, S. U. Khan, S. Dall’Acqua, A. Mollica, A. Bouyahya and D. Montesano (2022). “Chemical Composition, Biological Activities and In Silico Analysis of Essential Oils of Three Endemic Prangos Species from Turkey.” Molecules 27(5): 1676.
- Zengin, G., Á. Fernández-Ochoa, M. d. l. L. Cádiz-Gurrea, F. J. Leyva-Jiménez, A. Segura-Carretero, F. Elbasan, E. Yildiztugay, S. Malik, A. Khalid and A. N. Abdalla (2023). “Phytochemical Profile and Biological Activities of Different Extracts of Three Parts of Paliurus spina-christi: A Linkage between Structure and Ability.” Antioxidants 12(2): 255.
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105233.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







