Translate this page into:
An extended strategy of “Recursive Tree” for characterization of drug metabolites in vivo and in vitro and actional mechanism study based on network Pharmacology: Formononetin as a study case
⁎Corresponding author at: School of Traditional Chinese Medicine, Binzhou Medical University, Yantai, 264003, Shandong, China. zhangjiayu0615@163.com (Jiayu Zhang)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
The screening and identification of drug metabolites in biological matrices is challenging, and ultra-high performance liquid chromatography-Q-Exactive Orbitrap mass spectrometry (UHPLC-Q-Exactive Orbitrap MS) has become a powerful technological tool for drug metabolites analysis due to its high sensitivity. However, the spectral information contained in existing chemical standards and databases is very limited, and the UHPLC-Q-Exactive Orbitrap MS technique alone cannot satisfy the identification of complex and diverse metabolites. Therefore, there is an urgent need for a new strategy to achieve comprehensive drug metabolic profile. Based on this, we have innovatively constructed a “recursive tree” analysis strategy and bridged it with network pharmacology for elucidating the pharmacological mechanisms of drugs. In this paper, we investigated the overall metabolic profile of formononetin as an example and utilized the primary branching metabolites of formononetin as effective ingredients for the study of the anti-NAFLD mechanism. The results showed that a total of 131 metabolites (prototype drug included) were detected and identified. Among them, 106 metabolites were found in rats and 31 metabolites were found in liver microsomes. Glucose conjugation, demethylation, sulfation, glucuronidation, and their complex reactions were the major processes of formononetin biotransformation. Network pharmacology results screened 104 potential targets and 20 major signaling pathways. Their mechanisms may be additive and/or synergistic effects. In addition, the therapeutic effects of formononetin against NAFLD were investigated based on palmitic acid / oleic acid-induced HepG2 cells. In summary, the recursive tree analysis strategy provides a convenient method for the identification of metabolites, and its seamless integration with network pharmacology lays the foundation for studying the pharmacological activities of natural products.
Keywords
Recursive tree analytical strategy
Metabolism
Network pharmacology
Formononetin
NAFLD
UHPLC-Q-Exactive Orbitrap MS
- NAFLD
-
anti-nonalcoholic fatty liver disease
- DPIs
-
diagnostic product ions
- PRM
-
parallel reaction monitoring
- NLFs
-
neutral loss fragments
- SD
-
Sprague Dawley
- MgCl2
-
Magnesium chloride
- UDPGA
-
uridine 5′-diphosphoglucuronic acid trisodium salt
- NADPH
-
β-nicotinamine adenine dinucleotide phosphate
- PA/OA
-
palmitic acid / oleic acid
- BCA
-
bicinchoninic acid
- TG
-
total triglycerides
- TC
-
total cholesterol
- ALT
-
alanine transaminase
- AST
-
aspartate transaminase
- SOD
-
superoxide dismutase
- GSH
-
glutathione peroxidase
- CCK-8
-
Cell Counting Kit-8
- FA
-
formic acid
- RDB
-
ring double bond
- NF-κB
-
nuclear factor kappa-B
- AKT1
-
AKT serine/threonine kinase 1
- FoxO
-
forkhead box O3
- ALB
-
albumin
- MAPK
-
mitogen-activated protein kinase
- IL-17
-
interleukin 17
- STAT3
-
signal transducer and activator of transcription 3
- PI3K
-
phosphatidylinositol-3-kinases
- EGFR
-
epidermal growth factor receptor
- Akt
-
protein-serine-threonine kinase
- mTOR
-
mammalian target of rapamycin
Abbreviations
1 Introduction
Drug metabolism is the process by which exogenous compounds undergo chemical structural changes catalyzed by various enzymes in the body, also known as biotransformation (Almazroo et al., 2017). It can transform exogenous compounds into active metabolites or toxic metabolites, generally known as the duality of biotransformation (Klotz et al., 2009). Metabolites may be the “effective form” of drugs due to their high blood concentration (Knights et al., 2016). Therefore, metabolic profiles are essential for elucidating the actional mechanism of drugs. However, hundreds of metabolites make it difficult to conduct actional mechanism studies based on all metabolites. Thus, rational selection of metabolites for mechanism studies is particularly important.
In recent years, the research on drug metabolites has made deeper development. Ultra-high performance liquid chromatography-Q-Exactive Orbitrap mass spectrometry (UHPLC-Q-Exactive Orbitrap MS) has been widely used for metabolite research with its high resolution, high sensitivity, and high accuracy (Wang et al., 2019). Moreover, the use of techniques such as neutral loss fragments (NLFs), diagnostic product ions (DPIs), and parallel reaction monitoring (PRM) has enabled high-quality mass spectrometry data acquisition, but data mining strategies for metabolites remain to be further developed (Knights et al., 2016; Kaur et al., 2020). Based on this, we first propose the “recursive tree” analysis strategy, which draws on the ideas of induction and deduction, starting from the prototype drugs and progressing step by step. In addition, network pharmacology is used to reveal the interrelationships between effective ingredients and actional targets through multidisciplinary cross-applications and integration of holistic biological networks based on bioinformatics methods (Yin et al., 2019; Das et al., 2018). Primary branching metabolites have higher blood concentrations and biological activity compared to prototypical compounds, making them most likely to be the main material basis for the treatment of disease (Cui et al., 2022; Cai et al., 2020). Therefore, the combination of these metabolites with network pharmacology can elucidate the synergistic mechanism of action between metabolites and improve the accuracy of modern pharmacological studies and the mechanism of exploration of drugs.
Formononetin (7-hydroxy-4′-methoxy isoflavone) widely distributes in Leguminosae plants, such as Glycyrrhiza uralensis Fisch (Zhao et al., 2021), Astragalus membranaceus (Liu et al., 2021), Pueraria lobata (Li et al., 2016), Trifolium pretense (Muñoz et al., 2022), and Santalum album (Machado et al., 2021). Over the past decade, formononetin has been intensively studied and many biological activities have been reported, including anti-inflammatory (Yu et al., 2022), antioxidant (Li et al., 2022), antitumor (Ma et al., 2022), vasodilatory (Oza et al., 2019), neuroprotective effects (Aly et al., 2021), and so on. It is worth mentioning that the potential anti-NAFLD effects of formononetin have been reported, but the specific mechanism and the material basis are still unclear. Several studies have investigated the pharmacokinetics of formononetin after oral administration in humans, rats, or mice (Rao et al., 2019; Guo et al., 2015; Kim et al., 2022). However, the metabolic profile remains incomplete. Due to the potential therapeutic role of formononetin in NAFLD (Scorletti et al., 2022), a deeper understanding of its metabolic fate is essential. As there is no systematic analytical strategies or techniques to mine and characterize formononetin metabolites, the screening of formononetin metabolites from complex matrix systems remains a difficult task in metabolic studies.
In this study, the formononetin was used as a case study. For the first time, the recursive tree analysis strategy was proposed and combined with multiple data processing tools to quickly screen and identify the metabolites of formononetin in vivo and in vitro. What’s more, the primary branching metabolites of formononetin were bridged with network pharmacology, and the actional mechanism of formononetin against NAFLD was initially elucidated.
2 Materials and methods
2.1 Chemicals and reagents
Formononetin reference (Batch number: MUST-21033005, purity ≥ 99.03 %) and Ononin reference (Batch number: MUST-21080311, purity ≥ 98.48 %) were provided by Chengdu Must Biotechnology Co., Ltd. (Sichuan, China). HepG2 cells was obtained from Binzhou medical university (Yantai, China). Kits measuring the levels of bicinchoninic acid (BCA), total cholesterol (TC), total triglycerides (TG), alanine transaminase (ALT), aspartate transaminase (AST), superoxide dismutase (SOD), and glutathione peroxidase (GSH) were purchased from Jiancheng Institute of Biotechnology (Nanjing, China). Palmitic acid / oleic acid (PA/OA) were obtained Xi'an Kunchuang Technology Development Co., Ltd. (Xian, China). Oil Red O and Cell Counting Kit-8 (CCK-8) were purchased from Beyotime Biotechnology (Shanghai, China).
HPLC grade methanol, acetonitrile, and formic acid (FA) were purchased from Thermo Fisher Scientific (Fair Lawn, NJ, USA), and pure water for analysis was purchased from Watson Group Co., Ltd. (Jinan, China). All the other chemicals of analytical grade were available at the workstation, Shandong Academy of Chinese Medicine (Jinan, China). Grace PureTM SPE C18-Low solid phase extraction cartridges (200 mg·3 mL−1, 59 μm, 70 Å) were purchased from Grace Davison Discovery Science (Deerfield, IL, USA). Male Sprague Dawley (SD) rat liver microsomes (1 mL, Batch number: 20210305), Magnesium chloride (MgCl2), β-nicotinamine adenine dinucleotide phosphate (NADPH), and uridine 5′-diphosphoglucuronic acid trisodium salt (UDPGA) were all purchased from NEWGAINBIO Co., Ltd. (Wuxi, China).
2.2 In vivo animal experiment
2.2.1 Animals and drug administration
Eight male SD rats (200 ± 10 g) were acquired by Jinan Pengyue Experimental Animal Technology Co., Ltd. (Jinan, China). They were free to drink and eat at humidity (70 ± 5 %) and constant temperature (25 ± 2 °C) for 7 days. The rats were randomly divided into two groups including Formononetin Group (n = 6) and Control Group (n = 6) for test plasma, urine, faeces, and liver, respectively. They were fasted for 12 h with free access to water before the experiment. Formononetin was dissolved in 0.9 % saline solution. The rats in the formononetin group and the control group were orally administered with formononetin solution (150 mg·kg−1, 10 mL·kg−1) and 0.9 % saline solution, respectively. Pentobarbital sodium was used for anesthesia. The animal facilities and protocols were complied with the Guide for the Care and Use of Laboratory Animals (USA National Research Council, 1996). The animal experiment was approved by the Animal Care and Use Committee at Binzhou Medical University (2021–085).
2.2.2 Sample collection and pretreatment
Plasma samples were prepared by collecting blood samples from the infraorbital venous plexus within 24 h and centrifuging at 3,500 rpm for 10 min. Urine and faeces samples were collected from the metabolic cage within 24 h. Furthermore, liver samples were collected and quenched in liquid nitrogen from dissected rats. Finally, the collected homogeneous samples were mixed. (Li et al., 2022).
The main method of sample pretreatment is the SPE cartridges purification method. To obtain more metabolites, methanol precipitation, and acetonitrile precipitation methods were added to the pretreatment of plasma samples. Specific research methods were described in the team’s previous research. (Li et al., 2022; Dong et al., 2021; Wang et al., 2022; Jing et al., 2022).
2.3 In vitro liver microsomes incubation
Firstly, incubation mixtures were prepared in PBS buffer and divided into Drug Group and the Control Group. The buffer for the Drug Group contained rat liver microsomes (1 mg·mL−1), formononetin (0.1 mg·mL−1), and MgCl2 (3 mM, final concentration). The buffer for the Control Group did not contain formononetin. 900 μL of the above incubation mixture, 100 μL of NADPH (25 mg·mL−1) and 100 μL of UDPGA (25 mg·mL−1) were added to each of the 6-well plates to start the reaction. The reaction was incubated at 37 °C for 5, 10, 15, 30, 45, 60, 120, 240, and 480 min, and then 100 μL of incubation solution was removed and the reaction was terminated by adding 200 μL of cold acetonitrile. Finally, the supernatant was purified by acetonitrile precipitation.
2.4 Instruments and analytical conditions
UHPLC analysis was conducted on the Dionex Ultimate 3000 UHPLC system (Thermo Fisher Scientific, MA, USA). Separation was performed on a Waters ACQUITY UPLC BEH C18 column (2.1 × 100 mm, 1.7 μm). The column temperature was kept at 30℃, the flow rate was 0.3 mL·min−1. The mobile phase was composed of water containing 0.1 % formic acid (A) and acetonitrile (B). The elution gradient condition was as follows: 0–5 min, 5 %-30 % B; 5–10 min, 30 %-50 % B; 10–27 min, 50 %-90 % B; 27–27.1 min, 90 %-5% B; 27.1–30 min, 5 % B.
HRMS and MS/MS spectra were obtained using Q-Exactive Focus Orbitrap MS (Thermo Fisher, Waltham, MA, USA) equipped with HESI. The ion source parameters were set as follows: Scan range, m/z 80–1,200; Aux gas flow rate, 10 arbitrary units; Aux gas heater temperature, 320 °C; Sheath gas flow rate, 45 arbitrary units; Capillary temperature, 320 °C; Spray voltage, 3.8/3.5 kV (+/-); Scan modes, full MS resolution, 70,000; dd-MS2 resolution, 17,500; Collision energy, 15 %, 30 %, 45 %.
2.5 Data processing and peak selections
In order to obtain accurate metabolites of formononetin, the parameters of the Thermo Xcalibur 2.1 workstation were set within a reasonable range. The compositional structural formulae were set to C[0–30], N[0–5], O[0–15], H[0–60], S[0–1]. The mass error range was set to within ± 5 ppm. The ring double bond (RDB) equivalent was set to [0–15].
2.6 Cells and treatments
Logarithmic growth phase HepG2 cells were inoculated into 96-well plates at a density of 5 × 103 per well for cell viability. Subsequently, HepG2 cells were seeded into 6-well plates at a density of 2 × 105 per well. The cells were stimulated with PA (187.5 μM) / OA (375 μM) and formononetin (5, 10, and 50 μg·mL−1) for 24 h. After fixation with 4 % paraformaldehyde, Oil Red O staining was added for 20 min, and the images were captured using an Echo Laboratories RevolveFL imaging system (San Diego, USA). Finally, HepG2 cells were collected by inoculating 6-well plates in the same way to determine protein content. The levels of TC, TG, AST, ALT, SOD, and GSH were also determined according to the kit instructions.
2.7 Network pharmacology
2.7.1 Targets prediction and screening common targets
Formononetin and its primary branch metabolites were subjected to a SMILE search in the PubChem database and relevant targets were predicted using the Swiss Target Database. NAFLD-related targets were then searched for in the GeneCards database by using the term “nonalcoholic fatty liver disease”. Finally, the targets of formononetin and its primary branch metabolites and NAFLD were mapped by Venny 2.1 software to screen for overlapping targets.
2.7.2 Construction of protein–protein interaction (PPI) network
Overlapping targets were analyzed with the String Version 11.0 platform to establish relationships between target proteins. The PPI network diagram was constructed using Cytoscape 3.9.1 for topological analysis. Core targets in the network were visualized based on degree values.
2.7.3 Biological function and pathway enrichment analyses
Using the Metascape database, the top 20 relevant biological processes and KEGG pathways were selected as key pathways for the main targets of formononetin and its primary branch metabolites for the treatment of NAFLD.
2.7.4 Construction of disease-pathways-targets-metabolites network
Diseases, major pathways, metabolites and key targets were linked together using Cytoscape 3.9.1 software. Visual analysis was performed by adjusting parameters. The disease-pathways-targets-metabolites network diagram was constructed to visually express their interactions.
2.8 Statistical analysis
Cell experiments were repeated at least three times. GraphPad Prism 8.0 was used to analyze the results of biochemical indices and related graphs. Two-tailed Student’s t-test was used for statistical analysis. P < 0.05 was considered significant.
3 Results
3.1 The establishment of a recursive tree-based analytical strategy
We focused on a “recursive tree” analysis strategy to systematically screen and characterize the metabolites of drugs, using formononetin as an example (Fig. 1). Firstly, we defined the concept of “recursive tree”. The drug is compared to a tree trunk, the metabolites resulting from the addition or subtraction of a single functional group are the primary branches (main branches), the metabolites resulting from the addition or subtraction of a single functional group from the primary branches are the secondary branches (lateral branches), and so on. Among the addition and subtraction of a single functional group include methylation (demethylation), hydroxylation (dehydroxylation), hydrogenation (dehydrogenation), methoxylation (demethoxylation), decarbonylation, decarbonization, etc. During the recursion process, two noteworthy points aroused our interest. One was the emergence of duplicate metabolites as the recursion progresses, and it was essential to remove duplicate metabolites when summarizing. Secondly, after the screening of the tertiary branching metabolites, fewer and fewer metabolites could be identified, so we mainly performed the identification of the primary, secondary, and tertiary branching metabolites.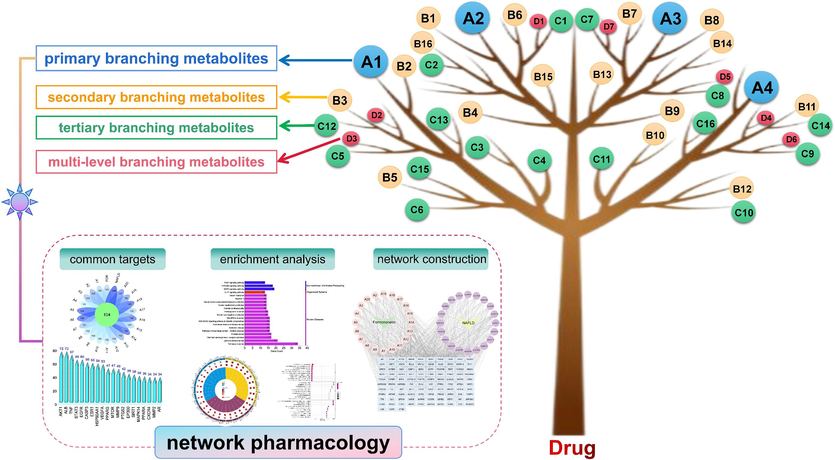
The summary diagram of recursive tree analysis strategy.
Not to be ignored, the diagnostic product ions (DPIs) played a key role in the “recursive tree” analysis strategy. The structural backbone of the prototype drug is largely preserved during metabolism, thus allowing rapid and precise structural identification of the metabolite using DPIs based on the mass fragmentation behavior of atomoxetine. In addition, the seamless integration of primary branching metabolites, which are used as effective forms of drugs due to their higher blood concentrations and well-defined structural formulas, with network pharmacology is key to the application of “recursive tree” analysis strategies.
3.2 Dpis construction based on formononetin mass cleavage pattern
Formononetin belongs to the class of isoflavones, and the DPIs are mainly produced by cleavage at different positions on the C-ring. It includes Retro-Diels-Alder (RDA) cleavage, deA-ring cleavage, and deB-ring cleavage, often accompanied by neutral loss of CO2, CO, and H2O. The prototype with the chemical formula C16H12O5 was rapidly identified based on the retention time of formononetin standard at 9.65 min and the ions at m/z 267.06 ([M−H]-) and m/z 269.08 ([M + H]+). In order to more conveniently analyze the metabolites of formononetin, we list its six cleavage pathways (Fig. 2). In the negative ion mode, four typical ions with m/z 251, m/z 239, m/z 223, and m/z 209 were obtained after the sequential loss of O, CO, CO2, and CH2, respectively. Three characteristic ions with m/z 252, 236, and 208 were generated after the successive loss of CH3, OCH3, and CO, respectively. Moreover, six product ions were observed at m/z 107, 159, 146, 120, 135, and 132 through the cleavage pathway (a)\(b)\(c). Among them, m/z 135 and m/z 132 were typical RDA cleavage ions. According to the cleavage pathways (d)\(e)\(f), the ions were produced at m/z 119, 147, 91, 175, 159, and 107. Among them, m/z 91 and m/z 175 were the key ions of flavonoid cleavage mode II, and the product ion with high abundance at m/z 161 was produced by demethylation of m/z 175. Thus, the DPIs of formononetin were summarized as m/z 91, 107, 119, 120, 132, 135, 146, 147, 159, 175, 208, 209, 223, 236, 239, 251, 252 in negative ion mode. Similarly, the DPIs in positive ion mode were m/z 93, 109, 121, 122, 134, 137, 148, 149, 161, 177, 210, 211, 225, 238, 241, 253, 254.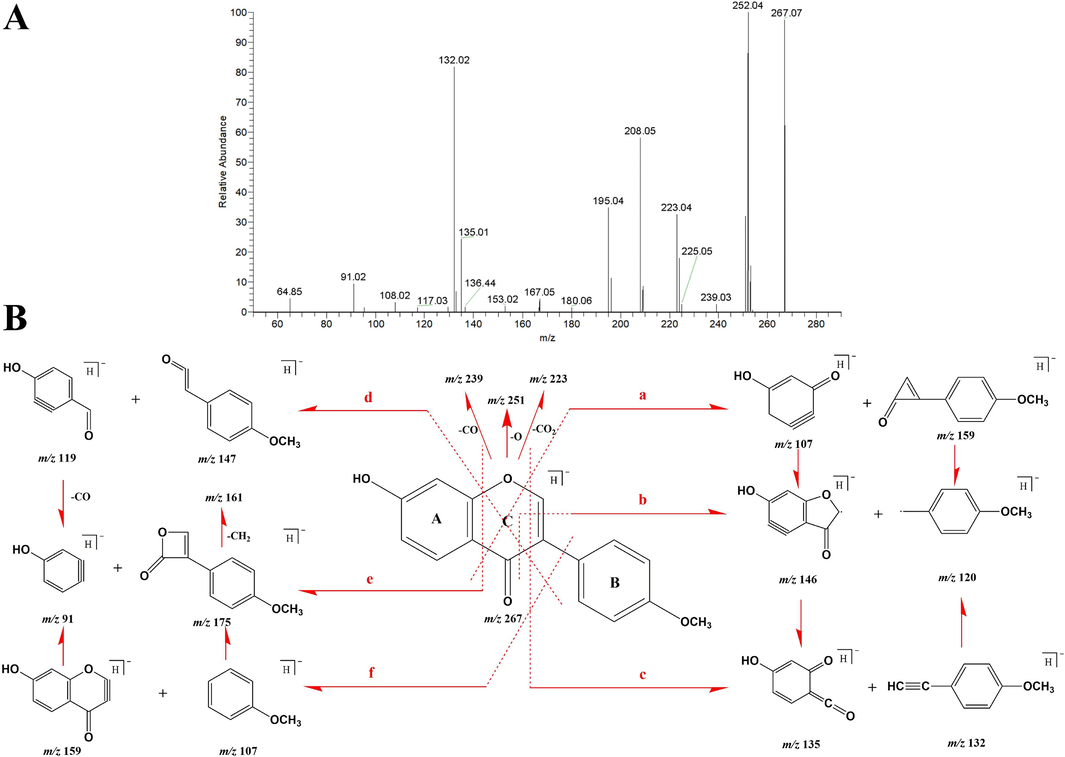
ESI-MS spectrum (A) and typical cleavage pathways (B) of formononetin in negative ion mode.
3.3 Dpis construction of primary, secondary, tertiary, and multi-level branching metabolites
Based on the “recursive tree” analysis strategy and the general rules of drug metabolism, formononetin was susceptible to methylation (demethylation), oxidation (deoxidation), glucuronidation, sulfation, and their composite reactions. With the addition or subtraction of a single functional group in the parent nucleus as the core of the analysis strategy, we classified these metabolites into primary, secondary, tertiary, and multi-level branching metabolites, which resulted in the generation of regular DPIs. For example, daidzein, a demethylated metabolite of formononetin (primary branching metabolite), might yield negative ions at m/z 253 (m/z 267 - CH2), m/z 237 (m/z 251 - CH2), m/z 215 (m/z 239 - CH2), m/z 209 (m/z 223 - CH2), m/z 145 (m/z 159 - CH2), m/z 118 (m/z 132 - CH2), m/z 161 (m/z 175 - CH2), and m/z 93 (m/z 107 - CH2). Another one, glucuronidation metabolites of daidzein (secondary branching metabolite) might produce negative ions at m/z 429 (m/z 253 + GluA), m/z 413 (m/z 237 + GluA), m/z 391 (m/z 215 + GluA), m/z 385 (m/z 209 + GluA), m/z 321 (m/z 145 + GluA), m/z 294 (m/z 118 + GluA), m/z 337 (m/z 161 + GluA), and m/z 269 (m/z 93 + GluA). By analogy, the DPIs ± nX [n indicated the presence of multiple functional groups, X indicated molecular weight of substituents, e.g. 14 (CH2), 16 (O), 18 (H2O), 30 (OCH2), 80 (SO3), 176 (GluA), etc.] could be used to identify formononetin metabolites if multi-level branching reactions occurred.
3.4 Identification of formononetin metabolites
This study identified 131 metabolites of formononetin, with 53, 27, 46, 9, and 31 metabolites detected in plasma, urine, faeces, liver, and liver microsomes samples, respectively. Based on the “recursive tree” analysis strategy, we divided the metabolites into four groups, including primary, secondary, tertiary, and multi-level branching metabolites group. Table 1 provided detailed information on the formononetin metabolites. Fig. 3 illustrated the structure of the metabolites. Representative metabolites of formononetin were listed and the identification analysis of all metabolites was detailed in the Supplementary Material.
ID
Ion
Mode
tR/
min
Formula
H]-/
[M + H]+
Theoretical
Mass (m/z)
Experimental
Mass (m/z)
RDB
Error
(ppm)
MS/MS Fragment Ions
PS
PM
PA
U
F
L
LM
formononetin
N
9.62
C16 H11 O4
267.06519
267.06631
11.5
0.104
252(1 0 0),267(89),221(2),206(1),134(2),121(5),194(2),102(1),92(2),77(1)
+
+
+
+
+
+
+
P
9.63
C16 H13O4
269.08083
269.08026
12.5
0.156
269(1 0 0),253(46),136(24),223(5),225(8),211(5),161
(11)+
+
+
+
+
–
+
A1
P
6.92
C22H23O9
431.13409
431.13365
10.5
1.000
269(1 0 0),431(2),253(1),136(1),161(1)
–
–
–
–
–
–
+
A2
N
7.09
C15H9O4
253.04953
253.05043
11.5
−0.799
253(1 0 0),225(18),209(5),237(2),221(2)
+
–
+
–
–
–
–
P
7.09
C15H11O4
255.06518
255.06427
10.5
−0.915
255(1 0 0),199(6),137(4),227(3),91(1)
+
+
+
+
+
–
–
A3
N
8.22
C16H11O7S
347.02199
347.02319
11.5
0.269
267(1 0 0),347(33),252(31),62(2),223(2)
+
+
+
–
–
–
–
P
8.23
C16H13O7S
349.03764
349.03650
10.5
−1.150
269(1 0 0),349(31),254(4),213(3),97(3)
+
+
–
–
–
–
–
A4
N
6.97
C22H19O10
443.09727
443.09888
13.5
0.045
267(1 0 0),107(73),252(53),209(35),443(32)
+
–
+
+
–
–
–
P
6.97
C22H21O10
445.11292
445.11172
12.5
−1.873
269(1 0 0),445(10),254(2),213(1),237(1)
+
+
–
+
–
–
–
A5
P
4.81
C16H13O5
285.07575
285.07498
10.5
−0.770
285(1 0 0),137(63),270(18),149(2),63(1)
+
–
–
–
–
–
–
A6
N
6.12
C16H11O5
283.06009
283.06119
11.5
−0.024
135(1 0 0),283(58),240(4),211(2),147(1)
–
–
–
–
–
–
+
A7
N
6.81
C16H11O5
283.06009
283.06125
11.5
0.188
283(1 0 0),135(63),163(18),120(2),147(1)
–
–
–
–
–
–
+
A8
N
7.30
C16H11O5
283.06009
283.06116
11.5
−0.130
268(1 0 0),283(72),135(22),147(17),61(6)
–
–
–
+
–
–
–
A9
N
7.85
C16H11O5
283.06009
283.06097
11.5
−0.801
283(1 0 0),268(61),135(20),175(2),147(1)
–
–
–
–
–
–
+
A10
N
8.76
C16H11O5
283.06009
283.06100
11.5
−0.695
283(1 0 0),268(77),224(5),132(3),152(0.6)
–
–
–
–
–
–
+
A11
N
9.52
C16H11O5
283.06009
283.06116
11.5
−0.130
283(1 0 0),268(74),224(11),152(9),132(6)
–
–
–
–
–
–
+
A12
P
10.66
C16H13O5
285.07575
285.07477
10.5
−0.980
285(1 0 0),270(54),134(11),154(9),63(6)
–
–
–
–
–
–
+
A13
N
10.84
C17H13O4
281.08083
281.08194
11.5
0.028
253(1 0 0),281(79),223(3),134(2),232(1)
–
–
–
–
–
–
+
P
10.87
C17H15O4
283.09648
283.09564
10.5
−2.986
283(1 0 0),255(26),93(9),163(7),206(6)
–
–
–
+
–
–
–
A14
N
7.21
C16H13O4
269.08083
269.08200
10.5
0.252
269(1 0 0),225(7),253(5),135(4),209(3)
–
–
–
+
–
–
–
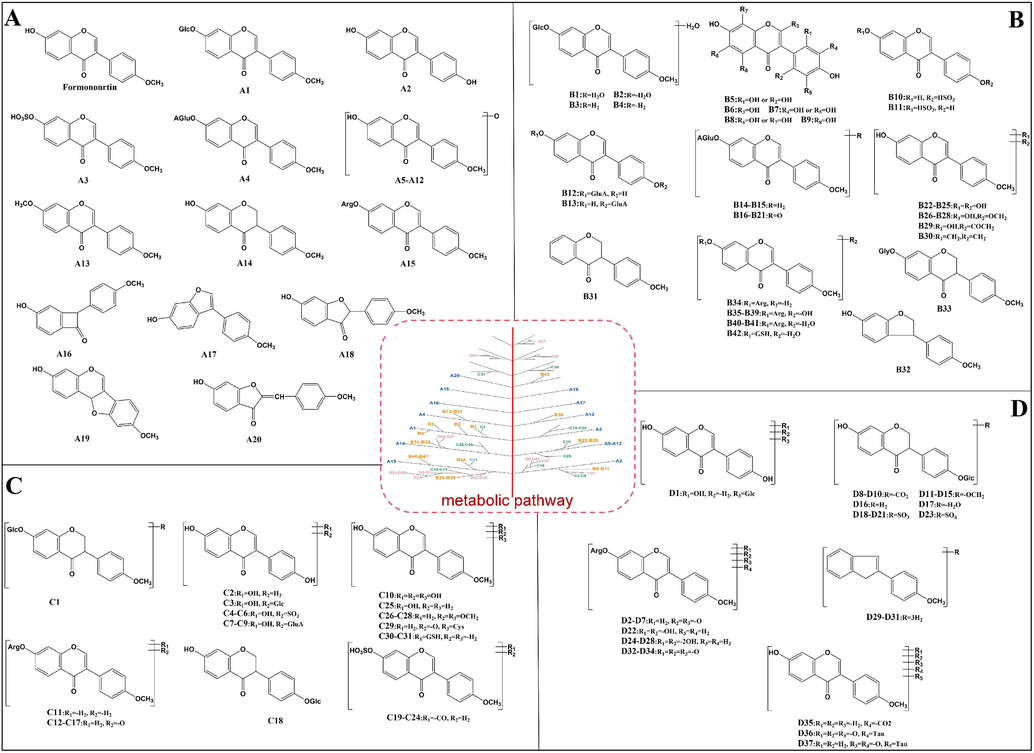
Structural identification and proposed metabolic pathway of formononetin metabolites. A: primary branching metabolites of formononetin (A1-A20); B: secondary branching metabolites of formononetin (B1-B42); C: tertiary branching metabolites of formononetin (C1-C31); D: Multi-level branching metabolites of formononetin (D1-D37).
3.4.1 Representative primary branching metabolites of formononetin (A1-A20)
The retention time of A1 in the positive ion mode was 6.92 min. Its theoretical [M + H]+ ion at 431.13365 (C22H23O9, mass error within ± 5 ppm), which was 162 Da greater than that of formononetin. In ESI-MS2 spectrum, the fragment ions at m/z 269.08 [M + H-Glc]+, m/z 253.05 [M + H-Glc-CH2]+, and m/z 237.06 [M + H-Glc-OCH2]+ were observed. Additionally, the DPIs of formononetin at m/z 107.09, m/z 136.06, and m/z 161.85 were also detected. It was inferred from the literature that A1 was ononin (Li et al., 2020).
A2 yielded significant [M−H]-/ [M + H]+ ions at 253.05043 (C15H9O4, mass error of −0.799 ppm) and m/z 255.06427 (C15H11O4, mass error of −0.915 ppm) in negative and positive ion modes, respectively. It was 14 Da less massive than formononetin. And then, the fragment ions at m/z 237.06 [M−H−O]-, m/z 225.06 [M−H−CO]-, m/z 221.07 [M−H−2O]-, and m/z 209.07 [M−H−CO2]- were produced by a series of neutral losses of O, CO, and CO2. These fragment ions confirmed that A2 was the demethylated metabolite of formononetin, daidzein.
The [M−H]- ions of A3, A4, and A15 were found at m/z 347.02319 (C16H11O7S, mass error of 0.269 ppm), m/z 443.09888 (C22H19O10, mass error of 0.045 ppm), and m/z 423.16559 (C22H23N4O5, mass error of −1.261 ppm). A3, A4, and A15 were 80 Da, 176 Da, and 156 Da larger than that of formononetin, respectively, suggesting that A3, A4, and A15 could be deduced to be sulfated metabolite, glucuronidated metabolite and arginine conjugated metabolite of formononetin, respectively. In their ESI-MS2 spectra, the significant fragment ion at m/z 267.06 was formed by the neutral loss of SO3, GluA, and Arg, respectively. Other DPIs of formononetin at m/z 91.06, m/z 107.09, m/z 135.05, m/z 147.06, m/z 209.03, m/z 223.85, and m/z 252.05 were observed. Therefore, A3, A4, and A15 were deduced as formononetin-7-O-sulfate, formononetin-7-O-GluA, and formononetin-7-O-Arg, respectively.
Eight isomeric metabolites, A5-A12 were 16 Da more massive than formononetin, and their theoretical [M−H]-/[M + H]+ ions at m/z 283.06009 (C16H11O5, mass errors within ± 5 ppm) and m/z 285.07575 (C16H13O5, mass errors within ± 5 ppm). Therefore, A5-A12 were initially identified as the mono-hydroxylated metabolites of formononetin. Among them, the MS2 spectra of A5-A9 all showed complementary fragment ions at m/z 135.01 (C7H3O3) and m/z 147.04 (C9H7O2), which were generated by cleavage pathway (c), indicating hydroxylation in the B-ring or C-ring. Notably, the pathway cleavage (b) of A7 produced fragment ions at m/z 163.00 (C8H3O4) and m/z 120.06 (C8H8O), indicating hydroxylation in the C-2 position. In general, the larger the ClogP value, the longer the retention time. ClogP values for the substituted hydroxyl groups of C-2′, C-6′, C-3′, and C-5′ were 1.11784, 1.11784, 1.48784, and 1.48784, respectively. Therefore, the hydroxyl substitution of C-2′ and C-6′ were A5 and A6 (or A6 and A5), and the hydroxyl substitution of C-3′ and C-5′ were A8 and A9 (or A9 and A8). Furthermore, A10, A11, and A12 all showed fragment ions at m/z 152.01 (C7H4O4) and m/z 132.06 (C9H8O), which were typical DPIs generated by RDA cleavage. The m/z 152.01 indicated the presence of two hydroxyl groups in the A-ring. This was also confirmed by the ions at m/z 136.06, m/z 114, m/z 91.06, and m/z 71.00 produced by the cleavage pathway (c)/(e). In other words, the hydroxyl substitution in the A-ring has three positions: C-5, C-6, and C-8. The hydroxyl group in the C-5 position readily formed an intramolecular hydrogen bond with the carbonyl group in the C-4 position, thus greatly reduced the polarity of the metabolite and its retention time. It was deduced that A12 was 5-hydroxyl-formononetin (Biochanin A). The ClogP values of A10 and A11 were both 1.57529, so it was difficult to distinguish between C-6 and C-8 substituted hydroxyl groups.
3.4.2 Representative secondary branching metabolites of formononetin (B1-B42)
B1 produced [M−H]- ion with a mass of m/z 447.13028 (C22H23O10, mass error of 1.364 ppm), which was 18 Da more than A1. In the ESI-MS2 spectrum, the fragment ions at m/z 429.12 [M−H−H2O]- and m/z 202.80 [M−H−H2O−Glc−CO2−OCH3]- were generated. Other DPIs were found at m/z 175.00, m/z 135.01, and m/z 121.06. Thus, B1 was a secondary branching metabolite resulting from the hydrolysis of A1.
Five isomeric metabolites, B5-B9 were 14 Da less massive than A5-A12. And thus,
they were tentatively inferred to be demethylated products of A5-A12. In their ESI-MS2 spectra, the DPIs at m/z 251.05 [M−H−O]-, m/z 241.88 [M−H−CO]-, and m/z 225.06 [M−H−CO2]- were obtained after the sequential loss of O, CO, and CO2, respectively. Based on the structural characteristics of A5-A12 and the ClogP values, the structures of B5-B9 were inferred (Fig. 3).
B10 and B11 afforded [M−H]- ion at m/z 333.00634 (C15H9O7S, mass errors within ± 5 ppm), which were 80 Da more massive than A2. The product ions of B10 and B11 at m/z 305.86 [M−H−CO]- and m/z 253.05 [M−H−SO3]- were observed. Furthermore., the ions of m/z 135.05 and m/z 117.93 (m/z 198-SO3) were generated in B10; m/z 214.88 and m/z 217.87 were generated in B11. The ClogP values for B10 and B11 were 0.4985 and 0.879, respectively. Therefore, B10 was daidzin-4′-O-sulfate, B11 was daidzin-7-O-sulfate.
The retention times of B12 and B13 were 4.66 and 5.31 min, respectively, with a mass 14 Da smaller than that of A4, and the [M−H]- ions produced in the negative ion mode were m/z 429.08377 and m/z 429.08231 (C21H17O10, mass errors within ± 5 ppm). Therefore, both B12 and B13 were deduced as glucuronidation and demethylation metabolites of formononetin. M/z 117.93 and m/z 293.80 (117 + GluA) were B-ring fragment ions produced by RDA cleavage of B12 and B13, respectively. It was determined that B12 was daidzin-7-O-GluA and B13 was daidzin-4′-O-GluA.
3.4.3 Representative tertiary branching metabolites of formononetin (C1-C31)
C1 showed theoretical deprotonated molecular ion at m/z 449.14703 (C22H25O10, mass error of 1.807 ppm), which was 2 Da more massive than B1. Two predominant fragment ions at m/z 405.30 [M−H−CO−H2O]- and m/z 374.43 [M−H−CO2−H2O−CH3]- were detected. Additionally, high abundance ions at m/z 447.14 (B1) and m/z 431.11 (B1-O) were generated. Based on this, C1 was identified as the hydrogenated metabolite of B1, a tertiary branching metabolite of formononetin.
C2 generated its [M−H]- ion at m/z 271.06100 (C15H11O5, mass error of 0.132 ppm) with the retention time of 7.20 min. It was 2 Da more massive than B5-B9, extrapolating that C2 was a hydrogenated metabolite of B1. The DPIs at m/z 251.05, m/z 241.88, and m/z 225.06 were observed. C3 afforded the deprotonated molecular ion at m/z 431.09869 (C21H19O10, mass error of 0.320 ppm) with a mass 162 Da larger than that of B5-B9, and generated fragment ions at m/z 269.08 [M−H−Glc]-, m/z 255.05 [M−H−Glc−CH2]-, m/z 241.88 [M−H−Glc−CO]-, and m/z 225.06 [M−H−Glc−CO2]-. Hence, C3 was characterized as a glucosylation product of B5-B9. Moreover, C4-C6 has the same theoretical [M−H]- ion at m/z 349.00238 (C15H9O8S, mass errors within ± 5 ppm), which were 80 Da more massive than B5-B9. The fragment ions at m/z 269.08 [M−H−SO3]-, m/z 241.88 [M−H−SO3−CO]-, and m/z 225.06 [M−H−CO2]- were exhibited. So, C4-C6 were speculated to be sulfated metabolites of B5-B9. Similarly, the dominant ion at m/z 269.08 [M−H−GluA]- was detected in the ESI-MS2 spectra of C7-C9. They were 176 Da more massive than B5-B9, inferring that they were glucuronided metabolites of B5-B9.
3.4.4 Representative mmulti-level branching metabolites of formononetin (D1-D37)
D1 had [M−H]- ion at m/z 433.11414 (C21H21O10, mass error of 0.120 ppm) in the ESI-MS2 spectrum. It was 2 Da less massive than C3. Besides, glucosylation reaction characterization was presented. The fragment ions at m/z 415.43 [M−H−H2O]-, m/z 271.05 [M−H−Glc]-, m/z 257.08 [M−H−Glc−CH2]-, and m/z 239.06 [M−H−Glc−CH2−H2O]- were presented. Based on the analysis above, D1 was a dehydrogenated metabolite of C3. However, it could be clearly stated that demethylation occurred at the C-4′ position.
D2-D7 with experimental [M−H]- ion at m/z 393.19211 (C22H25N4O3, mass errors within ± 5 ppm) were observed. They were 16 Da less massive than C12-C17, suggesting that D2-D7 were the deoxygenated products of C12-C17. Noteworthy ions at m/z 133.45 and m/z 104.07 (260-Arg) from RDA cleavage, and fragment ions at m/z 147.04 and m/z 246.00 from cleavage pathway (e), suggesting that arginine conjugation occurred in the C-7 position and two oxygen deletions in the C-ring, with hydrogenation of C-2 and C-3. Moreover, D22, D24-D28, and D32-D34 were also arginine conjugation metabolites with only a few structural changes. Their structures were predicted based on the results of fragment ions analysis and the DPIs of the primary and secondary branching arginine conjugation metabolites, as shown in Fig. 3. They were all defined as multi-level branching metabolites of formononetin.
3.5 Effects of formononetin on HepG2 cells induced by PA/OA
Literatures reported that PA/OA stimulation of HepG2 cells can mimic fatty hepatocytes for NAFLD studies. We successfully established a NAFLD cell model by stimulating HepG2 cells with PA (187.5 μM) / OA (375 μM). The effect of formononetin on HepG2 cells viability was assessed by CCK-8, and the results showed that cells viability was significantly reduced at formononetin concentrations greater than 100 μg·mL−1 (Fig. 4A). To ensure the anti-NAFLD efficacy of formononetin, the concentrations of 5, 10, and 50 μg·mL−1 were selected for subsequent experiments.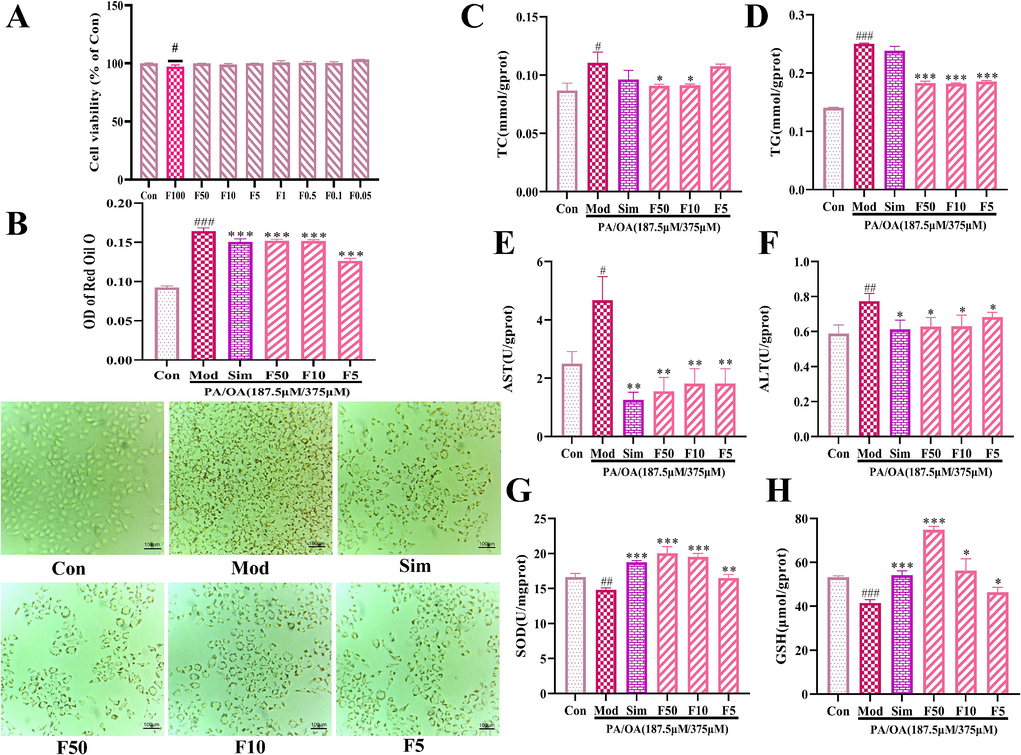
Effects of formononetin on NAFLD cell models. (A) Results of cell viability; (B) Results of Oil Red O staining (×200); (C–H) Results of TG, TC, AST, ALT, SOD, and GSH levels. #P < 0.05, ##P < 0.01, ###P < 0.001 vs the Con group; *P < 0.05, **P < 0.01, ***P < 0.001 vs the Mod group.
After 24 h of PA/OA stimulation, HepG2 cells showed severe lipid accumulation. However, the number of lipid droplets in the cells was significantly reduced after formononetin treatment (Fig. 4B), and the levels of TC and TG decreased, indicating that formononetin has a lipid-lowering effect (Fig. 4C, D). At the same time, the levels of AST and ALT were also significantly reduced (Fig. 4E, F), indicating that formononetin has hepatoprotective effects. Moreover, lipid accumulation easily leads to oxidative stress. Formononetin dose-dependently reversed the PA/OA-induced increase in SOD levels and decrease in GSH levels (Fig. 4G, H). Our data preliminarily demonstrated the ameliorative effect of formononetin in NAFLD.
3.6 Network pharmacology analysis
Primary branching metabolites play an important role in drug metabolism because they retain similar parent nucleus structures (Li et al., 2009). Moreover, in our study, the structures of the primary branching metabolites of formononetin were completely characterized. Therefore, we selected formononetin and its primary branching metabolites for a network pharmacological analysis to explore the mechanism of anti-NAFLD.
3.6.1 Relevant targets of formononetin and primary branching metabolites against NAFLD
Fig. 5A illustrates the metabolic process of the primary branching metabolites of formononetin. The targets of 21 (including formononetin) metabolites were gathered from the Swiss Target Prediction database, and a total of 566 targets were obtained after removing duplicate targets. Additionally, using the GeneCards database, 1409 targets associated with NAFLD were acquired. Through Venny 2.1 software, the metabolites and NAFLD targets were overlapped, and 104 common target genes were determined (Fig. 5B).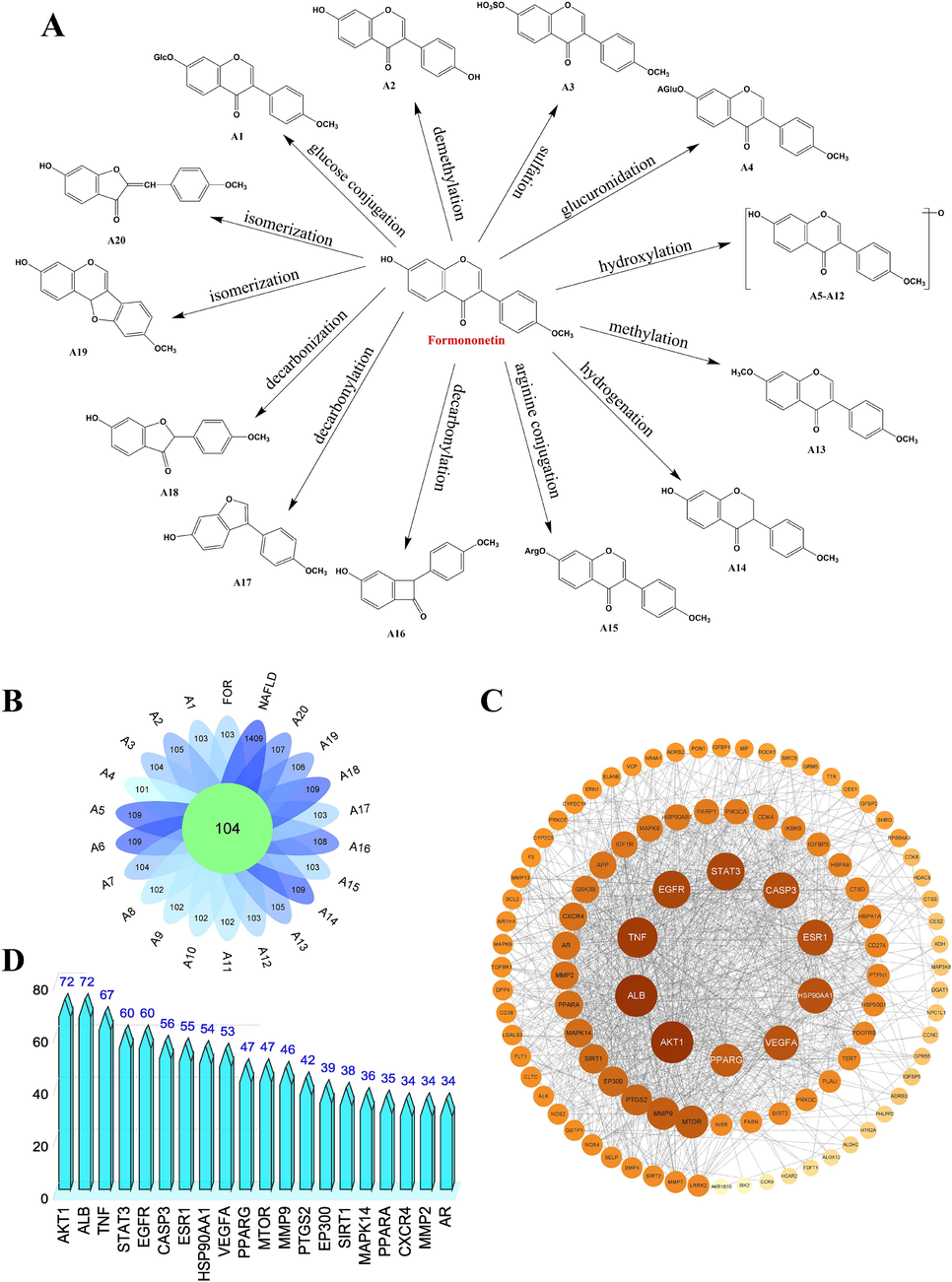
(A) Metabolic process of the primary branching metabolites of formononetin; (B) Common targets of NAFLD and 21 metabolite targets (including formononetin); (C) PPI network diagram; (D) The 20 targets with the highest degree valu.
3.6.2 Analysis of PPI network
These 104 candidate targets were entered into Cytoscape 3.9.1 software and a PPI network with 104 nodes and 1046 edges was constructed. Notably, we chose orange dots to represent nodes with higher degree values. They were more influential in the mechanism of formononetin anti-NAFLD (Fig. 5C). The top 20 targets with the highest degree values were listed in Fig. 5D, including AKT1, ALB, TNF, STAT3, EGFR, etc.
3.6.3 GO and KEGG enrichment analysis
GO functional annotation and KEGG pathway enrichment were performed on 104 targets using the Metascape database. The top 10 enriched BP (biological process), MF (molecular function), and CC (cellular compound) were graphically analyzed (Fig. 6A). BP was mainly involved in response to the hormone, positive regulation of phosphorylation, protein phosphorylation, and cellular response to the organonitrogen compound. MF was mainly involved in protein kinase activity, phosphotransferase activity, alcohol group as acceptor, kinase activity, and protein serine/threonine kinase activity. CC was mainly involved in the lytic vacuole, lysosome, vesicle lumen, and receptor complex. KEGG enrichment analysis showed that many targets in pathways in cancer, lipid and atherosclerosis, and MAPK signaling pathways were enriched. Fig. 6B provided the top 20 important signaling pathways.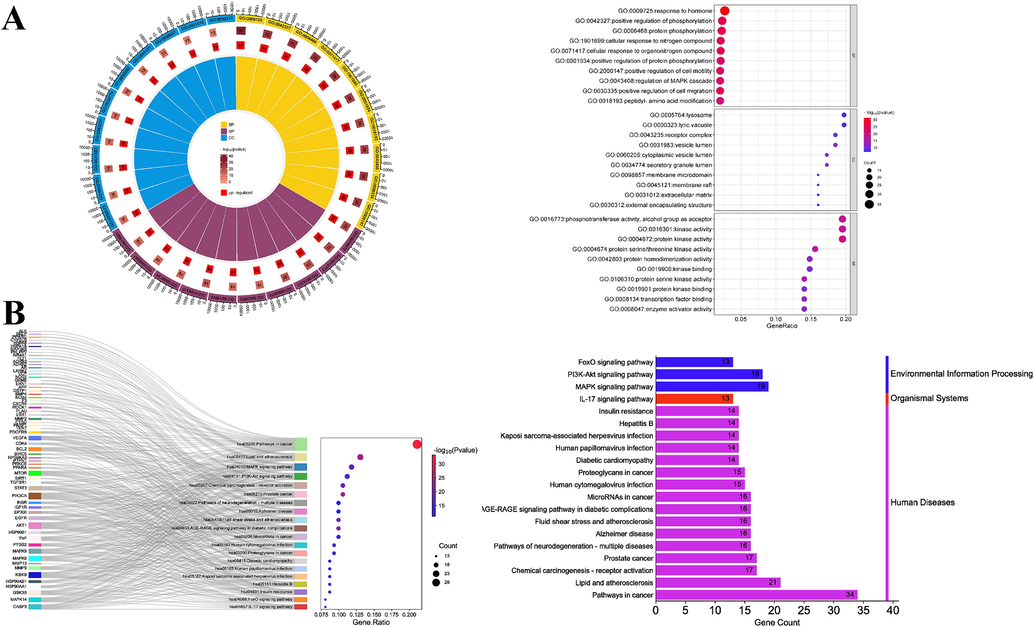
(A) Circle and dot bubble plots for GO analysis; (B) Sankey dot and category summary plots for KEGG.
3.6.4 Building a “disease-pathways-targets-metabolites” network
The “Disease-Pathway-Target-Metabolite” network diagram constructed by Cytoscape 3.9.1 software includes 20 pathways, 104 targets, 21 metabolites, and NAFLD. In the diagram, 20 primary branching metabolites were surrounded by formononetin as the center. With NAFLD as the center, 20 related pathways were encircled. Among them, NAFLD, formononetin, pathways, metabolites, and targets were represented by yellow triangles, green triangles, purple circles, pink circles, and baby blue quadrilaterals, respectively. Both metabolites and pathways were connected to the targets with lines (Fig. 7).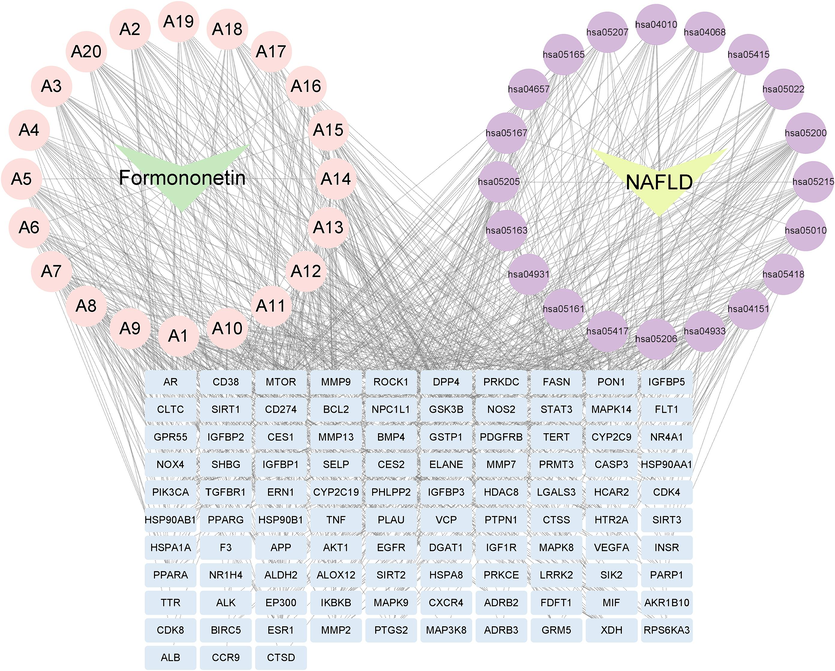
Disease-Pathways-Targets-Metabolites Network Diagram. Note: yellow triangles: NAFLD; Green triangles: formononetin; Purple circles: pathways; Pink circles: primary branching metabolites of formononetin; Baby blue quadrilaterals: targets.
4 Discussion
4.1 Feasibility analysis of recursive tree strategy establishment
Metabolite identification is a crucial step in drug metabolism research, as well as the material basis for pharmacology and pharmacodynamics research. Drug metabolites have the characteristics of complex background matrix, diverse species, and low content, which require strong detection capability of their analytical instruments. LC-HRMS has become a powerful technical tool for the analysis of drug metabolite due to its high sensitivity, high accuracy, and high separation capacity (Cui et al., 2022). By combining various data processing methods, it can be used for the characterization of metabolites, as well as the detection and structural identification of unknown trace metabolites. However, the spectral information included in the existing chemical standards and databases is very limited, and the complex and diverse metabolite structure identification cannot be satisfied by LC-HRMS technology alone. Therefore, more new techniques and research strategies are needed to meet this challenge.
In the face of this bottleneck, we have proposed the first “recursive tree” analysis strategy. Based on the addition or subtraction of a single functional group, the metabolites were classified into primary, secondary, tertiary, and multi-level branching metabolites. Taking formononetin as an example, formononetin metabolites were identified and analyzed using the recursive tree analysis strategy in vivo and in vitro. A total of 131 metabolites were identified, of which 106 were detected in vivo and 31 were detected in vitro. From the identification results, the implementation of the recursive tree analysis strategy could expand the scope of metabolite search and obtain as many metabolites as possible. In addition, the use of the recursive tree strategy and the analysis of parameters in the mass spectrometry profiles ensure the accuracy of metabolite identification.
Most importantly, the recursive tree strategy is simple, accurate, and convenient, and can be widely applied to the study of drug metabolites. The primary branching metabolites identified based on this strategy are regarded as the material basis for the treatment of diseases due to their high blood concentrations and well-defined molecular structures. In the present study, the recursive tree strategy was combined with network pharmacology, which not only comprehensively identified the metabolites of formononetin, but also improved the study of the mechanism of action of formononetin against NAFLD. The innovative initiative is of great significance as it lays the foundation for the identification of drug metabolites and their pharmacological activity studies.
4.2 Comparison of formononetin metabolites in vivo and in vitro
Biotransformation is a featured segment of drug metabolism research. Most of the metabolism studies in the crowd are mainly based on rats. Of interest is the rapidity and simplicity of microsome metabolism assays in vitro. The selective interaction between enzymes and substrates can be directly observed, reducing the interference of many internal factors (Knights et al., 2016). Therefore, microsomes can serve as a reliable source for in vivo and in vitro overall research. Among them, liver microsomes are the most effective means because they contain a variety of hepatic drug enzymes (Chen et al., 2023). At present, there have been no reports on the in vivo and in vitro metabolism of formononetin. Thus, a recursive tree analysis strategy and LC-HRMS were used to compare the metabolism of formononetin in rats and liver microsomes with a view to retrieve the metabolites of formononetin more comprehensively.
In the in vivo study, 53, 27, 46, and 9 metabolites were detected in rat plasma, urine, faeces, and liver, respectively. Although there were fewer metabolites in the liver, it might be the result of rapid metabolism by multiple enzymes. In addition, plasma, urine, and faeces have higher metabolic activity, and most of the metabolites were excreted through urine and faeces. Genistein (B9) was detected in plasma and urine as one of the important metabolites of formononetin. It has various pharmacological properties, including tyrosine and topoisomerase inhibition (Cheng et al., 2023; Jaiswal et al., 2019). In addition, the demethylated metabolite Daidzein (A2) was detected in plasma, urine, and faeces. The pharmacological effects of Daidzein, a typical isoflavone, have been extensively studied, including anti-inflammatory, hypolipidemic, antitumor, and neuroprotective effects (Pyo et al., 2009; Zhu et al., 2021; Grisley et al., 2022). It was worth mentioning that only 9 metabolites, mostly tertiary branching metabolites, were detected in the liver, even though it was the most important metabolic organ in the body. As a consequence, preliminary speculation suggested that formononetin was rapidly metabolized by hepatic drug enzymes, prone to multi-level reactions, and excreted from the body by urine and faeces via internal circulation.
In vitro study, liver microsomes also exhibited strong metabolism, and although fewer formononetin metabolites were found in liver microsomes than in rats, most phase II metabolites were shown. More importantly, Ononin (A1) was found only in liver microsomes. It indicated that the metabolic conversion of ononin to other metabolites occurred rapidly in the body. Currently, there were more studies on ononin, which was widely used in clinical practice and has clear pharmacological effects such as anti-inflammatory (Yu et al., 2023), anti-cancer (Ye et al., 2022), and anti-angiogenic (Fedoreyev et al., 2008). Overall, the metabolism of formononetin was much more complex in vivo than in vitro, but metabolism in vitro enriches the lack of biotransformation in vivo and provides a comprehensive database for the study of formononetin metabolism.
4.3 Comparison of the different biological treatment methods
The aim of sample preparation is to eliminate the interference of impurities with an appropriate recovery rate. Plasma samples are the most commonly used samples in metabolic assays and usually provide a good response to the overall level of the organism. It contains inorganic salts, oxygen, hormones, enzymes, antibodies, and cellular metabolites (Frampton et al., 2023). Due to the large number of proteins in plasma, precipitation is required prior to mass spectrometry analysis. Currently, researchers often use methanol and acetonitrile as protein precipitation reagents. The mechanism is to use organic solvents to disrupt the protein structure in biological samples, which reduces the solubility of proteins in aqueous solutions and thus converts them into solids for separation. Moreover, SPE is a sample preparation technique developed from chromatographic columns for extraction, separation, and concentration, using the difference in the interaction of different substances in the solid–liquid phases to achieve separation (Jordan et al., 2009). Hence, the plasma samples were first adsorbed onto the stationary phase and then eluted step by step using solvents with different elution capacities to achieve separation, purification, and enrichment.
The number and type of metabolites obtained by the three preparation methods used for the plasma samples were different. 49, 39, and 34 metabolites were obtained by the SPE method, methanol precipitation method, and acetonitrile precipitation method, respectively. Obviously, the SPE method yielded the highest number of metabolites and might be the best method for metabolite acquisition. Surprisingly, a large number of identical metabolites were obtained in both methanol and acetonitrile treatments. This also suggests that methanol can be chosen for protein precipitation in metabolite studies as it is cheap and easily available. The results in Table 1 showed that the samples prepared by SPE were mostly secondary and tertiary branching metabolites, specifically demethylated, sulfated, and glucuronidated metabolites. In contrast, arginine conjugation metabolites were predominantly captured in methanol and acetonitrile precipitations. (See Table 1a-1d.). Note: tR: retention time; PS: SPE column treated plasma; PM: Methanol treated plasma; PA: Acetonitrile treated plasma; U: urine; F: faeces; L: liver; LM: liver microsomes; “+”: detected; “-”: undetected. The recursive branching metabolites were marked in the lower right corner of the ID.
A15
N
8.05
C22H23N4O5
423.16629
423.16559
13.5
−1.261
269(1 0 0),304(93),205(90),178(81),423(15)
-
-
-
+
+
-
-
A16
P
6.43
C15H13O3
241.08592
241.08516
9.5
−0.750
241(1 0 0),107(49),131(30),119(2),93(2)
–
–
–
–
+
–
+
A17
P
7.89
C15H13O3
241.08592
241.08516
9.5
−3.156
109(1 0 0),241(9),131(5),147(2),95(2)
–
–
–
–
+
–
+
A18
P
5.52
C15H13O4
257.08083
257.08005
9.5
−0.785
123(1 0 0),257(62),137(12),107(24),121(3)
+
+
–
–
–
–
–
A19
N
8.29
C16H11O4
267.06519
267.06635
11.5
4.211
252(1 0 0),267(87),220(5),59(5),161(2)
+
+
+
–
–
–
–
A20
N
6.99
C16H11O4
267.06519
267.06631
11.5
4.211
252(1 0 0),267(96),220(6),201(4),132(2)
+
+
+
+
–
–
–
ID
Ion
Mode
tR/
min
Formula
[M−H]-/
[M + H]+
Theoretical
Mass (m/z)
Experimental
Mass (m/z)
RDB
Error
(ppm)
MS/MS Fragment Ions
PS
PM
PA
U
F
L
LM
B1 (A1)
N
6.54
C22H23O10
447.12857
447.13028
11.5
1.364
201(1 0 0),447(65),113(63),85(37),175(21)
–
–
–
+
–
–
–
P
6.53
C22H25O10
449.14422
449.14743
10.5
3.207
393(1 0 0),337(68),83(45),449(30),57(22)
–
–
–
–
–
+
–
B2 (A1)
N
4.31
C22H19O8
411.10744
411.10733
13.5
−2.945
255(1 0 0),375(68),212(15),411(12),184(1)
–
–
–
–
–
+
–
B3 (A1)
N
8.82
C22H23O9
431.13365
431.13458
11.5
−0.407
113(1 0 0),431(70),85(57),121(38),255(34)
–
–
–
+
–
–
–
B4 (A1)
P
9.21
C22H21O9
429.11800
429.11685
12.5
−1.159
429(1 0 0),193(58),149(31),237(31),385(21)
+
+
+
–
–
–
+
B5 (A2)
N
5.72
C15H9O5
269.04444
269.04550
11.5
−0.173
269(1 0 0),225(14),169(10),241(8),121(2)
–
–
–
–
–
–
+
P
5.70
C15H11O5
271.06009
271.05908
10.5
−2.123
271(1 0 0),215(9),197(4),136(2),153(2)
–
–
–
–
–
–
+
B6 (A2)
N
6.25
C15H9O5
269.04444
269.04565
11.5
0.384
269(1 0 0),253(32),225(2),209(8),173(1)
+
–
–
–
–
–
+
P
6.25
C15H11O5
271.06009
271.05948
10.5
−0.620
271(1 0 0),255(60),197(12),136(2),79(1)
+
–
–
–
–
–
+
B7 (A2)
N
6.43
C15H9O5
269.04444
269.04544
11.5
−0.396
269(1 0 0),213(2),225(2),197(2),173(1)
–
–
–
–
–
–
+
P
6.44
C15H11O5
271.06009
271.05902
10.5
−3.984
271(1 0 0),215(6),197(3),153(2),136(0.4)
–
–
–
–
–
–
+
B8 (A2)
N
6.61
C15H9O5
269.04444
269.04553
11.5
−0.062
269(1 0 0),225(3),240(2),197.06(1),69(1)
–
–
–
–
–
–
+
P
6.62
C15H11O5
271.06009
271.05917
10.5
−3.431
271.06(1 0 0),215(6),136(4),197(2),153(2)
–
–
–
–
–
–
+
B9 (A2)
N
8.39
C15H9O5
269.04444
269.04581
11.5
0.979
269(1 0 0),253(41),225(20),59(9),197(9)
+
–
–
–
–
–
–
P
8.40
C15H11O5
271.06009
271.05917
10.5
−3.431
271(1 0 0),167(6),153(5),215(4),253(1)
–
–
–
+
–
–
–
B10 (A2)
N
4.19
C15H9O7S
333.00634
333.00742
11.5
−0.027
214(4),253(79),333(43),81(27),305(2)
+
–
–
–
–
–
–
B11 (A2)
N
5.97
C15H9O7S
333.00634
333.00766
11.5
0.213
253(1 0 0),333(37),80(2),32(1),130(1)
+
–
–
–
–
–
–
P
5.98
C15H11O7S
335.02199
335.02109
10.5
−0.910
255(1 0 0),335(30),199(4),117(2),184(2)
+
+
–
+
–
–
–
B12
(A4)
N
4.66
C21H17O10
429.08162
429.08377
13.5
2.447
253(1 0 0),113(42),85(21),175(19),429(10)
+
+
+
–
–
–
–
P
4.65
C21H19O10
431.09727
431.09631
12.5
−0.963
255(1 0 0),98(12),421(9),178(5),348(3)
+
+
+
+
–
–
–
B13 (A4)
N
5.31
C21H17O10
429.08162
429.08231
13.5
−0.955
253(1 0 0),113(84),85(44),429(42),175(32)
+
+
+
–
–
–
–
P
5.30
C21H19O10
431.09727
431.09625
12.5
−1.023
255(1 0 0),431(12),85(3),70(2),199(2)
+
+
+
–
–
–
–
B14 (A4)
N
8.86
C22H21O10
445.11292
445.11426
12.5
0.539
121(1 0 0),445(84),225(51),181(34),357(24)
+
+
+
–
–
–
+
B15 (A4)
N
7.24
C22H21O10
445.11292
445.11523
12.5
2.718
269(1 0 0),113(71),175(35),85(33),445(17)
+
+
+
–
–
–
–
P
7.23
C22H23O10
447.12857
447.12518
11.5
−3.393
271(1 0 0),137(59),163(56),121(9),109(8)
+
–
+
–
–
–
–
B16 (A4)
N
4.79
C22H19O11
459.09218
459.09381
13.5
1.144
283(1 0 0),267(24),241(16),173(11),83(8)
+
+
+
–
–
–
–
P
4.79
C22H21O11
461.10783
461.10519
12.5
−2.648
285(1 0 0),269(94),113(66),85(32),175(28)
+
+
+
+
–
–
–
B17 (A4)
N
5.98
C22H19O11
459.09218
459.09399
13.5
1.537
268(1 0 0),459(92),283(53),113(47),85(38)
+
–
–
–
–
–
–
B18 (A4)
N
6.84
C22H19O11
459.09218
459.09409
13.5
1.754
283(1 0 0),268(25),459(10),87(5),175(5)
–
–
–
–
–
–
+
P
6.81
C22H21O11
461.10783
461.10611
12.5
−3.747
284(1 0 0),461(83),255(9),136(3),89(1)
–
–
–
–
–
–
+
B19 (A4)
P
7.03
C22H21O11
461.10783
461.10614
12.5
−3.682
284(1 0 0),461(60),126(9),337(5),136(4)
–
–
–
–
–
–
+
B20 (A4)
N
7.46
C22H19O11
459.09218
459.09390
13.5
1.341
283(1 0 0),459(41),268(37),113(3),69(3)
+
+
+
–
–
–
–
P
7.48
C22H21O11
461.10783
461.10660
12.5
−1.238
114(1 0 0),99(70),136(42),159(38),142(27)
+
–
–
–
–
–
–
B21 (A4)
N
9.54
C22H19O11
459.09218
459.09134
13.5
−1.945
121(1 0 0),401(79),313(57),225(14),459(6)
+
+
+
–
–
+
+
B22 (A5-A12)
N
6.06
C16H11O6
299.05501
299.05603
11.5
−0.272
299(1 0 0),240(25),228(9),148(3),176(2)
–
–
–
–
–
–
+
P
6.06
C16H13O6
301.07066
301.06946
10.5
−4.001
301(1 0 0),286(20),241(7),268(7),152(4)
–
–
–
–
–
–
+
B23 (A5-A12)
N
6.87
C16H11O6
299.05501
299.05603
11.5
−0.372
284.03(1 0 0),140(15),256(5),148(4),176(1)
–
–
–
–
–
–
+
P
6.88
C16H13O6
301.07066
301.06943
10.5
−4.101
301(1 0 0),286(26),241(5),153(4),136(2)
–
–
–
–
–
–
+
B24 (A5-A12)
P
7.28
C16H13O6
301.07066
301.07123
10.5
1.878
301(1 0 0),95(50),81(44),109(40),286(1)
–
–
–
–
+
–
–
B25 (A5-A12)
N
8.69
C16H11O6
299.05501
299.05624
11.5
0.430
299(1 0 0),191(11),240(8),268(8),134(5)
–
–
–
–
–
–
+
B26 (A5-A12)
N
9.01
C17H13O5
297.07575
297.07678
11.5
−0.225
297(1 0 0),268(82),224(21),134(15),196(2)
–
–
–
–
–
–
+
B27 (A5-A12)
P
9.73
C17H15O5
299.09140
299.09073
10.5
−2.240
299(1 0 0),81(44),136(36.59),95(32),121.(30)
–
–
–
+
–
–
–
B28 (A5-A12)
N
9.84
C17H13O5
297.07575
297.07669
11.5
−0.528
297(1 0 0),268(92),224(6),134(5),112(3)
–
–
–
–
–
–
+
B29 (A5-A12)
P
4.20
C18H15O6
327.08631
327.08569
11.5
−1.910
208(1 0 0),182(88),327(39),169(29),110(13)
–
–
–
+
–
–
–
B30 (A13)
N
12.44
C18H15O4
295.09648
295.09763
11.5
0.162
295(1 0 0),280(88),134(33),79(18),61(7)
–
–
–
–
–
–
+
P
12.47
C18H17O4
297.11213
297.11090
10.5
−4.158
297(1 0 0),270(4),81(3),95(3),137(2)
–
–
–
–
–
–
+
B31 (A14)
P
7.49
C16H15O3
255.10157
255.10184
9.5
1.055
255(1 0 0),142(66),119(10),133(10),237(8)
–
–
–
+
–
–
–
B32 (A14)
P
6.18
C15H15O3
243.10157
243.10086
8.5
−0.711
243(1 0 0),197(87),137(46),120(30),77(27)
+
+
+
+
–
–
–
B33 (A14)
P
7.99
C18H18NO5
328.11794
328.11768
10.5
−0.820
328(1 0 0),152(3),130(3),198(2),286(2)
–
–
–
+
–
–
–
B34 (A15)
N
12.67
C22H21N4O5
421.15064
421.15543
14.5
0.687
421(1 0 0),221(78),136(32),96(21),77(10)
–
+
+
–
–
–
–
B35 (A15)
N
6.88
C22H23N4O4
407.17138
407.17310
13.5
1.526
407(1 0 0),80(90),96(23),61(22),325(14)
–
–
–
–
+
–
–
B36 (A15)
N
7.26
C22H23N4O4
407.17138
407.17346
13.5
2.411
407(1 0 0),80(10),343(9),392(7),96(6)
–
–
–
–
+
–
–
B37 (A15)
N
8.82
C22H23N4O4
407.17138
407.17203
13.5
−1.101
407(1 0 0),80(4),392(3),345(2),78(2)
–
–
–
–
+
–
–
B38 (A15)
N
9.19
C22H23N4O4
407.17138
407.17383
13.5
3.319
407(1 0 0),389(10),80(9),377(6),96(3)
–
–
–
–
+
–
–
B39 (A15)
N
9.23
C22H23N4O4
407.17138
407.17349
13.5
2.484
407(1 0 0),80(43),377(35),307(14),125(13)
–
–
–
–
+
–
–
B40 (A15)
P
3.73
C22H23N4O4
407.17138
407.17023
13.5
−1.152
377(45),407(86),349(35),307(19),125(9)
–
+
–
–
–
–
+
B41 (A15)
N
6.58
C22H21N4O4
405.15573
405.15845
14.5
4.002
80(1 0 0),405(73),96(19),124(13),171(12)
–
–
–
–
+
–
–
P
6.54
C22H23N4O4
407.17138
407.17029
13.5
−1.092
201(1 0 0),407(72),330(40),130(29),348(13)
–
+
–
–
–
–
–
B42
N
9.45
C26H24N3O8S
538.12786
538.13043
16.5
2.734
538(1 0 0),421(1 0 0),389(10),330(40),221(16)
–
–
–
–
+
–
–
ID
Ion
Mode
tR/
min
Formula
[M−H]-/
[M + H]+
Theoretical
Mass (m/z)
Experimental
Mass (m/z)
RDB
Error
(ppm)
MS/MS Fragment Ions
PS
PM
PA
U
F
L
LM
C1 (B1)
N
6.97
C22H25O10
449.14422
449.14703
10.5
1.807
449(1 0 0),116(42),187(20),374(2),405(2)
–
–
–
+
–
–
–
P
6.96
C22H27O10
451.15987
451.15942
9.5
−1.005
158(1 0 0),334(37),207(27),120(22),451(11)
–
–
–
+
–
–
–
C2 (B5-B9)
N
7.20
C15H11O5
271.06009
271.06100
11.5
0.132
271(1 0 0),252(1 0 0),221(2),206(1),134(2),121(5)
–
–
–
–
–
–
+
C3 (B5-B9)
N
5.52
C21H19O10
431.09727
431.09869
12.5
0.320
431(1 0 0),113(77),85(48),255(47),149(43)
+
+
+
–
–
–
–
C4 (B5-B9)
N
4.97
C15H9O8S
349.00126
349.00238
11.5
0.054
269(1 0 0),349(57),116(3),190(3),80(2)
+
+
+
–
–
–
–
C5 (B5-B9)
N
6.96
C15H9O8S
349.00126
349.00269
11.5
0.942
269(1 0 0),349(57),116(3),190(3),80(2)
+
–
–
–
–
–
–
C6 (B5-B9)
N
4.79
C15H9O8S
349.00126
349.00269
11.5
0.942
269(1 0 0),349(52),219(6),80(4),62(4)
+
–
–
–
–
–
–
C7 (B5-B9)
N
4.42
C21H17O11
445.07653
445.07703
13.5
−1.358
269(1 0 0),445(8),225(5),78(5),134(3)
–
–
–
–
–
–
+
P
4.40
C21H19O11
447.09218
447.09030
12.5
−4.222
270(1 0 0),447(82),136(12),241(7),70(6)
–
–
–
–
–
–
+
C8 (B5-B9)
N
5.93
C21H17O11
445.07653
445.07849
13.5
1.922
269(1 0 0),445(28),113(13),85(7),71(5)
+
+
+
–
–
–
–
C9 (B5-B9)
N
5.63
C21H17O11
445.07653
445.07822
13.5
1.315
269(1 0 0),445(45),113(20),85(15),71(1)
+
+
+
–
–
–
–
P
5.63
C21H19O11
447.09218
447.09122
12.5
−0.968
114(1 0 0),99(70),139(42),159(38),447(1)
+
+
+
+
–
–
–
C10 (B22-B25)
P
4.87
C16H13O7
317.06557
317.06604
10.5
1.453
299(1 0 0),271(26),102(10),225(4),317(2)
–
–
–
–
–
–
+
C11 (B34)
N
6.96
C22H19O5N4
419.13499
419.13513
15.5
−0.963
419(1 0 0),221(28),243(43),373(2),263(1)
+
+
+
+
–
–
–
C12 (B35-B39)
N
6.64
C22H25N4O4
409.18703
409.18967
12.5
3.767
409(1 0 0),253(40),225(11),211(11),201(3)
–
–
–
–
+
–
–
C13 (B35-B39)
N
6.81
C22H25N4O4
409.18703
409.18945
12.5
3.229
409(1 0 0),211(42),253(16),325(6),201(4)
–
–
–
–
+
–
–
C14 (B35-B39)
N
7.59
C22H25N4O4
409.18703
409.18915
12.5
2.496
253(1 0 0),409(85),211(40),327(23),227(14)
–
–
–
–
+
–
–
C15 (B35-B39)
N
7.78
C22H25N4O4
409.18703
409.18924
12.5
2.716
409(1 0 0),253(95),96(44),327(28),201(26)
–
–
–
–
+
–
–
C16 (B35-B39)
N
7.96
C22H25N4O4
409.18703
409.18881
12.5
1.665
409(1 0 0),211(95),327(26),253(21),96(16)
–
–
–
–
+
–
–
C17 (B35-B39)
N
8.51
C22H25N4O4
409.18703
409.18915
12.5
2.496
409(1 0 0),211(73),253(48),171(18),329(12)
–
–
–
–
+
–
–
C18 (A2)
N
6.86
C21H21O9
417.11911
417.11948
11.5
0.898
417(1 0 0),255(53),241(38),163(36),59(19)
+
+
+
+
–
+
–
C19 (A3)
N
5.32
C15H13O6S
321.04273
321.04376
9.5
−0.225
321(1 0 0),121(55),241(33),80(26),135(15)
+
–
–
–
–
–
–
C20 (A3)
N
6.81
C15H13O6S
321.04273
321.04388
9.5
0.149
321(1 0 0),121(53),241(48),80(19),135(15)
+
+
+
–
–
+
–
C21 (A3)
N
6.98
C15H13O6S
321.04273
321.04379
9.5
−0.131
321(1 0 0),121(49),241(34),80(23),135(14)
+
+
+
–
–
+
–
C22 (A3)
N
7.16
C15H13O6S
321.04273
321.04391
9.5
0.243
321(1 0 0),121(48),241(34),219(27),80(20)
+
+
+
–
–
+
–
C23 (A3)
N
7.34
C15H13O6S
321.04273
321.04410
9.5
0.834
321(1 0 0),121(55),219(40),241(40),80(23)
+
+
+
–
–
+
–
C24 (A3)
N
7.53
C15H13O6S
321.04273
321.04407
9.5
0.741
321(1 0 0),121(46),241(44),219(30),80(25)
+
–
–
–
–
+
–
C25 (A5-A12)
N
5.71
C16H15O5
287.09140
287.09274
9.5
0.847
213(1 0 0),287(62),257(27),106(10),59(5)
–
–
–
–
+
–
–
C26 (A14)
P
5.93
C18H19O6
331.11761
331.11707
9.5
−1.645
211(57),121(48),133(20),86(20),331(1)
–
–
–
–
+
–
–
C27 (A14)
P
6.91
C18H19O6
331.11761
331.11768
9.5
0.197
95(1 0 0),81(93),121(71),107(67),331(2)
–
–
–
–
+
–
–
C28 (A14)
N
7.70
C18H17O6
329.10196
329.10303
10.5
−0.096
329(1 0 0),311(22),284(15),269(14),239(1)
–
–
–
–
+
–
–
C29 (A14)
P
2.79
C19H20NO4S
358.11075
358.11053
10.5
−0.629
84(1 0 0),129(94),179(58),358(48),105(48)
–
–
–
–
+
–
–
C30
N
9.98
C26H24O10N3S
570.11769
570.11871
16.5
−0.136
570(1 0 0),163(82),500(34),534(20),126(9)
+
–
+
–
–
–
–
C31
N
11.01
C26H24O10N3S
570.11769
570.11920
16.5
0.723
534(1 0 0),570(32),163(31),128(16),496(9)
+
–
–
–
–
–
–
ID
Ion
Mode
tR/
min
Formula
[M−H]-/
[M + H]+
Theoretical
Mass (m/z)
Experimental
Mass (m/z)
RDB
Error
(ppm)
MS/MS Fragment Ions
PS
PM
PA
U
F
L
LM
D1 (C3)
N
7.22
C21H21O10
433.11292
433.11414
11.5
0.120
433(1 0 0),257(94),271(64),415(40),175(30)
–
+
+
–
+
–
–
D2 (C12-C17)
N
7.27
C22H25N4O3
393.19211
393.19339
12.5
0.448
79(1 0 0),393(77),191(9),81(2),171(2)
–
–
–
–
+
–
–
D3 (C12-C17)
N
7.63
C22H25N4O3
393.19211
393.19339
12.5
0.448
79(1 0 0),393(88),191(8),96(3),171(2)
–
–
–
–
+
–
–
D4 (C12-C17)
N
7.82
C22H25N4O3
393.19211
393.19391
12.5
1.770
393(1 0 0),79(89),59(8),96(5),165(4)
–
–
–
–
+
–
–
D5 (C12-C17)
N
8.69
C22H25N4O3
393.19211
393.19296
12.5
−0.646
393(1 0 0),80(77),311(21),116(4),191(3)
–
–
–
–
+
–
–
D6 (C12-C17)
N
9.24
C22H25N4O3
393.19211
393.19354
12.5
0.829
80(1 0 0),393(68),311(40),62(16),171(3)
–
–
–
–
+
–
–
D7 (C12-C17)
N
10.00
C22H25N4O3
393.19211
393.19205
12.5
−2.960
80(1 0 0),393(82),311(32),293(20),61(11)
–
–
–
–
+
–
–
D8 (C18)
N
7.05
C20H21O7
373.12817
373.12967
10.5
0.394
329(1 0 0),373(84),284(58),147(30),175(25)
+
+
–
–
–
–
–
D9 (C18)
N
10.04
C20H21O7
373.12817
373.13278
10.5
3.504
373(1 0 0),293(98),275(66),80(28),64(13)
+
+
–
–
–
–
–
D10 (C18)
P
10.97
C20H23O7
375.14382
375.14459
9.5
2.027
375(1 0 0),189(34),213(11),199(11),357(5)
–
–
–
–
–
–
+
D11 (C18)
N
5.27
C20H19O8
387.10744
387.10858
11.5
0.101
387(1 0 0),134(65),96(42),149(26),80(26),
–
–
–
–
+
–
–
D12 (C18)
N
5.45
C20H19O8
387.10744
387.10794
11.5
−1.552
387(1 0 0),134(76),193(50),80(48),171(16)
–
–
–
–
+
–
–
D13 (C18)
N
5.83
C20H19O8
387.10744
387.10822
11.5
−0.829
387(1 0 0),343(34),80(31),134(24),284(18)
–
–
–
–
+
–
–
D14 (C18)
N
7.60
C20H19O8
387.10744
387.11099
11.5
2.449
113(1 0 0),387(71),196(56),85(54),59(30)
+
–
–
–
–
–
–
D15 (C18)
P
10.77
C20H21O8
389.12309
389.12308
10.5
−0.036
389(1 0 0),189(36),174(28),136(21),81(10)
–
–
–
–
–
–
+
D16 (C18)
N
6.96
C21H23O9
419.13365
419.13513
10.5
0.375
419(1 0 0),113(55),243(43),85(36),59(19)
+
+
+
+
–
–
–
D17 (C18)
P
6.22
C21H21O8
401.12309
401.12076
11.5
−2.334
401(80),131(18),145(14),84(13),119(13)
+
–
+
+
–
–
–
D18 (C18)
N
5.23
C21H21O12S
497.07482
497.07614
11.5
0.443
321(1 0 0),497(39),241(25),113(17),85(11)
+
+
+
+
–
–
–
D19 (C18)
N
5.41
C21H21O12S
497.07482
497.07645
11.5
1.066
321(1 0 0),497(39),241(25),113(17),85(11)
+
+
+
–
–
–
–
D20 (C18)
N
5.58
C21H21O12S
497.07482
497.07620
11.5
0.563
321(1 0 0),497(43),241(26),113(22),80(10)
+
+
+
–
–
–
–
D21 (C18)
N
5.76
C21H21O12S
497.07482
497.07697
11.5
2.112
321(1 0 0),497(40),241(26),113(20),85(8)
+
–
–
–
–
–
–
D22 (D2-D7)
N
12.30
C22H27N4O3
395.20776
395.20993
11.5
2.697
395(1 0 0),96(94),315(27),268(7),59(5)
–
–
–
+
+
–
–
D23 (D18-D21)
N
5.66
C21H21O13S
513.06973
513.07196
11.5
1.125
269(1 0 0),467(89),514(27),113(19),391(14)
+
+
+
–
–
–
–
D24 (D22)
N
7.31
C22H27N4O
363.21793
363.21829
11.5
−2.051
363(1 0 0),317(19),57(11),80(10),261(4)
–
–
–
–
+
–
–
D25 (D22)
N
8.38
C22H27N4O
363.21793
363.21814
11.5
−2.464
363(1 0 0),96(29),80(14),201(11),171(9)
–
–
–
–
+
–
–
D26 (D22)
N
8.75
C22H27N4O
363.21793
363.21909
11.5
0.152
363(1 0 0),80(31),345(21),96(15),123(6)
–
–
–
–
+
–
–
D27 (D22)
N
9.27
C22H27N4O
363.21793
363.21802
11.5
−2.794
363(1 0 0),317(17),171(4),201(3),80(1)
–
–
–
–
+
–
–
D28 (D22)
N
9.45
C22H27N4O
363.21793
363.21841
11.5
−1.720
363(1 0 0),96(7),80(5),191(4),317(2)
–
–
–
–
+
–
–
D29 (B31)
P
5.79
C16H19O
227.14304
227.14249
7.5
−2.429
213(1 0 0),150(92),210(79),107(52),227(36)
–
–
–
–
+
–
–
D30 (B31)
P
7.65
C16H19O
227.14304
227.14310
7.5
0.256
81(91),227(85),95(68),184(61),107(61)
–
–
–
–
+
–
–
D31 (B31)
P
11.16
C16H19O
227.14304
227.14328
7.5
1.049
71(1 0 0),81(29),95(15),135(11),227(6)
–
–
–
–
+
–
–
D32 (B35-B39)
N
9.78
C22H23N4O2
375.18155
375.18433
13.5
−1.224
80(1 0 0),375(94),293(46),63(21),171(5)
–
–
–
–
+
–
–
D33 (B35-B39)
N
9.95
C22H23N4O2
375.18155
375.18433
13.5
1.681
80(1 0 0),375(89),64(47),116(25),191(21)
+
–
–
–
+
–
–
D34 (B35-B39)
N
10.17
C22H23N4O2
375.18155
375.18423
13.5
1.581
375(1 0 0),80(90),64(49),191(28),293(19)
+
–
–
–
+
–
–
D35
P
7.29
C15H7O2
219.04405
219.04480
12.5
3.397
201(65),159(43),119(34),103(14),219(4)
–
–
–
–
+
–
–
D36
N
3.50
C18H16NO3S
326.08454
326.08807
11.5
2.433
108(1 0 0),326(90),150(85),173(66),85(37)
–
+
+
–
–
–
–
P
3.55
C18H18NO3S
328.10019
328.10138
10.5
3.625
152(1 0 0),110(85),85(8),120(4),328(4)
–
–
–
+
–
–
–
D37
N
2.36
C18H20NO4S
346.11075
346.10828
9.5
−3.572
346(1 0 0),316(46),241(38),127(28),104(24)
–
+
–
–
–
–
–
The differences in the metabolites obtained by the three methods were due to a variety of reasons, including the influence of the matrix, the strength of the purification ability, and the divergence in separation options. A series of factors resulted in different intensity and quantity signal peaks of formononetin metabolites after UPLC-HRMS detection.
4.4 Potential metabolic pathways of formononetin
The elucidation of the drug metabolic pathways can help the in-depth study of the pharmacological mechanisms. In the present study, the metabolites of formononetin were identified for the first time based on the recursive tree analysis strategy. There were 20, 42, 32, and 37 metabolites with primary, secondary, tertiary and multistage branches, respectively. The analysis revealed that there was a universal phase I and phase II metabolic process for formononetin, and the metabolic pathways included demethylation, decarbonylation, methylation, hydroxylation, sulfation, glucuronidation, arginine conjugation, glutathione conjugation, and their complex reactions, etc. The metabolic profile of formononetin was shown in Fig. 3. Significantly, the most metabolites were identified in plasma, followed by faeces and urine, while the fewest metabolites were identified in the liver. We speculated that after oral administration, formononetin might be biotransformed with the participation of gastric acid, digestive enzymes, and intestinal flora. After absorption into the blood, it is rapidly metabolized by the kidneys and liver and then excreted in the faeces and urine.
Understanding the metabolic properties of drugs is of great significance for drug development and clinical treatment. Formononetin is a typical isoflavone component with low polarity. Therefore, it mostly exists in organisms as hydroxylation products and conjugation products such as sulfation and glucuronidation, increasing the polarity to facilitate absorption and excretion. For example, Ononin (A1), Daidzein (A2), Formononetin-7-O-sulfate (A3), Formononetin-7-O-GluA (A4), Calycosin (A8 or A9), Biochanin A (A12), Dihydroformononetin (A14), Formononetin-7-O-Arg (A15), Genistein (B9), etc.
4.5 Formononetin attenuated PA/OA-induced lipid accumulation and oxidative stress
NAFLD is a disease caused by excessive fat deposition in the liver in addition to alcohol and other liver damage factors, and is recognized as the most common metabolic liver disease worldwide (Nian et al., 2023). NAFLD can progress from simple steatosis at the beginning to hepatitis and finally to cirrhosis and liver cancer, with a prevalence of about 25 % in adults (Jiang et al., 2023). Studies have proven that NAFLD has a long course and complex pathogenesis, which is still unclear. During the development of NAFLD, lipid metabolism disorders cause lipid accumulation. This process results in a high accumulation of ROS inducing oxidative stress in the liver. In turn, the mitochondrial dysfunction induced by oxidative stress may cause oxidative damage, which triggers a series of cascade signals leading to inflammation, cell death, and fibrosis (Loh et al., 2023). Therefore, inhibition of the inflammatory response and antioxidant may be effective in the treatment of NAFLD. At present, only some hepatoprotective and lipid-lowering drugs can be used clinically, and there is no specific drug on the market. The search for safe and effective anti-NAFLD drugs is imminent.
Our in vitro experiments preliminarily demonstrated that TG and TC levels were significantly elevated and lipid droplet aggregation was obvious in HepG2 cells induced by PA/OA. Formononetin treatment significantly reduced lipid accumulation. Oxidative stress is considered to be a key pathogenic link in the development of NAFLD (Fang et al., 2023). Dysregulation of SOD and GSH predisposes to mitochondrial dysfunction and the production of pro-inflammatory factors such as IL-6 and TNF-α (Meca et al., 2021). However, with the accumulation of lipids in the liver, the increased metabolism of free fatty acids and lipopolysaccharides leads to increased ROS production, accelerating oxidative stress and inflammatory responses in the liver (Gao et al., 2022). As stated, AST, ALT, and GSH levels were significantly elevated in PA/OA-induced HepG2 cells, whereas SOD levels were significantly decreased. However, the above indicators were reversed after treatment with formononetin. Our data suggest that formononetin ameliorated PA/OA-induced NAFLD.
4.6 Prediction of the mechanism of formononetin against NAFLD
To investigate the possible mechanisms of formononetin intervening in NAFLD, we evaluated the mechanism of action of formononetin and its primary branching metabolites against NAFLD using network pharmacology. As a result, it was indicated that AKT1, ALB, TNF, STAT3, and EGFR were more important core targets, which played pharmacological roles in NAFLD. Among them, AKT1 is involved in insulin resistance and FoxO signaling pathways, and its high level of expression contributed to the severity in patients with NAFLD (Abolfazli et al., 2022). Members of the tumor necrosis factor protein family are among the most typical triggers of hepatocyte death, and overexpression of TNF-α triggers pro-apoptotic signaling in hepatic adipocytes (Shirakami et al., 2023). Furthermore, STAT3 is a promising therapeutic target. Activation of phosphorylated STAT3 is one of the predisposing factors for the development of NAFLD to hepatocellular carcinoma (Krizanac et al., 2023). In addition, EGFR is a cell membrane protein that can be activated by its ligand, epidermal growth factor. It has been shown that epidermal growth factor receptor deletion in mice not only reduced the accumulation of macrophages in the liver, but also reduced the release of inflammatory factors (Li et al., 2023). The amelioration of NAFLD was verified by EGFR knockout mice (Shao et al., 2023). Thus, reducing the expression of EGFR may be an effective breakthrough point for the treatment of NAFLD.
To further explore the mechanism of action of formononetin and its metabolites against NAFLD and provide a scientific basis for related experimental studies, the researchers performed GO and KEGG pathway enrichment analysis. GO analysis enriched these targets in the categories such as “response to hormone”, “positive regulation of phosphorylation”, “protein kinase activity”, and “lytic vacuole”. KEGG analysis pooled targets across multiple pathways, and signaling pathways with high gene counts included: pathways in cancer, lipid and atherosclerosis, MAPK signaling pathway, PI3K-Akt signaling pathway, and prostate cancer, etc. Among these, regulation of the PI3K-AKT pathway ameliorates hepatic oxidative stress and further inhibits NF-κB mediated inflammatory responses in NAFLD. Fang et al. (2023) found that the MAPK signaling pathway has a regulatory role in oxidative stress, mitochondrial dysfunction, and hepatic steatosis. Inhibiting the activation of the MAPK signaling pathway, abnormal metabolism, and oxidative stress could be alleviated to improve NAFLD. Lipid autophagy was considered a novel mechanism to regulate cellular lipid metabolism and lipid deposition (Liu et al., 2023). Activation of the AMPK/mTOR signaling pathway enhanced hepatic macrophage autophagy, which could further inhibit IL-17 mediated inflammatory response and thus significantly improved NAFLD-related liver injury (Meng et al., 2021). In addition, TNF-α has been found to play an important role in lipid accumulation. It first promotes lipid synthesis in the liver and activates adipokines leading to obesity. At the same time, it regulates the activity of lipid metabolizing enzymes and inhibits the uptake of free fatty acids leading to lipid accumulation in the liver (Peng et al., 2023). Therefore, the TNF signaling pathway has been concluded to be a predisposing pathway in the development of NAFLD and contributes to the advancement of NAFLD to NASH.
The results of the study revealed the potential mechanism of formononetin against NAFLD from the perspective of preliminary in vitro experiments and network pharmacology, but bioinformatics has limitations. We will next validate the network pharmacology results through pharmacological experiments and molecular biology techniques.
5 Conclusion
In the present study, an analytical strategy of recursive tree bridging network pharmacology provides a powerful integrated approach to identify drug metabolites and predict the pharmacological activity of metabolites. The validity of the strategy was validated for the first time by analyzing formononetin. As a result, a total of 131 metabolites were identified. Metabolic pathways included oxidation, methylation, hydrogenation, glucuronidation, sulfation, and their complex reactions. The primary branching metabolites collectively covered the candidate targets and enrichment pathways of formononetin against NAFLD. Thus, the bioactivity of formononetin against NAFLD might be mediated by formononetin itself and/or its multiple metabolites. Additionally, the anti-NAFLD effects of formononetin was verified by in vitro cellular assays. In brief, we have not only established a new strategy for metabolite analysis, but also enriched the database of metabolic transformations of formononetin. Its combination with network pharmacology provided effective information for subsequent pharmacological studies in NAFLD.
CRediT authorship contribution statement
Yanan Li: Data curation, Writing – original draft, Writing – review & editing, Investigation, Formal analysis. Shaoping Wang: Visualization. Hong Wang: Resources. Long Dai: Methodology, Supervision. Jiayu Zhang: Conceptualization, Funding acquisition.
Acknowledgements
This work was supported by the National Natural Science Foundation of China, China (82174039), Natural Science Foundation of Shandong Province, China (ZR2020MH371), Taishan Young Scholar Program of Shandong, China (TSQN202103110), Chinese Medicine Science and Technology Program of Shandong Province, China (Z-2022085), and Matching Support Program for Provincial and Above Leading Talents in Yantai City, China (10073801).
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Bioinformatics analysis reveals molecular connections between non-alcoholic fatty liver disease (NAFLD) and COVID-19. J Cell Commun Signal.. 2022;16(4):609-619.
- [Google Scholar]
- Neuroprotective effects of Sophora secundiflora, Sophora tomentosa leaves and formononetin on scopolamine-induced dementia. Nat Prod Res.. 2021;35(24):5848-5852.
- [Google Scholar]
- A systematic strategy for rapid identification of chlorogenic acids derivatives in Duhaldea nervosa using UHPLC-Q-Exactive Orbitrap mass spectrometry. Arabian Journal of Chemistry.. 2020;13(2):3751-3761.
- [Google Scholar]
- Chen, F., Zou, H., Zhang, P., Yan, Y., 2023. Investigation on the in vitro metabolism of bicyclol using liver microsomes, hepatocytes and human recombinant cytochrome P450 enzymes. Xenobiotica. 1–31. Advance online publication.
- KCNK9 mediates the inhibitory effects of genistein on hepatic metastasis from colon cancer. Clinics (sao Paulo).. 2023;78:100141.
- [Google Scholar]
- The Analytical Strategy of “Ion Induction and Deduction Based on Net-Hubs” for the Comprehensive Characterization of Naringenin Metabolites In Vivo and In Vitro Using a UHPLC-Q-Exactive Orbitrap Mass Spectrometer. Molecules.. 2022;27(21):7282.
- [Google Scholar]
- Daidzein, its effects on impaired glucose and lipid metabolism and vascular inflammation associated with type 2 diabetes. Biofactors.. 2018;44(5):407-417.
- [Google Scholar]
- Characterization of Metabolites of α-mangostin in Bio-samples from SD Rats by UHPLC-Q-exactive Orbitrap MS. Curr Drug Metab.. 2021;22(13):1065-1073.
- [Google Scholar]
- Isoflavonoid composition of a callus culture of the relict tree Maackia amurensis Rupr. et Maxim. J Agric Food Chem.. 2008;56(16):7023-7031.
- [Google Scholar]
- The metabolic interplay between dietary carbohydrate and exercise and its role in acute appetite regulation in males: a randomized controlled study. J Physiol 2023
- [CrossRef] [Google Scholar]
- Baicalin Attenuates Oxidative Stress in a Tissue-Engineered Liver Model of NAFLD by Scavenging Reactive Oxygen Species. Nutrients.. 2022;14(3):541.
- [Google Scholar]
- Effects of Dietary Soy Protein Isolate Versus Isoflavones Alone on Poststroke Skilled Ladder Rung Walking and Cortical mRNA Expression Differ in Adult Male Rats. J Med Food.. 2022;25(2):158-165.
- [Google Scholar]
- Simultaneous determination of linarin, naringenin and formononetin in rat plasma by LC-MS/MS and its application to a pharmacokinetic study after oral administration of Bushen Guchi Pill. Biomed Chromatogr.. 2015;29(2):246-253.
- [Google Scholar]
- An Overview on Genistein and its Various Formulations. Drug Res.. 2019;69(6):305-313.
- [Google Scholar]
- Global Burden of Nonalcoholic Fatty Liver Disease, 1990 to 2019: Findings From the Global Burden of Disease Study 2019. J Clin Gastroenterol.. 2023;57(6):631-639.
- [Google Scholar]
- Comprehensive analysis of dihydromyricetin metabolites in rats using ultra-high-performance liquid chromatography coupled with high-resolution mass spectrometry. J Sep Sci.. 2022;45(21):3930-3941.
- [Google Scholar]
- Selection of SPE cartridge for automated solid-phase extraction of pesticides from water followed by liquid chromatography-tandem mass spectrometry. Anal Bioanal Chem.. 2009;394(8):2257-2266.
- [Google Scholar]
- Drug-metabolizing enzymes: role in drug resistance in cancer. Clin Transl Oncol.. 2020;22(10):1667-1680.
- [Google Scholar]
- Parent-Metabolite Pharmacokinetic Modeling of Formononetin and Its Active Metabolites in Rats after Oral Administration of Formononetin Formulations. Pharmaceutics.. 2022;15(1):45.
- [Google Scholar]
- Knights, K. M., Stresser, D. M., Miners, J. O., Crespi, C. L., 2016. In Vitro Drug Metabolism Using Liver Microsomes. Curr Protoc Pharmacol. 74, 7.8.1–7.8.24.
- Lipid-Independent Regulation of PLIN5 via IL-6 through the JAK/STAT3 Axis in Hep3B Cells. Int J Mol Scis.. 2023;24(8):7219.
- [Google Scholar]
- An Alternative Pathway for Formononetin Biosynthesis in Pueraria lobata. Front Plant Sci.. 2016;7:861.
- [Google Scholar]
- Global Profiling and Structural Characterization of Metabolites of Ononin Using HPLC-ESI-IT-TOF-MSn After Oral Administration to Rats. J Agric Food Chem.. 2020;68(51):15164-15175.
- [Google Scholar]
- Epidermal growth factor receptor promotes high-fructose nonalcoholic fatty liver disease by inducing mitochondrial fission in zebrafish. Biochem Biophys Res Commun.. 2023;652:112-120.
- [Google Scholar]
- Oleic acid improves hepatic lipotoxicity injury by alleviating autophagy dysfunction. Exp Cell Res.. 2023;429(2):113655. Advance online publication
- [Google Scholar]
- Characterization of Formononetin Sulfonation in SULT1A3 Overexpressing HKE293 Cells: Involvement of Multidrug Resistance-Associated Protein 4 in Excretion of Sulfate. Front Pharmacol.. 2021;11:614756.
- [Google Scholar]
- PKC-δ-dependent mitochondrial ROS attenuation is involved as 9-OAHSA combats lipoapotosis in rat hepatocytes induced by palmitic acid and in Syrian hamsters induced by high-fat high-cholesterol high-fructose diet. Toxicol Appl Pharmacol.. 2023;470:116557.
- [Google Scholar]
- Variations of Serum Oxidative Stress Biomarkers under First-Line Antituberculosis Treatment: A Pilot Study. J Pers Med.. 2021;11(2):112.
- [Google Scholar]
- The SGLT2 inhibitor empagliflozin negatively regulates IL-17/IL-23 axis-mediated inflammatory responses in T2DM with NAFLD via the AMPK/mTOR/autophagy pathway. Int Immunopharmacol.. 2021;94:107492.
- [Google Scholar]
- Optimization of enzymatic parameters for the production of formononetin from red clover (Trifolium pratense L.) through a response surface methodology. Nat Prod Res.. 2022;36(18):4719-4724.
- [Google Scholar]
- Gut microbiota metabolite TMAO promoted lipid deposition and fibrosis process via KRT17 in fatty liver cells in vitro. Biochem Biophys Res Commun.. 2023;669:134-142.
- [Google Scholar]
- In silico and in vivo demonstration of the regulatory mechanism of Qi-Ge decoction in treating NAFLD. Ann Med.. 2023;55(1):2200258.
- [Google Scholar]
- Comparative pharmacokinetic study on three formulations of Astragali Radix by an LC-MS/MS method for determination of formononetin in human plasma. Biomed Chromatogr.. 2019;33(9):e4563.
- [Google Scholar]
- Loss of EGFR contributes to high-fat diet-induced nonalcoholic fatty liver disease. FEBS Lett.. 2023;597(11):1503-1516.
- [Google Scholar]
- Skeletal muscle atrophy is exacerbated by steatotic and fibrotic liver-derived TNF-α in senescence-accelerated mice. J Gastroenterol Hepatol.. 2023;38(5):800-808.
- [Google Scholar]
- Comprehensive Analysis of Pterostilbene Metabolites In Vivo and In Vitro Using a UHPLC-Q-Exactive Plus Mass Spectrometer with Multiple Data-Mining Methods. ACS Omega.. 2022;7(43):38561-38575.
- [Google Scholar]
- Formononetin alleviates hepatic steatosis by facilitating TFEB-mediated lysosome biogenesis and lipophagy. J Nutr Biochem.. 2019;73:108214.
- [Google Scholar]
- Ononin Shows Anticancer Activity Against Laryngeal Cancer via the Inhibition of ERK/JNK/p38 Signaling Pathway. Front Oncol.. 2022;12:939646.
- [Google Scholar]
- Comprehensive Study of the in Vivo and in Vitro Metabolism of Dietary Isoflavone Biochanin A Based on UHPLC-Q-TOF-MS/MS. J Agric Food Chem.. 2019;67(45):12481-12495.
- [Google Scholar]
- Ononin alleviates DSS-induced colitis through inhibiting NLRP3 inflammasome via triggering mitophagy. Immun Inflamm Dis.. 2023;11(2):e776.
- [Google Scholar]
- Formononetin protects against inflammation associated with cerebral ischemia-reperfusion injury in rats by targeting the JAK2/STAT3 signaling pathway. Biomed Pharmacother.. 2022;149:112836.
- [Google Scholar]
- A Novel Formononetin Derivative Promotes Anti-ischemic Effects on Acute Ischemic Injury in Mice. Front Microbiol.. 2021;12:786464.
- [Google Scholar]
- Investigation of inhibition effect of daidzein on osteosarcoma cells based on experimental validation and systematic pharmacology analysis. PeerJ.. 2021;9:e12072.
- [Google Scholar]
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2024.105761.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary Data 1
Supplementary Data 1







