Translate this page into:
Design, synthesis, and biological evaluation of evodiamine-indolequinone hybrids as novel NQO1 agonists against non-small cell lung cancer
⁎Corresponding authors. guohui@sntcm.edu.cn (Hui Guo), yupingtang@sntcm.edu.cn (YuPing Tang)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
NQO1 is a FAD containing NAD(P)H-dependent oxidoreductase that catalyzes the reduction of quinones and related substrates, which plays an important role in the treatment of non-small cell lung cancer (NSCLC). Based on the indolequinone structure from 5-methoxy-2-methylindole, the indolequinone of NQO1 agonists was first coupled with amino-evodiamine derivatives by esterification reaction, and sixteen new compounds targeting NQO1 were developed. Among them, compounds 11b and 12d (IC50 = 2.72 or 3.66 µM, respectively) were showed better activity by cytotoxicity assay than the reference drug EVO (IC50 = 19.65 µM). Furthermore, the results of flow cytometry analysis showed that compounds 11b and 12d promoted apoptosis in A549 cells, blocked the cell cycle to the G2/M stage and caused a burst of reactive oxygen species. Western blotting experiments revealed that compounds 11b and 12d, after 24 h of treatment in A549 cells, downregulate the expression of Keap1 while upregulating the expression of Nrf2, NQO1, and HO-1. This suggests that compounds 11b and 12d increase cellular antioxidant capacity by regulating the Keap1/Nrf2/NQO1 antioxidant pathway. In vivo anti-tumor experiments showed that the reference drugs EVO (TGI = 15.94 %) and 5-Fu (TGI = 27.54 %) inhibited the proliferation of tumor tissue, while compound 11b could better inhibit the proliferation of tumor tissue (TGI = 39.13 %). In conclusion, our research results suggest that compounds 11b and 12d are potent agonism of the NQO1 signaling pathway and provide a potential opportunity to improve the treatment of NSCLC.
Keywords
Evodiamine
Indolequinone
Synthesis
Non-small cell lung cancer (NSCLC)
NQO1
1 Introduction
The transcription factor Nrf2 (nuclear factor E2-related factor 2) is the master regulator of cellular antioxidant responses, which is repressed through interaction with a redox-sensitive protein Keap1 (Kelch-like ECH-associated protein 1). The Keap1/Nrf2 pathway is an important signaling cascade responsible for cellular resistance to oxidative damage induced by exogenous chemicals and electrophiles in almost all cell types (Wu et al., 2019; Dempke and Reck, 2021). Dysregulation of Keap1/ Nrf2 transcriptional activity has been implicated in pathogenesis the Keap1/Nrf2 axis has emerged as a key modulator of cellular homeostasis (Rojo de la Vega et al., 2018). Nrf2 is highly sensitive to oxidative and electrophilic products, including reactive oxygen species (ROS) and reactive nitrogen species (RNS) (Frank et al., 2018). It is noteworthy that under normal condition, cellular Nrf2 levels are very low, but increase dramatically when exposed to varying levels of stress from electrophilic chemicals, RNS or ROS (Wu et al., 2019). The Keap1/Nrf2 system is a promising intracellular pathway to target common pathological mechanisms of many chronic diseases and cancers including NSCLC (Yu et al., 2022). NAD(P)H: Quinone Oxidoreductase 1 (NQO1) is a downstream gene of Nrf2 (Zhang et al., 2018).
NQO1 is a cyto plasmic flavoenzyme overexpressed in various tumors (lung, liver, colon, etc.) compared to normal tissue (Yang et al., 2022c; Grieco et al., 2023). This flavoenzyme can catalyze the reduction of various biologically reduced prodrugs using NADH or NADPH as cofactors (Preethi et al., 2022). NQO1 uses reduced pyridine nucleotides NADH or NADPH as cofactors to catalyze the two-electron reduction of indoquinone-based prodrugs (Fig. 1) with potential leaving groups at the indole 3-position to dihydroxyindole (2) (Rashid et al., 2021), triggering a significant increase in electron density on the indole nitrogen due to unstable drug efflux (Zhang et al., 2020). Thus, the released drug, together with the active imine electrophilic reagent 3, leads to apoptosis of cancer cells overexpressing NQO1 (Yap et al., 2022). Numerous studies have shown that NQO1-activated prodrugs exhibit significant antitumor effects (Qu et al., 2020; Zhu et al., 2022). Therefore, NQO1 agonists indoquinone prodrugs functionalized at the indole three position have emerged as an attractive strategy for antitumor prodrug design.
The mechanism of action of indoquinone NAD(P)H: NQO1 responsive prodrugs.
Lung cancer remains the leading cause of cancer death (Peters et al., 2022), which includes two main classifications based on treatment response and prognosis: small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), with the latter accounting for 85–90 % of cases (Zappa and Mousa, 2016). Clinically, a significant proportion of lung cancer patients exhibit poor a response to conventional chemo- and radio-therapeutic approach hes, resulting in a dismal 5-year survival rate of approximately 15 % (Dai et al., 2024). The recent emergence of targeted therapies and immunotherapy has offered renewed hope to NSCLC patients, but the results are still unsatisfactory. With increasing research on natural products, many natural drug monomers, such as camptothecin (Tsuchihashi et al., 2020), podophyllotoxins (Guo and Jiang, 2021), paclitaxel (Yang et al., 2020), and zebularine (Taguchi and Yamamoto, 2017), have been used for tumor treatment. Camptothecin, Podophyllotoxins, Paclitaxel, Zebularine, and Evodiamine (EVO) share structural features (Fig. 2), including benzene ring and nitrogen atoms, and all exhibit significant anticancer activity. Camptothecin exhibits excellent anti-tumor activity by inhibiting Top I, and EVO also demonstrates significant anti-tumor activity. However, the use of camptothecin is limited by side effects such as nausea, vomiting, diarrhea, and hair loss, and its anti-tumor activity decreases when formulated as a water-soluble sodium salt. Podophyllotoxin and Paclitaxel target microtubules, showing antitumor effects on testicular and breast cancers. Zebularine inhibits cancer cell growth by reversing DNA methylation. EVO induces apoptosis and inhibits proliferation, with particularly notable activity against lung cancer. The structure–activity relationships (SAR) of these compounds provide crucial insights for anticancer drug development.
Chemical structure of evodiamine, camptothecin, zebularine, podophyllotoxins, paclitaxel.
EVO (Fig. 2) is an indole alkaloid and has been shown to exert various medicinal effects, such as anti-inflammatory (Wang et al., 2021a), anti-obesity (Li and Wang, 2020), anti-tumor (Jiang et al., 2020), and anti-bacterial (Li et al., 2019). In particular, EVO has been reported to inhibit the proliferation of various tumor cells by inducing apoptosis (Liu et al., 2022; Kim et al., 2019; Yang et al., 2022a). In recent years, researchers have devoted themselves to the development of derivatives of EVO to enhance its antitumor activity and investigate its mechanism of action. Previous studies have shown that EV408, a derivative of evodiamine, has the potential to effectively target HSP70 with limited toxicity while inhibits populations of non-cancer stem-like cell (non-CSC) and cancer stem-like cell (CSC) in NSCLC (Min et al., 2023). Additionally, our team previously combined EVO with a benzosulfonamide fragment, a key component of PGAM1 inhibitors, which can cause cytotoxicity and targeting of PGAM1 enzymes against NSCLC (Wei et al., 2023). Meanwhile, EVO can induce apoptosis of lung cancer cells by regulating the PI3K/AKT (Wang et al., 2021b), MUC1-C/PD-L1 (Jiang et al., 2020), ERS (Li et al., 2022), AKT/NF-κB and SHH/GLI1 (Lin et al., 2016) signaling pathways. Although the anti-proliferative effects of EVO have been extensively studied, research on the oxidative stress pathway is relatively limited. The previous literature reported that EVO induces ROS-dependent cytotoxicity in human gastric cancer cells via TRPV1/Ca2+ pathway (Liu et al., 2022) and alleviates DEHP-induced hepatocyte pyroptosis, necroptosis and immunosuppression in grass carp through ROS-regulated TLR4/MyD88/NF-κB pathway (Lei et al., 2023). Therefore, to further expand the scope of EVO research, we introduced indolequinone into EVO to develop a compound targeting NQO1.
In this study, 3-methylindoquinone was selected for its effective NQO1-targeting activity and linked with amino-EVO derivatives through amide bonds formed using various hydrolyzable binary carboxylic acids (Fig. 3). The amino-EVO derivatives, including 2-amino-evodiamine (2-NH-EVO), 2-amino-10-methoxy-evodiamine (2-NH-10-OCH3-EVO), 3-amino-evodiamine (3-NH-EVO), and 3-amino-10-methoxy-evodiamine (3-NH-10-OCH3-EVO), were synthesized in-house (Yang et al., 2022b). Our findings suggest for the first time that the combination of EVO and indoquinone could exhibit anti-NSCLC activity through NQO1 pathway agonism.
Pharmacophore based on Evodiamine.
2 Results and discussion
2.1 Synthesis of NQO1-targeted amine evodiamine prodrugs
The indolequinone shelf was synthesized according to the route shown in Scheme 1 (Xu et al., 2017). Briefly, intermediate 2 was obtained from 5-methoxy-2-methylindole (1) as starting material by treatment with Vilsmeier's reagent made from POCl3; subsequently, the indole nitrogen was agonisted by passing sodium hydride and the methyl group was introduced by adding iodomethane to obtain intermediate 3; intermediate 4 was obtained by introducing nitro on the benzene ring using concentrated nitric acid in glacial acetic acid as solvent; intermediate 4 was dissolved in anhydrous ethanol and the addition of Fremy'salt oxidized the aniline structure to intermediate 6 with quinone structure; The aldehyde group was reduced using a reducing reagent NaBH4, and finally different diacid carboxylic acids were introduced on the alcohol hydroxyl group of intermediate 7 under DMAP conditions to obtain intermediate compounds 8a-d with different carbon chain lengths having a carboxyl group at the end. The target compounds 9a-d, 10a-d, 11a-d, and 12a-d were then produced by coupling 8a-d with these four amino-EVO derivatives via condensation reactions in the presence of EDCI and DMAP (Scheme 2).
Synthesis of indolequinone skeleton. Reagents and conditions: (i) DMF/POCl3. (ii) DMF/NaH/CH3I. (iii) AcOH/HNO3. (iv) Sn/HCl/EtOH, reflux. (v) Fremy’s salt/Acetone. (vi) NaBH4/MeOH/THF. (vii) Anhydride/DMAP/DCM.

Synthesis of compounds 9a, 9b, 9c, 9d, 10a, 10b, 10c, 10d, 11a, 11b, 11c, 11d, 12a, 12b, 12c, and 12d. Reagents and conditions: (i) EDCI/DMAP/DCM, rt, 6–8 h.
2.2 Relative efficacies of various compounds at inhibiting cancer cell proliferation
To analyze the cytotoxicity of these targeting compounds, the antiproliferative activity of compounds 9a-9d, 10a-10d, 11a-11d, and 12a-12d containing indoquinone nuclei against human lung cancer cells A549, H460, PC9, and gefitinib-resistant cells PC9/GR was initially screened by MTT assay. The compounds also acted on human normal liver cell line LO2 for 48 h. EVO was employed as the positive control. As shown in Table 1, most compounds had an antitumor efficacy on these cancer cells with IC50 ranging from 2.54 to 34.48 µM. Compared with unmodified EVO, IC50 decreased significantly. In particular, compounds 11b and 12d on the A549 cell line and PC9 cell line were more effective than other compounds. On A549 cells, the IC50 of 2.72 µM and 3.66 µM, respectively, even more than EVO with IC50 of 19.65 µM. On PC9 cells, the IC50 of 3.80 µM and 2.54 µM, respectively, even more than EVO with IC50 of 15.21 µM. Further observation revealed that compounds 11a-11d which are derivatives of 3-NH-EVO, compounds 12a-12d which are derivatives of 3-NH-10-OCH3-EVO, exhibit better activity. These two series of EVO derivatives, when conjugated with indolequinone, demonstrate superior antitumor activity compared to that of the conjugation of indolequinone with derivatives of 2-NH-EVO and 2-NH-10-OCH3-EVO. Previously, Dong et al. (2012) conducted structural modifications on EVO and found that derivatives substituted with methoxy or chlorine at the C-2 position exhibited weaker anticancer activity, indicating that the introduction of substituents at the C-2 position led to a loss of activity. Substitution at the C-3 position had the most significant impact on anticancer activity, with substitutions such as –F, –Cl, –OH, –NH, and –NHR enhancing the anticancer activity of the compounds. Subsequently, the research group reported a series of novel EVO derivatives substituted with different groups or modified scaffolds.
Compounds
IC50 (µM) a
SI b
A549
H460
PC9
PC9/GR
LO2
9a
13.88 ± 0.19
11.53 ± 0.28
3.34 ± 0.16
8.84 ± 0.23
8.41 ± 0.29
0.61
9b
16.36 ± 0.14
24.25 ± 0.25
7.28 ± 0.23
34.48 ± 0.11
29.14 ± 0.28
1.78
9c
8.54 ± 0.07
10.17 ± 0.26
2.85 ± 0.15
7.61 ± 0.15
3.18 ± 0.18
0.37
9d
9.19 ± 0.25
11.59 ± 0.19
3.86 ± 0.16
12.42 ± 0.13
6.46 ± 0.20
0.70
10a
24.11 ± 0.12
15.63 ± 0.29
4.40 ± 0.19
19.69 ± 0.27
18.56 ± 0.42
0.77
10b
5.52 ± 0.15
11.83 ± 0.27
3.76 ± 0.15
10.5 ± 0.27
8.28 ± 0.26
1.50
10c
3.39 ± 0.10
7.47 ± 0.31
3.80 ± 0.24
7.83 ± 0.19
4.73 ± 0.27
1.40
10d
27.82 ± 0.48
20.74 ± 0.13
4.42 ± 0.07
24.03 ± 0.13
18.40 ± 0.16
0.66
11a
3.16 ± 0.37
13.10 ± 0.10
8.10 ± 0.11
10.52 ± 0.11
11.02 ± 0.19
3.49
11b
2.72 ± 0.12
4.32 ± 0.16
3.66 ± 0.11
5.76 ± 0.28
8.34 ± 0.24
3.07
11c
12.07 ± 0.16
8.78 ± 0.50
7.28 ± 0.21
27.39 ± 0.13
22.00 ± 0.17
1.82
11d
8.55 ± 0.17
8.496 ± 0.12
3.45 ± 0.11
16.34 ± 0.16
15.61 ± 0.12
1.83
12a
18.62 ± 0.16
14.12 ± 0.30
5.50 ± 0.23
16.61 ± 0.17
9.71 ± 0.29
0.52
12b
4.61 ± 0.13
8.39 ± 0.25
2.64 ± 0.15
6.06 ± 0.23
6.74 ± 0.25
1.46
12c
11.78 ± 0.31
16.94 ± 0.15
4.25 ± 0.12
8.756 ± 0.39
11.27 ± 0.16
0.96
12d
3.80 ± 0.10
6.96 ± 0.24
2.54 ± 0.15
17.53 ± 0.20
6.83 ± 0.27
1.80
EVO
19.65 ± 0.27
16.19 ± 0.16
15.21 ± 0.11
27.06 ± 0.20
3.05 ± 0.43
0.16
We then calculated the selectivity index (SI) of the compounds between normal and tumor cells (Table 1). The SI, defined as the ratio of the IC50 value in normal liver cells LO2 to that in A549 cells, was 0.16 for EVO, and the SI value of compounds 11b and 12d were increased to 3.07 (19.2-fold) and 1.80 (11.3-fold), respectively. Compounds 11b and 12d exhibited weak cytotoxicity on LO2 cells, and was shown that these compounds kill cancer cells selectively over normal cells.
2.3 Bioreductive agonism of compounds 11b and 12d by NQO1
To verify that the compounds can target NQO1 to trigger anticancer activity, we selected the typical competitive inhibitor of the NQO1 enzyme, dicumarol (DIC), which inhibits its catalytic efficiency by interacting with the NAD(P)H binding site on the NQO1 enzyme (Lewis et al., 2017). In our literature research, we found that A549 cells exhibited a high expression of NQO1 protein (Xu et al., 2017), so we chose A549 cells to validate NQO1 targeting.
First, we verified the toxicity of DIC on A549 cells by exposing them to DIC for 48 h. The results are shown in Fig. 4A. The IC50 value of DIC on tumor cells after 48 h of incubation was 140.5 ± 0.15 µM, indicating that the effect of pre-incubating A549 cells with 10 µM DIC for 1 h on the proliferation of tumor cells was small. Therefore, A549 cells were co-incubated with 10 µM of DIC followed by the addition of compound 11b and compound 12d. A549 cells incubated without DIC were used as a control. The inhibition was measured at 24 h, 48 h, and 72 h (Fig. 4B). IC50 values are shown in Table 2. Without DIC, the IC50 of compound 11b at 24 h, 48 h, and 72 h was 4.17 µM, 1.86 µM, and 1.25 µM, respectively. The IC50 of compound 12d at 24 h, 48 h, and 72 h was 5.52 µM, 3.00 µM, and 1.72 µM, respectively. At the same time, the IC50 of compound 11b at 24 h, 48 h, and 72 h was 7.65 µM, 3.95 µM, and 1.84 µM after pretreatment with the NQO1 enzyme competitive inhibitor DIC for 1 h and then addition of compound 11b. The antiproliferative activity of compound 11b against A549 decreased by 1.83, 2.12, and 1.47-fold, respectively (Table 2). The IC50 of compound 12d was 8.77 µM, 4.80 µM, and 2.62 µM at 24 h, 48 h, and 72 h, respectively, and the antiproliferative activity of compound 12d against A549 decreased by 1.59, 1.60, and 1.52-fold, respectively. This indicates that the antiproliferative activity decreased after pretreating A549 cells with enzyme inhibitors followed by the action of compounds 11b and 12d, suggesting that the antitumor proliferative activity of compounds was closely related to NQO1.
(A) Antiproliferative activity of dicumarol acting on A549 cells. (B) Evaluation of NQO1-dependent cytotoxicity of compounds 11b and 12d.
Compounds
DIC (10 μM)
24 h
48 h
72 h
11b
With
7.65 ± 0.12 µM
3.95 ± 0.21 µM
1.84 ± 0.26 µM
11b
Without
4.17 ± 0.14 µM
1.86 ± 0.28 µM
1.25 ± 0.21 µM
Multiple
1.83
2.12
1.47
12d
With
8.77 ± 0.21 µM
4.80 ± 0.19 µM
2.62 ± 0.32 µM
12d
Without
5.52 ± 0.21 µM
3.00 ± 0.24 µM
1.72 ± 0.26 µM
Multiple
1.59
1.6
1.52
2.4 Effect of NQO1 on the stability of compound 11b
We used HPLC to assess the stability of compound 11b in the presence and absence of NQO1 in a pH 7.4 buffer (Fig. 5). The results showed that the decomposition of compound 11b is time-dependent. In the presence of NQO1, approximately 39 % of compound 11b decomposed within 24 h. In the absence of NQO1, about 83 % of compound 11b broke down within 24 h in the pH 7.4 buffer. In conclusion, compound 11b is a good substrate for NQO1. We detected the release process by LC-MS method (Fig. 6). After co-incubation with NQO1 enzyme and NADPH, the potential leaving group at position 3 of indole quinone nucleus underwent double electron reduction and split into two fragments ([M]+ m/z = 218.2) and ([M]+ m/z = 431.5). These findings suggest that NQO1 plays a role in promoting the release of compound 11b.
The stability of 11b in the presence and absence of NQO1 was determined by HPLC.

LC-MS analysis of 11b after 30 min co-incubation with NQO1.
2.5 Effect of compounds 11b and 12d on apoptosis
To better observe whether the inhibition of tumor cells by compounds 11b and 12d in MTT assay was related to the ability to induce apoptosis, we used Annexin V-FITC/PI kit to detect the effect of compounds on apoptosis. The assay results are shown in Fig. 7. Compound 11b and 12d were able to show concentration-dependent induction of apoptosis in A549 cells, and the total apoptosis rates were 2.18 %, 5.00 %, 14.30 %, and 27.86 % after compound 11b treated A549 cells using 0, 2, 4, and 8 µM, respectively. Compound 12d was assayed at the same concentration gradient, and compound 12d treated A549 cells using 2, 4, and 8 µM treatment of A549 cells resulted in total apoptosis rates of 5.98, 11.49 % and 34.54 %, respectively, indicating that compounds 11b and 12d could effectively induce apoptosis in A549 cells.
Compounds 11b and 12d induced apoptosis. A549 cells were incubated with varying concentrations of 11b and 12d (0, 2, 4, and 8 µM) for 48 h. At least three independent experiments were done for each condition.
2.6 Effects of compounds 11b and 12d on the cell cycle
Most antitumor drugs inhibit cell proliferation by inducing cell cycle arrest (Yang et al., 2015). To investigate the effect of compounds on the cycle of A549 cells during proliferation, we used propidium iodide (PI) staining to detect the effect of 11b and 12d on the cell cycle. A549 cells were incubated at concentrations of 0, 2, 4, and 8 µM of 11b and 12d for 48 h. The cells were stained with PI alone and the cell cycle distribution was detected using flow cytometry. The results are shown in Fig. 8. Compound 11b caused an increase in the percentage of G2/M phase cells from 12.52 % to 31.52 %. Compound 12d caused an increase in the percentage of cells in the G2/M phase from 12.52 % to 24.65 %. EVO also increased the proportion of cells in the G2/M phase to 21.00 %, which is consistent with previously reported (Hong et al., 2014). And the above results suggest that these compounds may inhibit cancer cell proliferation in the G2/M phase through cell cycle arrest. At the same time, the percentage of G1 phase cells decreased, and the percentage of S phase cells remained basically unchanged.
Compounds 11b and 12d induced G2/M arrest. A549 cells were incubated with varying concentrations of 11b and 12d (0, 2, 4, and 8 µM) for 48 h. At least three independent experiments were done for each condition.
2.7 Compounds 11b and 12d caused ROS burst in A549 cells
At lower concentrations, reactive oxygen species (ROS) function as signaling molecules, stimulating and hastening tumor proliferation and progression (Srinivas et al., 2019; Moloney and Cotter, 2018). Certain chemotherapeutic agents intensify oxidative stress and exhibit selective cytotoxicity against tumor cells (Maity et al., 2018). We used a 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA) fluorescence probe to detect ROS levels in cancer cells subjected to compounds 11b and 12d. Fig. 9 shows that A549 cells were treated at 2 µM, 4 µM, and 8 µM for 48 h, respectively, and the ROS ratio varied with the concentration. The ROS response rates of compound 11b in A549 cells at 2 µM, 4 µM, and 8 µM were 65.1 %, 87.9 % and 93.4 %, respectively. The ROS response rates of compound 12d in A549 cells at 2 µM, 4 µM, and 8 µM were 54.2 %, 62.7 % and 95.1 %, respectively. The ROS response rate in A549 cells was 88.4 % under 8 µM EVO treatment. The above results indicate that compounds 11b and 12d can both stimulate ROS generation in A549 cells, with the proportion increasing in a concentration-dependent manner.
Compounds 11b and 12d caused ROS burst. A549 cells were incubated with varying concentrations of 11b and 12d (0, 2, 4, and 8 µM) for 48 h. At least three independent experiments were done for each condition.
2.8 Western blot
To comprehend the mechanism of cell apoptosis induced by compounds 11b and 12d, western blot analysis was performed on related proteins. The experimental results (Fig. 10, S1) indicate that compounds 11b and 12d, after 24 h of treatment in A549 cells, downregulate the expression of Keap1 while upregulating the expression of Nrf2, NQO1, and HO-1. This suggests that compounds 11b and 12d increase cellular antioxidant capacity by agonizing the Keap1/Nrf2/NQO1 antioxidant pathway. Nrf2 is a transcription factor that regulates the expression of a series of antioxidant genes, including NQO1 and HO-1 (Younis, 2022). Keap1 is a negative regulator protein of Nrf2, which typically inhibits Nrf2 activity (Guo et al., 2015). Therefore, agonism of the Keap1/Nrf2/NQO1 pathway may be due to the compounds promoting oxidative stress within cells. Additionally, the upregulation of proteins P53, Bax, Caspase3, and Caspase9, along with the downregulation of Bcl2, suggests that the compounds induce apoptosis by agonizing g the apoptotic pathway in cells. Proteins Cyclin A2, CDK2, and CDK4 are all downregulated, and they participate in cell cycle regulation, particularly in the transition from the G1/S to G2/M phase (Oakes et al., 2014; Li et al., 2015). Thus, the downregulation of these proteins indicates that the compounds inhibit cell proliferation by suppressing the cell cycle regulation pathway.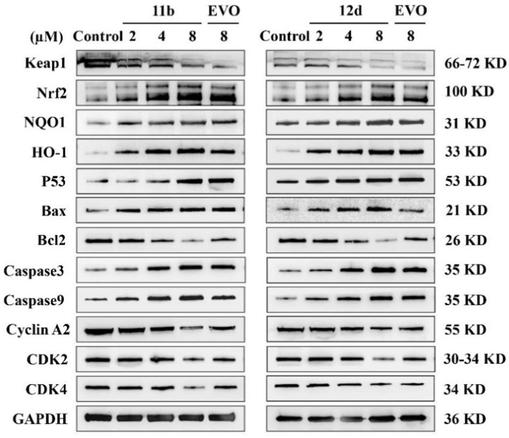
A549 cells were separately incubated with compound 11b (0, 2, 4, 8 µM), compound 12d (0, 2, 4, 8 µM) and EVO (8 µM) for 24 h, respectively. The expressions of Keap1, Nrf2, NQO1, HO-1, P53, Bax, Bcl2, Caspase3, Caspase9, Cyclin A2, CDK2 and CDK4 were determined respectively by western blotting assay. GAPDH was used as an internal loading control. At least three independent experiments were done for each condition.
ROS plays important signaling and regulatory roles within cells, excessive ROS can lead to oxidative stress and cell damage (Chio and Tuveson, 2017; Sies et al., 2017). The Western blot experiment results indicate upregulation of these antioxidant genes (Nrf2, NQO1, HO-1). It has been demonstrated that activation of NQO1 can cause DNA damage and cell death, which is associated with accumulation of reactive oxygen species (ROS) in the cells. The increase of NQO1 can further promote the production of ROS (Al-Gubory et al., 2010). This dual action may be one of the important mechanisms of drug pharmacology, but further research is needed to determine the specific cellular responses and biological effects.
2.9 Binding mode of the compounds to NQO1 protein
To gain insight into the possible interaction patterns of the synthesized compounds with indoquinone structures with NQO1 protein (PDB ID: 2F1O), molecular simulations of compounds 11b, 12d and EVO were performed. The docking results showed (Fig. 11) that the carbonyl oxygen in the amide bond of compound 11b forms two hydrogen bonds with GLY150 and TYR155, while another carbonyl group on the linker forms a hydrogen bond with PHE106. The oxygen atom near the indoloquinone moiety forms a hydrogen bond with TRP105, and one carbonyl oxygen on the indoloquinone ring forms a hydrogen bond with GLU117 (Fig. 11A), and the docking score was −11.256 kcal·mol−1. In compound 12d, the carbonyl oxygen in the ester bond on the linker forms two hydrogen bonds with GLY150 and TYR155, and the oxygen atom in the single bond on the linker forms a hydrogen bond with HIS161. Additionally, one carbonyl oxygen on the indoloquinone ring forms a hydrogen bond with TYR128 (Fig. 11B), and the docking score was −11.930 kcal·mol−1. For EVO, one hydrogen bond is formed between the side-chain NH on the indole ring and GLN104, and π-π interactions occur between the EVO skeleton and PRO68 (Fig. 11C), and the docking score was −8.070 kcal·mol−1. The Docking Score can be used to evaluate the binding of small molecule ligands to receptors, the results of the calculations are shown in Table 3. The docking study was a beneficial complement and explanation to the mode of intermolecular action. The results of this study seem very promising, as the two new compounds showed important interactions with different residues and can also serve as a basis for further studies on the design of drug candidates against cancer.
Binding models of NQO1 with compound 11b (A), 12d (B) and EVO (C), respectively. The hydrogen bonds are indicated with red dashed lines. The carbons of compounds 11b, 12d and EVO are colored in orange. The oxygen atoms are colored in red, nitrogen atoms in dark blue, sulfur atoms in yellow and hydrogen atom is grayish-white. The figure was generated using Pymol. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
Compounds
Docking Score (kcal/mol)
Compounds
Docking Score (kcal/mol)
9a
−9.648
11b
−11.256
9b
−11.348
11c
−9.593
9c
−10.442
11d
−11.113
9d
−10.686
12a
−9.315
10a
−8.314
12b
−9.689
10b
−11.273
12c
−8.104
10c
−9.238
12d
−11.930
10d
−9.271
EVO
−8.070
11a
−8.288
2.10 In vivo antitumor activity of compound 11b
In the study of the anti-proliferative activity of tumor cells, compound 11b was found to have the lowest IC50 value against A549 cells (2.72 ± 0.12 µM), and is a potential antitumor agent, based on this, the antitumor activity of compound 11b in vivo was evaluated. A preliminary efficacy experiment on the LLC-xenograft mode was prepared. Using 5-fluorouracil (5-Fu) as a positive control, EVO and compound 11b were injected intraperitoneally for 13 days (20 mg/kg, once a day), as exhibited in Fig. 12A. The changes in tumor volume and tumor weight comparison among tumor-bearing mice in each group are shown in Fig. 12B and 12D, respectively. The ex vivo tumor tissue of the mouse model is depicted in Fig. 12E. From the data in the figures, it can be observed that with the increase in time, the tumor volume of mice in each group exhibited a growing trend. Although the tumor volume of mice treated with compound 11b and 5-Fu also showed a growing trend, the rate of tumor volume growth was significantly slower compared to the control group. At the end of the treatment, the tumor volume of mice in the compound 11b and 5-Fu groups was smaller than that of the control group and the EVO group. Compared to EVO, EVO modified with an indolequinone structure demonstrated the best inhibitory effect. It was remarkable that compounds 11b and 5-Fu had no significant difference in antitumor activity under the same administration route, dose, and time, and the tumor growth inhibition rate (TGI) was 39.13 and 27.54 %, respectively (Table 4).
Compound 11b inhibited tumor growth. C57BL/6 mice were divided into four groups (n = 6) including the control, 11b, 5-Fu, and EVO. Each group except the control group was given 20 mg/kg once a day for 13 days (i.p.). (A) Experimental scheme. (B) Plot of tumor volumes over time plotted as mean ± SEM. (C) Diagram of mice weight over time, shown as mean ± SD. (D, E) Weight and size of tumors on the last day. (F) Liver index in mice. (G) Kidney index in mice. (H) IHC staining of Bax, Bcl2, Caspase3. (I) Pathological sections of liver, kidney, and tumor tissue.
Groups
Dose (mg/kg)
Tumor weight(g)
Tumor inhibition rate (%)
Control
–
0.69
–
EVO
20
0.58
15.94
5-Fu
20
0.50
27.54
11b
20
0.42
39.13
From the changes in mouse body weight (Fig. 12C), it can be seen that the body weight of each group slightly increased but without significant differences. Calculating the liver-kidney index (Fig. 12F, G) showed that compared to the control, the liver-kidney index of the EVO group was slightly larger, indicating that administration of the drug EVO could cause liver and kidney damage in mice. Compared to the control, EVO group, and positive drug 5-Fu group, compound 11b caused a decrease in the liver-kidney index, indicating that compound 11b induced less liver and kidney damage in mice, with no significant toxicity.
By slicing, staining, and observing the pathological status of ex vivo tissues, the preliminary assessment of the toxicity of synthesized compounds on normal tissues of model animals and their effects on tumor tissues was made (Fig. 12I). Observation of liver tissue slices showed that compared to the control group, the liver tissues in the 5-FU and EVO groups exhibited relaxation, while the liver tissue of compound 11b did not show significant differences from the control group. Observation of kidney tissue slices revealed that the interstitial space of renal glomeruli in the 5-FU and EVO groups tended to increase compared to the control group, while compound 11b showed little change compared to the control group. Observation of tumor slices showed that the tumor tissues in the control and EVO groups had larger nuclei, disordered cell arrangement, and darker staining, while the tumor tissues affected by compound 11b appeared loose, with a disproportionate nucleus-cytoplasm ratio and vacuolation. It can be concluded that compound 11b can inhibit tumor growth and cause minimal liver and kidney damage.
Immunohistochemistry was performed on ex vivo tumor tissues from mice in each group, and the staining results are shown in Fig. 12H. The pro-apoptotic protein Bax is a cytoplasmic protein. When cells are exposed to apoptotic stimuli, Bax enters the mitochondria through a specific translocation pathway, promoting the release of cytochrome C, further agonizing Caspase3, and initiating cell apoptosis (Légiot et al., 2019; Jürgensmeier et al., 1998). Therefore, Caspase3 plays a crucial role in cell apoptosis (Wang et al., 2017). In Fig. 12H, it can be observed that the compound 11b group exhibits significantly brown staining for Bax and Caspase3 compared to the blank and EVO groups, and is stronger than the 5-Fu group. Regarding the expression of the anti-apoptotic protein Bcl2, compound 11b shows weaker brown staining compared to the blank and EVO groups. Therefore, it can be concluded that compound 11b inhibits tumor cell proliferation by promoting the expression of Bax and Caspase3 while suppressing the expression of Bcl2.
3 Materials and methods
3.1 Chemistry
Most chemicals and solvents are purchased commercially and further purified and dried. The solvent was dried using a 4 Å molecular sieve. 1H and 13C NMR raw data were collected by a Bruker AVANCE400 and AVANCE500 spectrometer and spectrums were exported through MestReNova software. Deuterated solvents included DMSO‑d6, CDCl3, and D2O, and tetramethylsilane was used as an internal standard. The chemical shifts (δ) are reported in parts per million (ppm) and the coupling constant (J) unit is Hz. ESI mass spectra were performed on an Agilent 7250&JEOL-JMS-T100LP AccuTOF. TLC analysis was carried out on silica gel plates GF254 (Qingdao Haiyang Chemical, China). Silica gel column chromatography was performed with Silica gel 60 G (Qingdao Haiyang Chemical, China). The synthetic route of the indoloquinone structure started from 5-methyl-2-methoxyindole and was obtained by a series of substitution and oxygen reduction reactions (Zhang et al., 2005). The purity of all tested compounds was ≥95 % by HPLC analysis (S2).
3.2 General experimental procedure for the synthesis of target compounds
3.2.1 5-Methoxy-2-methyl-1H-indole-3-carbaldehyde (2)
Vilsmeier reagent was prepared by adding POCl3 (0.85 mL, 9.28 mmol) to 3 mL of anhydrous DMF and stirring for 10 min at 0 °C. Then dissolve 5-methoxy-2-methylindole (1.04 g, 6.45 mmol) in 3 mL anhydrous DMF, cool to 0 °C, add the prepared Vilsmeier reagent dropwise, and after the dropwise addition, stir the reaction for 30 min at 0 °C. Subsequently, drop the reaction mixture into an ice-based 2 M NaOH (50 mL) solution, add DCM (100 mL), extract and separate. The aqueous layer was then extracted with DCM (50 mL), the organic layer was combined, rinsed with brine, dried with anhydrous Na2SO4, the solvent was removed, and the residue was washed using ice-ethyl acetate to give 1.05 g (90 %) of light brown compound 2. 1H NMR (400 MHz, DMSO‑d6) δ 10.04 (s, 1H), 7.74 (d, J = 2.2 Hz, 1H), 7.23 (d, J = 8.8 Hz, 1H), 6.88 (dd, J = 8.8, 2.5 Hz, 1H), 3.88 (s, 3H), 2.72 (s, 3H). 13C NMR (101 MHz, DMSO) δ 184.47, 162.71, 156.40, 147.94, 130.22, 126.91, 113.07, 111.82, 102.88, 55.87, 36.61.
3.2.2 5-Methoxy-1,2-dimethyl-1H-indole-3-carbaldehyde (3)
Compound 2 (1.4 g, 7.40 mmol) and NaH (0.443 g, 60 % mineral oil, 11.10 mmol) were dissolved in anhydrous DMF under N2 protection, and the mixture solution was stirred at room temperature for 2 h. Subsequently, iodomethane (0.91 mL, 14.6 mmol) was added dropwise under an ice bath at 0 °C, and the reaction was warmed to room temperature and stirred for another 2 h. The mixture was dissolved in DCM, extracted sequentially using water and NaCl solution, dried with anhydrous Na2SO4, and after removing the solvent, purified using a silica gel column, eluting with ethyl acetate/petroleum ether (1:2, Rf = 0.3) to give 3.68 g (97.0 %) of a white solid 3. 1H NMR (400 MHz, DMSO‑d6) δ 10.09 (s, 1H), 7.79 (d, J = 2.4 Hz, 1H), 7.19 (d, J = 8.8 Hz, 1H), 6.91 (dd, J = 8.8, 2.5 Hz, 1H), 3.89 (s, 3H), 3.68 (s, 3H), 2.66 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 183.86, 162.56, 156.60, 147.92, 131.88, 126.31, 112.84, 109.97, 102.97, 55.88, 29.73, 10.42.
3.2.3 4-AMINO-5-methoxy-1,2-dimethyl-1H-indole-3-carbaldehyde (5)
Compound 3 (1.48 g, 5.38 mmol) was dissolved in acetic acid (88.7 mL) and cooled to 10–15 °C. Nitric acid solution (3.2 mL nitric acid dissolved in 18.8 mL acetic acid) was added dropwise using a dropping funnel and the temperature was maintained at 10–15 °C. Subsequently, the mixture was transferred to room temperature and stirred for another 2 h. The reaction mixture was then poured into ice (200 g) and extracted with DCM (150 mL × 3), the combined organic layers were extracted sequentially with water (200 mL) and brine (200 mL), dried over anhydrous Na2SO4 and the solvent was removed by vacuum concentration to give a mixture of 4-nitro and 6-nitro. The mixture (1.67 g) was suspended in ethanol (100 mL), tin pellets (2.23 g, 18.8 mmol) were added, followed by HCl (3.0 M, 23 mL), and the reaction was heated at reflux for 1 h, water (20 mL) was added, pH was adjusted to 6 using saturated Na2CO3 (aq), DCM (200 mL × 3) was extracted, and the organic layer was combined, sequentially, with water (100 mL), brine (50 mL) extraction, anhydrous Na2SO4 drying, solvent removal, silica gel column purification, elution with ethyl acetate/petroleum ether (1.5:1, Rf = 0.15) to give 0.43 g (47 %, two steps) of 5 as a yellow solid. 1H NMR (400 MHz, DMSO‑d6) δ 9.76 (s, 1H), 6.85 (d, J = 8.5 Hz, 1H), 6.45 (d, J = 8.5 Hz, 1H), 5.91 (s, 2H), 3.86 (s, 3H), 3.56 (s, 3H), 2.57 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 183.08, 149.82, 141.76, 134.63, 132.34, 115.58, 113.24, 110.13, 96.17, 29.91, 10.66.
3.2.4 5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indole-3-carbaldehyde (6)
Compound 5 (0.20 g, 0.68 mmol) was dissolved in acetone, then NaH2PO4/Na2HPO4 buffer (18.9 mL, 0.3 M, pH 6.0) was added with fremy's salt (0.59 g, 2.2 mmol) under stirring. The reaction was monitored at room temperature for 3 h, TCL, extracted with DCM (100 mL × 3), the organic layer was combined, extracted sequentially with water and NaCl solution, and dried over Na2SO4. The solvent was removed and 6 (0.134 g, 97 %) was obtained as a red solid without purification. 1H NMR (400 MHz, DMSO‑d6) δ 10.55 (s, 1H), 5.71 (s, 1H), 3.94 (s, 3H), 3.85 (s, 3H), 2.62 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 188.34, 179.06, 177.87, 159.84, 142.71, 120.00, 106.75, 56.77, 32.26, 11.33.
3.2.5 3-(Hydroxymethyl)-5-methoxy-1,2-dimethyl-1H-indole-4,7-dione (7)
Compound 6 (0.113 g, 0.48 mmol) was dissolved in an equal mixture of anhydrous methanol and anhydrous tetrahydrofuran, and NaBH4 (0.91, 2.40 mmol) was added under an ice bath, stirred for 8 min at 0 °C, and the reaction progress was monitored by TCL, followed by quenching with saturated NH4Cl solution, DCM extraction, and then washing with water and brine sequentially. Dried with anhydrous Na2SO4, concentrated, purified by silica gel column, and eluted with EA/PE (1:1, Rf = 0.21) to give 0.41 g (67.7 %) of 7 as an orange-red solid. 1H NMR (400 MHz, DMSO‑d6) δ 5.64 (s, 1H), 4.62 (d, J = 7.1 Hz, 2H), 3.89 (s, 3H), 3.83 (s, 3H), 2.23 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 179.69, 179.05, 160.11, 135.05, 129.87, 123.20, 122.49, 107.53, 57.00, 56.34, 32.82, 9.97.
3.2.6 Compounds 8
To improve the rate and yield of compound 8, adipic acid and 3,3′-dithiodipropionic acid were first prepared as anhydride using oxalyl chloride, respectively (He et al., 2017). Adipic acid (4 g, 27.2 mmol) and 3,3′-dithiodipropionic acid (4 g, 19 mmol) were dissolved in 12 mL of oxalyl chloride, respectively, and refluxed at 67 °C for 1.5 h. The solvent was removed by concentration under reduced pressure, and the residue was precipitated in cold ether (80 mL) for at least 4 h. The mixture was filtered, the ether was removed and dried at room temperature.
The anhydrous DCM was used to dissolve the anhydride (85 mg, 0.85 mmol) and DMAP (20.7 mg, 0.17 mmol), and the mixture was stirred at room temperature for 30 min, followed by the addition of compound 7, and the reaction was continued for 12 h. The solvent was removed, and the residue was dissolved in DCM, extracted with water and NaCl solution in turn, dried over anhydrous Na2SO4, concentrated, and purified on a silica gel column. Purification, PE/EA (1:1, Rf = 0.3) for elution, and finally obtained orange-yellow solid compound 8 (68 %-82 %).
3.2.6.1 4-((5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methoxy)-4-oxobutanoic acid (8a)
1H NMR (400 MHz, DMSO‑d6) δ 5.62 (s, 1H), 5.28 (s, 2H), 3.90 (s, 3H), 3.80 (s, 3H), 2.69 ∼ 2.65 (m, 2H), 2.63 ∼ 2.59 (m, 2H), 2.27 (s, 3H); 13C NMR (101 MHz, DMSO‑d6) δ 179.03, 177.74,177.36, 172.20, 159.83, 138.11, 129.25, 124.96, 115.75,106.79, 57.06, 56.59, 32.55, 29.01, 28.80, 9.65 ppm.
3.2.6.2 5-((5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methoxy)-5-oxopentanoic acid (8b
1H NMR (400 MHz, DMSO‑d6) δ 5.62 (s, 1H), 5.25 (s, 2H), 3.90 (s, 3H), 3.80 (s, 3H), 2.42 ∼ 2.37 (m, 4H), 2.28 (s, 3H), 1.97 ∼ 1.92 (m, 2H); 13C NMR (101 MHz, DMSO‑d6) δ 179.04, 178.31, 177.74, 172.98, 159.85, 137.99, 129.32, 121.89, 115.88, 106.79, 56.82, 56.59, 33.24, 33.04, 32.56, 20.02, 9.68 ppm.
3.2.6.3 6-((5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methoxy)-6-oxohexanoic acid (8c)
1H NMR (400 MHz, DMSO‑d6) δ 5.73 (s, 1H), 4.57 (s, 2H), 3.83 (s, 3H), 3.75 (s, 3H), 2.74 ∼ 2.66 (m, 2H), 2.24 (s, 3H), 2.04 ∼ 1.93 (m, 1H), 1.23 (s, 3H), 1.10 (t, J = 7.5 Hz, 2H); 13C NMR (101 MHz, DMSO‑d6) δ 177.99, 177.37, 159.19, 137.08, 127.52, 121.67, 120.42, 106.56, 56.37, 53.20, 37.96, 31.87, 30.89, 16.33, 13.61, 9.15 ppm.
3.2.6.4 3-((3-((5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methoxy)-3-oxopropyl)disulfanyl)propanoic acid (8d)
1H NMR (400 MHz, DMSO‑d6) δ 5.73 (s, 1H), 4.56 (s, 2H), 3.83 (s, 3H), 3.75 (s, 3H), 2.75 ∼ 2.65 (m, 2H), 2.24 (s, 3H), 1.23 (s, 2H), 1.10 (t, J = 7.5 Hz, 2H); 13C NMR (101 MHz, DMSO‑d6) δ 177.98, 177.38, 159.20, 142.28, 137.09, 127.53, 121.66, 120.42, 106.55, 56.37, 53.21, 31.88, 16.34, 13.61, 9.15 ppm.
3.2.7 Compounds 9
2-Aminowoodine (100 mg, 0.314 mmol), was dissolved in 5 mL anhydrous DCM, followed by the addition of compound 8 (109 mg, 0.314 mmol) and EDCI (300.98 mg, 1.57 mmol), the mixture was cooled to 0 °C and then DMAP (38.36 mg, 0.314 mmol) was added at room temperature The reaction was monitored by TCL with stirring for 24–48 h. After the reaction, the solvent was removed and the mixture was dissolved in DCM, washed with water, saturated saline, dried with anhydrous Na2SO4, concentrated under reduced pressure, purified by silica gel column separation, and eluted with CH3OH/DCM (2 ∼ 4 %, Rf = 0.4), respectively. A yellow solid compound 9 was obtained (48–62 % yield).
3.2.7.1 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 4-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-4-oxobutanoate (9a)
1H NMR (400 MHz, DMSO‑d6) δ 10.96 (s, 1H), 10.17 (d, J = 16.5 Hz, 1H), 7.69–7.58 (m, 1H), 7.50–7.33 (m, 3H), 7.12–6.91 (m, 3H), 6.13 (s, 1H), 5.73 (s, 1H), 5.15 (s, 2H), 3.76 (d, J = 18.3 Hz, 6H), 3.56 (d, J = 39.3 Hz, 2H), 3.18 (s, 2H), 2.96 (s, 3H), 2.59 (s, 4H), 2.21 (s, 3H);13C NMR (101 MHz, DMSO‑d6) δ 178.61, 177.54, 172.56, 170.83, 164.67, 159.68, 149.54, 144.34, 139.03, 136.82, 131.79, 129.23, 128.63, 126.54, 122.23, 121.30, 119.33, 118.57, 115.21, 113.73, 112.09, 111.79, 110.87, 107.13, 105.84, 70.85, 56.95, 56.72, 51.85, 36.99, 32.53, 31.63, 29.07, 19.86, 9.54. MS (ESI, positive) found (M+H) 636.24, calc (C35H33N5O7, m/z): 635.23. HPLC (MeOH:H2O = 70:30): 97.771 %.
3.2.7.2 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 5-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-5-oxopentanoate (9b)
1H NMR (400 MHz, DMSO‑d6) δ 10.98 (s, 1H), 10.10 (d, J = 9.0 Hz, 1H), 7.67 (d, J = 8.5 Hz, 1H), 7.52 (s, 1H), 7.44 (d, J = 7.8 Hz, 1H), 7.35 (d, J = 8.1 Hz, 1H), 7.10 (t, J = 7.5 Hz, 1H), 6.98 (q, J = 9.7, 8.7 Hz, 2H), 6.13 (s, 1H), 5.15 (s, 1H), 3.82 (s, 1H), 3.73 (s, 1H), 3.60 (s, 3H), 3.35 (s, 3H), 2.96 (s, 5H), 2.50 (s, 2H), 2.42–2.28 (m, 5H), 2.23 (s, 1H), 1.85 (p, J = 7.4 Hz, 3H);13C NMR (101 MHz, DMSO‑d6) δ 178.65, 177.54, 173.48, 172.91, 171.65, 164.66, 159.70, 149.61, 144.39, 138.96, 136.84, 131.73, 129.24, 128.71, 126.54, 122.23, 121.31, 119.33, 118.58, 115.26, 113.87, 112.09, 111.80, 111.08, 107.16, 106.10, 70.82, 56.95, 56.67, 51.73, 33.23, 33.08, 20.78, 20.73, 19.87, 9.54. MS (ESI, positive) found (M+H) 650.26, calc (C36H35N5O7, m/z): 649.69. HPLC (MeOH:H2O = 70:30): 95.211 %.
3.2.7.3 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 6-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-6-oxohexanoate (9c)
1H NMR (400 MHz, DMSO‑d6) δ 10.96 (s, 1H), 10.07 (d, J = 8.9 Hz, 1H), 7.66 (d, J = 8.4 Hz, 1H), 7.52 (s, 1H), 7.44 (d, J = 7.7 Hz, 1H), 7.35 (d, J = 8.0 Hz, 1H), 7.09 (t, J = 7.4 Hz, 1H), 7.03–6.94 (m, 2H), 6.13 (s, 1H), 5.75 (d, J = 4.2 Hz, 1H), 5.13 (s, 2H), 3.82 (s, 3H), 3.73 (s, 3H), 3.24–3.13 (m, 2H), 2.96 (s, 3H), 2.78–2.64 (m, 2H), 2.30 (d, J = 10.6 Hz, 4H), 2.22 (s, 3H), 1.56 (s, 4H).; 13C NMR (101 MHz, DMSO‑d6) δ 178.65, 177.52, 173.12, 172.02, 164.67, 159.69, 149.58, 144.44, 138.96, 136.83, 131.76, 129.23, 128.70, 126.54, 122.23, 121.31, 119.33, 118.58, 115.31, 113.79, 112.08, 111.79, 111.02, 107.16, 106.01, 70.83, 56.96, 56.57, 41.65, 37.00, 33.62, 32.57, 24.86, 24.55, 19.87, 9.53. MS (ESI, positive) found (M+H) 664.27, calc (C37H37N5O7, m/z): 663.72. HPLC (MeOH:H2O = 70:30): 99.725 %.
3.2.7.4 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 3-((3-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-3-oxopropyl)disulfaneyl)propanoate (9d)
1H NMR (400 MHz, DMSO‑d6) δ 10.96 (s, 1H), 10.21 (s, 1H), 7.66 (d, J = 8.5 Hz, 1H), 7.50 (s, 1H), 7.44 (d, J = 7.7 Hz, 1H), 7.34 (d, J = 8.1 Hz, 1H), 7.09 (t, J = 7.5 Hz, 1H), 7.01–6.94 (m, 2H), 6.13 (s, 1H), 5.75 (d, J = 1.8 Hz, 2H), 5.17 (s, 2H), 3.82 (s, 3H), 3.74 (s, 3H), 3.19 (td, J = 12.4, 4.6 Hz, 2H), 2.97–2.90 (m, 9H), 2.73–2.66 (m, 4H), 2.22 (s, 3H); 13C NMR (101 MHz, DMSO‑d6) δ 178.66, 177.56, 171.53, 170.11, 164.63, 159.70, 149.55, 144.20, 139.08, 136.82, 131.76, 129.29, 128.70, 126.54, 122.24, 121.32, 119.34, 118.58, 115.06, 113.91, 112.09, 111.80, 111.01, 107.17, 70.83, 56.98, 36.99, 36.61, 33.89, 33.24, 32.59, 19.86, 9.58. MS (ESI, positive) found (M+H) 728.22, calc (C37H37N5O7S2, m/z): 727.85. HPLC (MeOH:H2O = 70:30): 97.812 %.
3.2.7.5 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 4-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-4-oxobutanoate (10a)
1H NMR (400 MHz, DMSO‑d6) δ 11.00 (s, 1H), 9.96 (s, 1H), 8.01 (s, 1H), 7.69 (d, J = 11.1 Hz, 1H), 7.25 (d, J = 8.8 Hz, 1H), 7.05 (d, J = 8.8 Hz, 1H), 7.01–6.97 (m, 1H), 6.75 (dd, J = 8.7, 2.3 Hz, 1H), 6.01 (s, 1H), 5.75 (s, 1H), 5.14 (s, 2H), 3.78 (s, 3H), 3.76 (s, 3H), 3.73 (s, 3H), 3.22–3.13 (m, 2H), 2.87–2.78 (m, 3H), 2.63 (s, 2H), 2.55 (s, 4H), 2.20 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.60, 177.52, 172.56, 169.93, 164.27, 159.65, 153.82, 145.52, 139.00, 134.02, 132.18, 130.89, 128.60, 126.59, 124.88, 121.49, 121.27, 120.35, 118.66, 115.24, 112.75, 111.84, 107.10, 100.65, 70.26, 69.73, 56.92, 56.61, 55.84, 55.35, 36.95, 32.52, 31.34, 29.27, 20.21, 9.53. MS (ESI, positive) found (M+H) 666.25, calc (C36H35N5O8, m/z): 665.69. HPLC (MeOH:H2O = 70:30): 99.404 %.
3.2.7.6 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 5-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-5-oxopentanoate (10b)
1H NMR (400 MHz, DMSO‑d6) δ 10.99 (s, 1H), 9.91 (d, J = 11.2 Hz, 1H), 8.03 (dd, J = 6.0, 2.4 Hz, 1H), 7.78–7.72 (m, 1H), 7.24 (d, J = 8.8 Hz, 1H), 7.08 (d, J = 8.7 Hz, 1H), 6.99 (d, J = 2.1 Hz, 1H), 6.76 (d, J = 8.8 Hz, 1H), 6.01 (s, 1H), 5.75 (s, 2H), 5.15 (s, 1H), 3.85–3.71 (m, 6H), 3.59 (s, 3H), 3.21–3.12 (m, 2H), 2.86––2.79 (m, 2H), 2.63 (s, 3H), 2.39–2.28 (m, 5H), 2.23 (s, 1H), 1.83 (s, 2H), 1.23 (s, 1H). 13C NMR (101 MHz, DMSO‑d6) δ 178.67, 173.50, 172.93, 170.80, 164.30, 159.71, 153.82, 145.57, 139.01, 133.99, 132.16, 130.92, 126.59, 125.15, 121.44, 120.33, 118.87, 112.75, 112.45, 111.83, 107.16, 100.65, 69.76, 56.96, 56.65, 55.84, 55.36, 51.72, 36.97, 35.63, 33.11, 32.59, 20.90, 20.19, 9.56. MS (ESI, positive) found (M+H) 680.27, calc (C37H37N5O8, m/z): 679.72. HPLC (MeOH:H2O = 70:30): 99.590 %.
3.2.7.7 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 6-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-6-oxohexanoate (10c)
1H NMR (400 MHz, DMSO‑d6) δ 10.96 (s, 1H), 10.06 (s, 1H), 7.65 (d, J = 8.5 Hz, 1H), 7.52 (s, 1H), 7.34 (d, J = 8.1 Hz, 1H), 7.09 (t, J = 7.5 Hz, 1H), 6.97 (dd, J = 11.5, 7.8 Hz, 2H), 6.13 (s, 1H), 5.75 (s, 1H), 5.14 (s, 2H), 3.83 (s, 3H), 3.73 (s, 3H), 3.59 (s, 1H), 3.19 (d q, J = 12.2, 4.2, 3.7 Hz, 1H), 2.93 (d, J = 20.2 Hz, 5H), 2.76–2.69 (m, 1H), 2.35–2.26 (m, 4H), 2.23 (s, 3H), 1.61–1.51 (m, 4H). 13C NMR (101 MHz, DMSO‑d6) δ 178.67, 173.13, 172.04, 164.67, 159.71, 149.58, 139.00, 136.82, 131.78, 129.23, 126.54, 119.33, 118.59, 113.78, 112.08, 111.79, 111.02, 107.18, 70.83, 56.97, 56.58, 37.01, 36.60, 33.62, 32.60, 24.85, 24.54, 9.55. MS (ESI, positive) found (M+H) 694.29, calc (C38H39N5O8, m/z): 693.74. HPLC (MeOH:H2O = 70:30): 99.850 %.
3.2.7.8 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 3-((3-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-2-yl)amino)-3-oxopropyl)disulfaneyl)propanoate (10d)
1H NMR (400 MHz, DMSO‑d6) δ 10.99 (s, 1H), 10.03 (s, 1H), 8.03 (d, J = 2.2 Hz, 1H), 7.75 (dd, J = 8.7, 2.3 Hz, 1H), 7.24 (d, J = 8.7 Hz, 1H), 7.07 (d, J = 8.7 Hz, 1H), 6.98 (s, 1H), 6.75 (d, J = 10.9 Hz, 1H), 6.01 (s, 1H), 5.74 (d, J = 5.1 Hz, 1H), 5.16 (s, 2H), 3.82 (s, 3H), 3.74 (d, J = 7.0 Hz, 6H), 3.21–3.12 (m, 2H), 3.00–2.87 (m, 6H), 2.81 (s, 3H), 2.50 (s, 4H), 2.22 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.64, 177.54, 171.53, 169.23, 164.27, 159.70, 153.81, 145.64, 139.08, 133.75, 132.16, 130.93, 128.67, 126.59, 125.13, 121.30, 120.23, 118.89, 115.06, 112.75, 112.45, 111.83, 107.15, 100.63, 69.78, 56.97, 55.84, 36.95, 36.34, 34.05, 33.89, 33.27, 32.58, 20.19, 9.59. MS (ESI, positive) found (M+H) 758.23, calc (C38H39N5O8S2, m/z): 757.87. HPLC (MeOH:H2O = 70:30): 96.225 %.
3.2.7.9 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 4-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-4-oxobutanoate (11a)
1H NMR (400 MHz, DMSO‑d6) δ 11.20 (s, 1H), 9.97 (s, 1H), 8.01 (s, 1H), 7.69 (s, 1H), 7.49 (s, 1H), 7.36 (s, 1H), 7.12 (s, 1H), 7.01 (d, J = 7.4 Hz, 2H), 6.04 (s, 1H), 5.72 (s, 1H), 5.16 (s, 2H), 3.80 (s, 3H), 3.74 (s, 3H), 3.50 (s, 2H), 3.17 (s, 3H), 2.86 (s, 2H), 2.62 (s, 2H), 2.55 (s, 2H), 2.21 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.61, 177.55, 172.56, 170.83, 164.67, 159.68, 149.55, 144.34, 139.03, 136.83, 131.79, 129.23, 128.63, 126.54, 122.23, 121.30, 119.33, 118.57, 115.21, 113.73, 112.09, 111.79, 110.87, 107.13, 105.84, 70.85, 56.95, 56.72, 51.85, 36.99, 32.53, 31.63, 29.07, 28.84, 19.86, 9.53. MS (ESI, positive) found (M+H) 636.24, calc (C35H33N5O7, m/z): 635.67. HPLC (MeOH:H2O = 70:30): 98.224 %.
3.2.7.10 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 5-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-5-oxopentanoate (11b)
1H NMR (400 MHz, DMSO‑d6) δ 10.98 (s, 1H), 10.11 (s, 1H), 7.68 (s, 1H), 7.52 (s, 1H), 7.43 (s, 1H), 7.34 (s, 1H), 7.10 (s, 1H), 6.97 (s, 2H), 6.13 (s, 1H), 5.15 (s, 1H), 4.63 (s, 1H), 3.71 (d, J = 88.3 Hz, 6H), 3.35 (s, 5H), 2.75 (s, 2H), 2.50 (s, 2H), 2.36 (d, J = 23.1 Hz, 5H), 2.23 (s, 1H), 1.87 (s, 2H). 13C NMR (101 MHz, DMSO‑d6) δ 178.24, 177.12, 172.49, 170.38, 163.86, 159.26, 146.84, 145.15, 138.71 (d, J = 25.0 Hz), 136.66, 133.61, 129.83, 128.26, 125.83, 124.68, 121.88, 121.08, 120.85, 120.03, 118.87, 118.39, 114.83, 111.59, 106.72, 69.23, 56.52, 56.20, 36.56, 35.14, 32.80, 32.15, 20.49, 19.65, 9.11. MS (ESI, positive) found (M+H) 649.22, calc (C36H35N5O7, m/z): 649.69. HPLC (MeOH:H2O = 70:30): 99.139 %.
3.2.7.11 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 6-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-6-oxohexanoate (11c)
1H NMR (400 MHz, DMSO‑d6) δ 11.19 (s, 1H), 9.90 (s, 1H), 8.04 (s, 1H), 7.49 (s, 1H), 7.35 (s, 1H), 7.11 (s, 1H), 7.10 (s, 1H), 7.01 (s, 1H), 6.94 (s, 1H), 6.04 (s, 1H), 5.75 (s, 1H), 5.14 (s, 2H), 3.83 (s, 3H), 3.73 (s, 3H), 3.17 (s, 2H), 2.86 (s, 3H), 2.62 (s, 2H), 2.28 (s, 4H), 2.23 (s, 3H), 1.56 (s, 4H). 13C NMR (101 MHz, DMSO‑d6) δ 178.65, 177.52, 173.12, 172.02, 164.67, 159.69, 149.58, 144.44, 138.96, 136.83, 131.76, 129.23, 128.70, 126.54, 122.23, 121.31, 119.33, 118.58, 115.31, 113.79, 112.08, 111.79, 111.02, 107.16, 106.01, 70.83, 56.96, 56.57, 37.00, 36.61, 33.62, 32.57, 24.86, 24.55, 19.87, 9.53. MS (ESI, positive) found (M+2H) 664.27, calc (C37H37N5O7, m/z): 663.72. HPLC (MeOH:H2O = 70:30): 98.468 %.
3.2.7.12 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 3-((3-((14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-3-oxopropyl)disulfaneyl)propanoate (11d)
1H NMR (400 MHz, DMSO‑d6) δ 11.18 (s, 1H), 10.05 (s, 1H), 8.04 (s, 1H), 7.76 (d, J = 11.1 Hz, 1H), 7.49 (d, J = 7.8 Hz, 1H), 7.36 (d, J = 8.1 Hz, 1H), 7.14–7.07 (m, 2H), 7.01 (t, J = 7.4 Hz, 1H), 6.04 (s, 1H), 5.73 (s, 1H), 5.16 (s, 2H), 3.81 (s, 3H), 3.73 (s, 3H), 3.36 (s, 6H), 3.23–3.12 (m, 1H), 2.94 (dt, J = 14.7, 6.9 Hz, 4H), 2.68 (t, J = 6.5 Hz, 4H), 2.22 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.62, 177.53, 171.53, 169.25, 164.27, 159.68, 145.68, 139.07, 137.11, 133.87, 130.27, 128.66, 126.28, 125.10, 122.33, 121.48, 121.29, 120.42, 119.30, 118.87, 118.74, 115.05, 112.08, 112.03, 107.15, 69.69, 56.96, 36.99, 36.34, 34.04, 33.88, 33.26, 32.58, 20.11, 9.59. MS (ESI, positive) found (M−H) 726.20, calc (C37H37N5O7S2, m/z): 727.85. HPLC (MeOH:H2O = 70:30): 98.960 %.
3.2.7.13 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 4-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-4-oxobutanoate (12a)
1H NMR (400 MHz, DMSO‑d6) δ 11.01 (s, 1H), 10.02 (s, 1H), 8.01 (s, 1H), 7.74 (s, 1H), 7.24 (s, 1H), 7.07 (s, 1H), 6.77 (s, 1H), 6.59 (s, 1H), 6.01 (s, 1H), 5.71 (s, 1H), 5.15 (s, 1H), 4.65 (s, 1H), 3.76 (t, J = 10.9 Hz, 9H), 3.59 (s, 2H), 2.94 (s, 3H), 2.83 (s, 2H), 2.63 (s, 2H), 2.59 (s, 2H), 2.21 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.62, 177.55, 173.29, 172.57, 170.01, 169.94, 164.28, 159.67, 153.82, 149.57, 145.53, 139.03, 132.17, 130.89, 128.62, 126.59, 121.49, 121.28, 120.36, 118.74, 118.66, 115.25, 112.75, 112.46, 111.84, 107.12, 100.65, 69.74, 56.93, 56.62, 55.84, 51.81, 32.54, 31.22, 29.27, 29.01, 20.20, 9.54. MS (ESI, positive) found (M+H) 666.25, calc (C36H35N5O8, m/z): 665.69. HPLC (MeOH:H2O = 70:30): 98.904 %.
3.2.7.14 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 5-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-5-oxopentanoate (12b)
1H NMR (400 MHz, DMSO‑d6) δ 11.00 (s, 1H), 9.88 (s, 1H), 8.03 (s, 1H), 7.74 (s, 1H), 7.23 (s, 1H), 7.08 (s, 1H), 6.99 (s, 1H), 6.77 (s, 1H), 6.01 (s, 1H), 5.74 (s, 1H), 5.13 (s, 2H), 3.82 (s, 3H), 3.76 (s, 3H), 3.73 (s, 3H), 3.58 (s, 2H), 3.16 (s, 3H), 2.83 (s, 2H), 2.63 (s, 2H), 2.28 (s, 2H), 2.27 (s, 2H), 2.22 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.65, 177.53, 173.14, 171.18, 164.30, 159.70, 153.81, 145.54, 138.99, 134.08, 132.17, 130.91, 128.69, 126.58, 125.09, 121.47, 121.30, 120.36, 118.81, 115.32, 112.75, 112.45, 111.83, 107.16, 100.64, 69.76, 56.96, 56.56, 55.84, 36.97, 36.35, 33.62, 32.58, 25.00, 24.56, 20.20, 9.55. MS (ESI, positive) found (M+H) 680.27, calc (C37H37N5O8, m/z): 679.72. HPLC (MeOH:H2O = 70:30): 97.319 %.
3.2.7.15 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 6-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-6-oxohexanoate (12c)
1H NMR (400 MHz, DMSO‑d6) δ 10.99 (s, 1H), 10.03 (s, 1H), 8.03 (d, J = 2.3 Hz, 1H), 7.75 (dd, J = 8.7, 2.3 Hz, 1H), 7.24 (d, J = 8.8 Hz, 1H), 7.08 (d, J = 8.8 Hz, 1H), 6.98 (d, J = 1.9 Hz, 1H), 6.75 (dd, J = 8.8, 2.2 Hz, 1H), 6.02 (s, 1H), 5.75 (d, J = 2.0 Hz, 1H), 5.17 (s, 2H), 3.83 (s, 3H), 3.75 (d, J = 5.9 Hz, 6H), 3.17 (d, J = 5.1 Hz, 2H), 2.94 (dt, J = 14.7, 6.9 Hz, 5H), 2.71–2.63 (m, 8H), 2.23 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.66, 177.56, 169.23, 164.26, 159.71, 153.81, 145.64, 139.10, 133.76, 132.16, 130.93, 128.68, 126.59, 125.13, 121.36, 120.25, 118.89, 115.07, 112.75, 112.45, 111.83, 107.17, 100.63, 69.77, 56.98, 55.84, 36.95, 36.34, 34.05, 33.88, 33.26, 32.60, 20.19, 9.60. MS (ESI, positive) found (M+H) 694.29, calc (C38H39N5O8, m/z): 693.74. HPLC (MeOH:H2O = 70:30): 95.095 %.
3.2.7.16 (5-Methoxy-1,2-dimethyl-4,7-dioxo-4,7-dihydro-1H-indol-3-yl)methyl 3-((3-((10-methoxy-14-methyl-5-oxo-5,7,8,13,13b,14-hexahydroindolo[2′,3′:3,4]pyrido[2,1-b]quinazolin-3-yl)amino)-3-oxopropyl)disulfaneyl)propanoate (12d)
1H NMR (400 MHz, DMSO‑d6) δ 11.00 (s, 1H), 9.89 (s, 1H), 8.03 (d, J = 2.3 Hz, 1H), 7.75 (dd, J = 8.7, 2.4 Hz, 1H), 7.24 (d, J = 8.8 Hz, 1H), 7.07 (d, J = 8.8 Hz, 1H), 6.99 (d, J = 2.1 Hz, 1H), 6.75 (dd, J = 8.8, 2.3 Hz, 1H), 6.01 (s, 1H), 5.74 (s, 1H), 5.13 (s, 2H), 3.82 (s, 3H), 3.74 (d, J = 10.0 Hz, 6H), 3.17 (d, J = 4.9 Hz, 4H), 2.85–2.78 (m, 2H), 2.63 (s, 3H), 2.27 (q, J = 7.3 Hz, 4H), 2.22 (s, 3H). 13C NMR (101 MHz, DMSO‑d6) δ 178.65, 177.53, 173.14, 171.18, 164.30, 159.70, 153.81, 145.55, 138.99, 134.09, 132.17, 130.91, 128.69, 126.58, 125.08, 121.48, 121.30, 120.36, 118.82, 115.32, 112.75, 112.45, 111.83, 107.16, 100.64, 69.76, 56.96, 56.55, 55.84, 49.06, 36.97, 36.34, 33.62, 32.58, 25.00, 24.56, 20.20, 9.55. MS (ESI, positive) found (M+H) 758.23, calc (C38H39N5O8, m/z): 757.87. HPLC (MeOH:H2O = 70:30): 98.566 %.
3.3 Cell viability assays
A549, H460, PC9, PC9/GR, and LO2 cell lines were acquired from Procell Life Science & Technology Co. Ltd. (Wuhan, China). The A549, H460, PC9, and LO2 cells were maintained in Roswell Park Memorial Institute (RPMI)-1640 medium. The PC9/GR cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) containing 10 % (v/v) fetal bovine serum (FBS), 100 U/mL penicillin, and 100 mg/mL streptomycin. For the viability assays, 4000 cells were seeded in each well of a 96-well plate. The cells were allowed to attach for 12–24 h, subjected to the indicated inhibitors for 48 h, and incubated with 0.5 mg/mL MTT for 4 h. Cell viability was then determined from OD492. All experiments were performed in triplicate.
3.4 Effect of NQO1 enzyme on the stability of compound 11b
A stock solution of 11b (10 mM) was prepared in DMSO and diluted using a PBS solution. Compound 11b (10 mM) was added to PBS (10 mM; pH = 7.4) buffer solution configured to 50 μM, and NADPH (100 μM) and NQO1 (10 mg/mL) were added. The group without the NQO1 enzyme was incubated at 37 °C as a control. Samples were taken at 0, 2, 6, 12, and 24 h; incubation was terminated by adding 100 µL of cold acetonitrile. HPLC was performed on Shimadzu liquid chromatograph LC-2030 Plus equipment. Mass spectrometry was performed on Shimadzu LCMS-2020. The mobile phase was acetonitrile: the linear gradient of water was 40:60 (v/v), the flow rate was 0.5 mL/min, and the detection wavelength was 265 nm.
3.5 Cell apoptosis analysis
A549 cells (2.5 × 105/well) were seeded in six-well plates for 12–24 h. After adhesion, the cells were subjected to 2 µM, 4 µM, and 8 µM of compounds 11b and 12d for 48 h, harvested by trypsinization, and washed twice with cold phosphate-buffered saline (PBS). The cells were then centrifuged and their supernatants were removed. The washed cells were resuspended in 1 × binding buffer (500 µL) (Beyotime), subjected to Annexin V-FITC (5 µL), and incubated at room temperature for 5 min. The cells were then subjected to propidium iodide (PI; 10 µL) and incubated in the dark at room temperature for 5 min. The stained cells were analyzed by flow cytometry (NovoCyte; Agilent Technologies, Santa Clara, CA, USA).
3.6 Cell cycle analysis
A549 cells (2.5 × 105/well) were seeded in six-well plates for 12–24 h. After adhesion, the cells were subjected to 2 µM, 4 µM, and 8 µM of compounds 11b and 12d for 48 h, harvested by trypsinization, and washed with cold PBS. The cells were then centrifuged and their supernatants were removed. Then 1 mL ice was used to pre-cool 70 % (v/v) ethanol and the latter was used to fix the cells at 4 °C for >2h. Staining buffer (0.5 mL) (Beyotime), propidium iodide (PI) staining solution (20×; 25 µL), and RNase A (50X; 10 µL) were added to the cells and they were incubated in the dark at room temperature for 30 min. The stained cells were analyzed by flow cytometry. (NovoCyte; Agilent Technologies, Santa Clara, CA, USA).
3.7 ROS burst assay
A549 cells (3 × 105/well) were inoculated into six-well plates and attached overnight before treatment with diverse concentrations of specified compounds for 48 h. Then, the cell culture medium was discarded and cells were dyed with 10 µM DCFH-DA (Beyotime) dye for another 30 min at 37 °C. Next, labeled cells were cleaned using PBS three times. Eventually, the cells were resuspended with 500 µL of PBS before using flow cytometry. (BD Accuri C6).
3.8 Western blotting assay
A549 cells were seeded in six-well plates at a density of 2 × 105 cells/well. Then the cells were treated with compounds in different concentrations and cultured at 37 °C for 24 h. Proteins were extracted by lysis buffer and stored at −20 °C. The protein concentrations were quantified by the BCA Protein Concentration Detection Kit. Then the proteins were separated by 10 % sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto polyvinylidene difluoride (PVDF). The membranes were blocked with 5 % non-fat milk in TBST (Tris buffered saline with 0.1 % Tween-20) for 2 h and incubated with primary antibodies at 4 °C overnight under gentle shaking. Then, the membranes were washed with TBST and further incubated with the secondary antibodies at 37 °C for 1.5 h. All membranes were washed with TBST three times for 30 min and protein blots were detected with chemiluminescence reagent (Thermo Fischer Scientifics Ltd.) and Tanon automated chemiluminescence imaging analysis system.
3.9 Molecular docking
The crystal structure of NQO1 (PDB: 2F1O) was obtained from a protein database. The protein preparation tool in Maestro ver. 11.5 was used for docking. Ligands and water were removed from the structure and hydrogens were added to it. Staged minimization was performed using the default setting. All docking studies were conducted in Maestro ver. 11.5. The image representing the best pose was prepared with PyMol (https://www.pymol.org/).
3.10 In vivo anti-tumor evaluation
Six-week-old male C57 mice were purchased from Chengdu Dashuo Experimental Animal Co. LTD. Adaptive feeding 1 week later, a total of 1 × 106 LCC cells were subcutaneously inoculated into the right flank of C57 mice according to protocols of tumor transplant research, to initiate tumor growth. After the subcutaneous transplantation tumor formation, the transplantation tumor model C57 mice were randomly grouped into 6 mice per group. The groups treated with 11b were administered in a vehicle of 10 % DMSO/2% Tween 80/88 % saline every day by intraperitoneal injection, respectively. The positive control groups were treated with EVO every day and 5-Fu every day by intraperitoneal injection, respectively. The negative control group received a vehicle of 10 % DMSO/2% Tween 80/88 % saline through intraperitoneal injection. The mice were sacrificed after the treatments for a total of 13 consecutive days and the tumors were excised and weighed. The inhibition rate was calculated as follows: Tumor inhibitory ratio (%) = (1-average tumor weight of treated group/average tumor weight of control group) × 100 %. After execution, liver tissue, kidney tissue, and tumor tissue specimens were taken, fixed by 4 % paraformaldehyde, dehydrated in ethanol, embedded in paraffin, stained with HE, and observed microscopically. Paraffin sections of tumor tissues were eluted, incubated with antibodies, colored with DAB, restained with hematoxylin, observed on the immunohistochemical scanner, and saved with the corresponding results.
3.11 Statistical analysis
Data are means ± standard deviation (SD) for three independent experiments. T-test was used to identify statistically significant differences among cells exposed to the various target compounds and control drugs. GraphPad Prism ver. 8.0.2 (GraphPad Software, Inc., La Jolla, CA, USA) was utilized for all statistical analyses and graph plotting. Differences were considered statistically significant at p < 0.05.
4 Conclusion
In conclusion, according to previous studies, the NQO1 protein is highly expressed in the tumor microenvironment. Based on this finding, we designed and synthesized a series of EVO derivatives with NQO1 targeting specificity. In vitro, anti-proliferative activity results showed that compounds 11b and 12d exhibited anticancer activity against A549 cells (with IC50 values of 2.72 and 3.80 µM, respectively), which was significantly stronger than the parent compound EVO. Flow cytometry analysis results demonstrated that compounds 11b and 12d promoted the apoptosis of A549 cells, arrested the cell cycle at the G2/M phase and induced reactive oxygen species bursts. Further mechanistic studies revealed that compounds 11b and 12d could agonist the NQO1 pathway, thereby inducing intracellular oxidative stress responses and promoting cell apoptosis. Given that NQO1 is a well-known antioxidant enzyme that is involved in the detoxification of ROS (Ross and Siegel, 2021). Therefore, it seems counterintuitive that its activation would lead to increased ROS production. However, NQO1 has a diverse functionality. As a phase II enzyme, during the detoxification of some substrates, NQO1 mediated a large amount of ROS generation. It has been well established that β-lapachone induced cell death was mediated by excessive ROS through NQO1 (Oh and Park, 2015). Interestingly, NQO1 is overexpressed in providing an opportunity to preferentially damage cancers relative to normal tissues, using bioreductive anticancer drugs. Given that chemically 11b and 12d are indoquinones, we speculated that it might have a similar mechanism to that of β-lapachone. Importantly, compound 11b significantly inhibited tumor growth in the LCC xenograft model (TGI = 39.13 %), with no significant difference compared to 5-Fu. All of the above results indicate that compounds 11b and 12d are potential drug candidates for non-small cell lung cancer and deserve further investigation.
Author Contributions
Z. Y., H. G. and J. Z. conceived and designed the experiments; Z. Y. and B.-B. W. performed the experiments; Y.-W. W., R.-Y. J. and W.-Z. C. analyzed the data; Y.-P. T. contributed reagents/materials/analysis tools; B.-B. W., Z. Y. and H. G. wrote the paper.
CRediT authorship contribution statement
BinBin Wei: Writing – original draft. Zheng Yang: Writing – original draft. Hui Guo: Writing – review & editing, Supervision. YuWei Wang: Software, Conceptualization. WenZhuo Chen: Formal analysis. Jing Zhou: Methodology, Funding acquisition. RuYi Jin: Project administration. Zheng Wang: Writing – original draft. YuPing Tang: Resources, Investigation, Funding acquisition.
Acknowledgments
This research was financially supported by the National Natural Science Foundation of China (82003653, 81001669). Shaanxi Provincial Health Research Fund project (2022A018). Subject Innovation Team of Shaanxi University of Chinese Medicine (2019-PY02). Quality Improvement Project for Postgraduates of Shaanxi University of Chinese Medicine (JGCX006). Xianyang Innovation Team of Innovation Capability Support Program (L2023-CXNL-CXTD-004).
Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- The roles of cellular reactive oxygen species, oxidative stress and antioxidants in pregnancy outcomes. Int. J. Biochem. Cell Biol.. 2010;42:1634-1650.
- [CrossRef] [Google Scholar]
- Developing new ceramide analogs against non-small cell lung cancer (NSCLC) Am. J. Cancer Res.. 2024;14:86-96.
- [CrossRef] [Google Scholar]
- KEAP1/NRF2 (NFE2L2) mutations in NSCLC - Fuel for a superresistant phenotype? Lung Cancer. 2021;159:10-17.
- [CrossRef] [Google Scholar]
- New tricks for an old natural product: discovery of highly potent evodiamine derivatives as novel antitumor agents by systemic structure-activity relationship analysis and biological evaluations. J. Med. Chem.. 2012;55:7593-7613.
- [CrossRef] [Google Scholar]
- Clinical and pathological characteristics of KEAP1- and NFE2L2-mutated non-small cell lung carcinoma (NSCLC) Clin. Cancer Res.. 2018;24:3087-3096.
- [CrossRef] [Google Scholar]
- Structural dynamics at the active site of the cancer‐associated flavoenzyme NQO1 probed by chemical modification with PMSF. FEBS Lett.. 2023;597:2687-2698.
- [CrossRef] [Google Scholar]
- Recent advances in the application of podophyllotoxin derivatives to fight against multidrug-resistant cancer cells. Curr. Top. Med. Chem.. 2021;21:1712-1724.
- [CrossRef] [Google Scholar]
- Epigenetic regulation of Keap1-Nrf2 signaling. Free Radic. Biol. Med.. 2015;88:337-349.
- [CrossRef] [Google Scholar]
- Rational design of a new self-codelivery system from redox-sensitive camptothecin-cytarabine conjugate assembly for effectively synergistic anticancer therapy. Adv. Healthc. Mater.. 2017;6
- [CrossRef] [Google Scholar]
- Anti-proliferative effects of evodiamine in human lung cancer cells. J. Cancer Prev.. 2014;19:7-13.
- [CrossRef] [Google Scholar]
- Evodiamine suppresses non-small cell lung cancer by elevating CD8+ T cells and downregulating the MUC1-C/PD-L1 axis. J. Exp. Clin. Cancer Res.. 2020;39:249.
- [CrossRef] [Google Scholar]
- Bax directly induces release of cytochrome c from isolated mitochondria. Proc. Natl. Acad. Sci. U. S. A.. 1998;95:4997-5002.
- [CrossRef] [Google Scholar]
- Evodiamine eliminates colon cancer stem cells via suppressing Notch and Wnt signaling. Molecules. 2019;24:4520.
- [CrossRef] [Google Scholar]
- Mitochondria-Associated Membranes (MAMs) are involved in Bax mitochondrial localization and cytochrome c release. Microb. Cell. 2019;6:257-266.
- [CrossRef] [Google Scholar]
- Evodiamine alleviates DEHP-induced hepatocyte pyroptosis, necroptosis and immunosuppression in grass carp through ROS-regulated TLR4 / MyD88 / NF-κB pathway. Fish Shellfish Immunol.. 2023;140:108995
- [CrossRef] [Google Scholar]
- Targeting NAD(P)H:quinone oxidoreductase (NQO1) in pancreatic cancer. Mol. Carcinog.. 2017;56:1825-1834.
- [CrossRef] [Google Scholar]
- Histone deacetylase 10 regulates the cell cycle G2/M phase transition via a novel let-7-HMGA2-cyclin A2 pathway. Mol. Cell Biol.. 2015;35:3547-3565.
- [CrossRef] [Google Scholar]
- Traditional uses, phytochemistry, pharmacology, pharmacokinetics and toxicology of the fruit of Tetradium ruticarpum: A review. J. Ethnopharmacol.. 2020;263:113231
- [CrossRef] [Google Scholar]
- Evodiamine suppresses the progression of non-small cell lung carcinoma via endoplasmic reticulum stress-mediated apoptosis pathway in vivo and in vitro. Int. J. Immunopathol. Pharmacol.. 2022;36
- [CrossRef] [Google Scholar]
- Evodiamine augments NLRP3 inflammasome activation and anti-bacterial responses through inducing α-tubulin acetylation. Front. Pharmacol.. 2019;10:290.
- [CrossRef] [Google Scholar]
- Effect of evodiamine on the proliferation and apoptosis of A549 human lung cancer cells. Mol. Med. Rep.. 2016;14:2832-2838.
- [CrossRef] [Google Scholar]
- Evodiamine induces ROS-dependent cytotoxicity in human gastric cancer cells via TRPV1/Ca2+ pathway. Chem. Biol. Interact.. 2022;351:109756
- [CrossRef] [Google Scholar]
- Synthesis and characterization of biogenic metal nanoparticles and its cytotoxicity and anti-neoplasticity through the induction of oxidative stress, mitochondrial dysfunction and apoptosis. Colloids Surf. B Biointerfaces. 2018;161:111-120.
- [CrossRef] [Google Scholar]
- An A-ring substituted evodiamine derivative with potent anticancer activity against human non-small cell lung cancer cells by targeting heat shock protein 70. Biochem. Pharmacol.. 2023;211:115507
- [CrossRef] [Google Scholar]
- ROS signalling in the biology of cancer. Semin. Cell Dev. Biol.. 2018;80:50-64.
- [CrossRef] [Google Scholar]
- Cyclin A/Cdk2 regulates Cdh1 and claspin during late S/G2 phase of the cell cycle. Cell Cycle. 2014;13:3302-3311.
- [CrossRef] [Google Scholar]
- Addressing CPI resistance in NSCLC: targeting TAM receptors to modulate the tumor microenvironment and future prospects. J. Immunother. Cancer. 2022;10:e004863
- [CrossRef] [Google Scholar]
- Review on NAD(P)H dehydrogenase quinone 1 (NQO1) pathway. Mol. Biol. Rep.. 2022;49:8907-8924.
- [CrossRef] [Google Scholar]
- Synthesis and biological evaluation of NQO1-activated prodrugs of podophyllotoxin as antitumor agents. Bioorg. Med. Chem.. 2020;28:115821
- [CrossRef] [Google Scholar]
- Interactions of the antioxidant enzymes NAD(P)H: Quinone oxidoreductase 1 (NQO1) and NRH: Quinone oxidoreductase 2 (NQO2) with pharmacological agents, endogenous biochemicals and environmental contaminants. Chem. Biol. Interact.. 2021;345:109574
- [CrossRef] [Google Scholar]
- The diverse functionality of NQO1 and its roles in redox control. Redox Biol.. 2021;41:101950
- [CrossRef] [Google Scholar]
- Novel hydrophilic camptothecin derivatives conjugated to branched glycerol trimer suppress tumor growth without causing diarrhea in murine xenograft models of human lung cancer. Mol. Pharm.. 2020;17:1049-1058.
- [CrossRef] [Google Scholar]
- A novel evodiamine amino derivative as a PI3K/AKT signaling pathway modulator that induces apoptosis in small cell lung cancer cells. Eur. J. Pharmacol.. 2021;906:174215
- [CrossRef] [Google Scholar]
- Fluoride-induced oxidative stress and apoptosis are involved in the reducing of oocytes development potential in mice. Chemosphere. 2017;186:911-918.
- [CrossRef] [Google Scholar]
- Amelioration of AOM/DSS-induced murine colitis-associated cancer by evodiamine intervention is primarily associated with gut microbiota-metabolism-inflammatory signaling axis. Front. Pharmacol.. 2021;12:797605
- [CrossRef] [Google Scholar]
- Design, synthesis, and biological evaluation of benzenesulfonyl chloride-substituted evodiamine derivatives as potential PGAM1 inhibitors. Arab. J. Chem.. 2023;16:105295
- [CrossRef] [Google Scholar]
- Design, synthesis, and biological evaluation of NAD(P)H: Quinone oxidoreductase (NQO1)-targeted oridonin prodrugs possessing indolequinone moiety for hypoxia-selective activation. Eur. J. Med. Chem.. 2017;132:310-321.
- [CrossRef] [Google Scholar]
- Research advances in NQO1-responsive prodrugs and nanocarriers for cancer treatment. Future Med. Chem.. 2022;14:363-383.
- [CrossRef] [Google Scholar]
- A novel class I histone deacetylase inhibitor, I-7ab, induces apoptosis and arrests cell cycle progression in human colorectal cancer cells. Biomed. Pharmacother.. 2015;71:70-78.
- [CrossRef] [Google Scholar]
- Synthesis and antitumor activity of evodiamine derivatives with nitro, amino, and methoxy groups. Nat. Prod. Commun.. 2022;17
- [CrossRef] [Google Scholar]
- Research progress on the source, production, and anti-cancer mechanisms of paclitaxel. Chin. J. Nat. Med.. 2020;18:890-897.
- [CrossRef] [Google Scholar]
- Induction of apoptosis and effect on the FAK/AKT/mTOR signal pathway by evodiamine in gastric cancer cells. Curr. Issues Mol. Biol.. 2022;44:4339-4349.
- [CrossRef] [Google Scholar]
- Anti-cancer effects of epigenetics drugs scriptaid and zebularine in human breast adenocarcinoma cells. Anticancer Agents Med Chem.. 2022;22:1582-1591.
- [CrossRef] [Google Scholar]
- β-Caryophyllene ameliorates cyclophosphamide induced cardiac injury: the association of TLR4/NFκB and Nrf2/HO1/NQO1 pathways. J Cardiovasc Dev Dis. 2022;9:133.
- [CrossRef] [Google Scholar]
- Upregulation of wild-type p53 by small molecule-induced elevation of NQO1 in non-small cell lung cancer cells. Acta Pharmacol. Sin.. 2022;43:692-702.
- [CrossRef] [Google Scholar]
- NAD(P)H:quinone oxidoreductase 1 (NQO1) as a therapeutic and diagnostic target in cancer. J. Med. Chem.. 2018;61:6983-7003.
- [CrossRef] [Google Scholar]
- NQO1-selective activated prodrugs of combretastatin A-4: synthesis and biological evaluation. Bioorg. Chem.. 2020;103:104200
- [CrossRef] [Google Scholar]
- Bioreduction activated prodrugs of camptothecin: molecular design, synthesis, activation mechanism and hypoxia selective cytotoxicity. Org. Biomol. Chem.. 2005;3:1905-1910.
- [CrossRef] [Google Scholar]
- Design and synthesis of NAD(P)H: Quinone oxidoreductase (NQO1)-activated prodrugs of 23-hydroxybetulinic acid with enhanced antitumor properties. Eur. J. Med. Chem.. 2022;240:114575
- [CrossRef] [Google Scholar]
Appendix A
Supplementary data
Supplementary data (Data sets supporting the conclusions of this article are included in this article (and its additional files)) to this article can be found online at https://doi.org/10.1016/j.arabjc.2024.106075.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary Data 1
Supplementary Data 1
Data sets supporting the conclusions of this article are included in this article (and its additional files).
Supplementary Data 2
Supplementary Data 2







