Translate this page into:
Inhibition mechanism of silver nanoparticle-kaempferol against methicillin-resistant Staphylococcus aureus
⁎Corresponding author at: Department of Basic Sciences, Faculty of Dentistry, Universiti Sains Islam Malaysia, Kuala Lumpur 55100, Malaysia. rohazila@usim.edu.my (Rohazila Mohamad Hanafiah)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Methicillin-resistant Staphylococcus aureus (MRSA), a multidrug resistant strain, is known to cause a threat to public health due to its limited therapeutic treatment. Kaempferol (K) is a natural flavonoid that shows antibacterial activities toward MRSA, but its effectiveness is limited due to its low water solubility. However, poorly aqueous soluble drugs displayed better solubility through nano formulation. Hence, kaempferols were incorporated with silver nanoparticles (AgNPs) to enhance their solubility and antibacterial activity. Previous study showed that AgNPs incorporated with kaempferol (AgNPs-K) exhibited antibacterial activity against MRSA. However, the knowledge regarding the mechanism of action AgNPs-K against MRSA is still limited. The objective of the study is to unravel the inhibition mechanism of silver nanoparticles-kaempferol (AgNPs-K) on treated MRSA. The scanning electron microscopy (SEM) result showed significant difference in morphology between treated and non-treated MRSA which suggest the effectiveness of the AgNPs-K. Non-treated MRSA has an oval shape while MRSA treated with AgNPs-K showed a disrupted cell wall with contents leakage. The transcriptomic profile analysis by Next Generation Sequencing (NGS) showed that various genes and pathways related to biofilm, virulent activity and glycolysis pathway are differently expressed, with 581 genes were downregulated and 641 were upregulated. The affected genes of icab, clfa and eno which involved in biofilm, clumping factor A (virulent) and glycolysis pathway were validated by RT-PCR technique. The results were consistent with the NGS outcome. In conclusion, AgNPs-K possesses antibacterial activity against MRSA and its mechanism of action are reflected in the gene expression of biofilm pathway, virulent and glycolysis activity. Therefore, AgNPs-K can be suggested as a potential alternative to combat MRSA infection.
Keywords
Silver nanoparticles
Kaempferol
Methicillin-resistant Staphylococcus aureus
Transcriptomic profile analysis
1 Introduction
Antimicrobial resistance is still one of the major problems threatening human health. Following the advent of penicillin, Staphylococcus aureus (S. aureus) has been a prominent concern within the approach to antibiotic resistance since the 1960s (Jevons, 1961). An aerobic Gram-positive bacteria called methicillin-resistant Staphylococcus aureus (MRSA) is a subspecies strain of S. aureus, which has developed resistance to β-lactam antibiotics such as methicillin, oxacillin, and cephalosporins (Shariati et al., 2020). It is one of the opportunistic pathogens that is often linked to serious nosocomial infections. Whereas an individual is infected by antibiotic resistant bacteria, it lengthens the hospital stays and increases the cost of care. Thus, nanotechnology is suggested as one way to overcome the advancement of MRSA.
Nanotechnology is an emerging area of science dealing with nanoparticles of various materials. The materials could be particularly useful in optoelectronic devices, energy, and biomedicines (Munsif & Shah, 2023). Previous study used metals complexes to determine antibacterial activity against bacteria pathogen (Junaid et al., 2022). The high solubility and the small size of nanoparticles give advantages to cross tissue barriers of drug-resistant bacteria. Moreover, the unique attribute of nanoparticles such as their small size, various morphologies, and distribution enhance their functional properties including antioxidant, antiproliferative, antibiofilm activities, and antibacterial activities (Merghni et al., 2022). These characteristics of nanoparticles create a great potential to reduce or limit the development of new mechanisms of resistance against them.
Many different metallic nanoparticles are employed in drug delivery, fluorescent biological labels, and the detection of pathogens and proteins (Yaqoob et al., 2020). In this research, silver nanoparticles (AgNPs) were chosen as they showed excellent biological activities, such as anti-inflammatory, anti-bacterial, and antifungal effects (Fehaid et al., 2020). According to Yin et al., (2020), AgNPs exhibit antibacterial activity against Staphylococcus aureus and Streptococcus mutans. Eco-friendly synthesis of silver nanoparticles can be used as a potential strategy for multifunctional applications. Effective methods for green synthesis of metallic nanoparticles using natural resources such as plants, algae, bacteria, and fungi have been extensively reported. The effectiveness of AgNPs obtained by green synthesis against various pathogenic microorganisms is largely investigated. Moreover, plant extract functionalized AgNPs have drawn considerable attention due to their intensive biological properties, low toxicity, and good stability. One of the promising compounds from plants extract is kaempferol.
Kaempferol (K), a natural flavanol derived from plant sources is known for its antibacterial activity. Their antibacterial activities, unfortunately, are limited due to their large size and water insolubility (Santhosh Kumar et al., 2021). However, poorly aqueous soluble drugs displayed better solubility through nano formulation. From previous studies, it has been reported that the incorporation of kaempferol with other metallic nanoparticles has shown an increase of antibacterial activity. Thus, to address the issue of antibiotics-resistant bacteria, silver nanoparticles' key features can be modified with kaempferol and developed by green synthesis.
Recent study has reported that AgNPs were successfully incorporated with kaempferol and was characterize by using six instrument including UV–Visible Spectroscopy (UV–Vis), Zetasizer, transmission electron microscope (TEM), scanning electron microscope with a dispersive X-ray spectrometer (SEM-EDX), X-ray diffraction (XRD) and fourier transform infrared (FTIR) (Hairil Anuar et al., 2022). It has been reported that the formation of AgNPs-K was indicated with the presence of dark brown colour in the solution. The UV–Vis spectrum of the synthesized AgNPs-K showed a broad peak at range 350–500 nm wavelength. The results from TEM, Zetasizer and SEM-EDX reported that AgNPs-K shown a nearly spherical shape with the size of 120.4 nm and 14.42 ± 3.19 nm. During XRD analysis, it has been confirmed that AgNPs-K were in crystalline phase structure while FTIR showed (—OH) group was absent which indicated that kaempferol was successfully incorporated with AgNPs.
The study also concluded that AgNPs-K successfully exhibits antibacterial activities against methicillin-resistant Staphylococcus aureus (MRSA). In the disc diffusion (DDA) analysis, AgNPs-K showed greater antibacterial activity against MRSA than kaempferol or commercial AgNPs. The MIC and MBC values for AgNPs-K against MRSA were 1.25 mg/mL and 2.5 mg/mL, respectively. The time kill assay results showed that AgNPs-K displayed bacteriostatic activity against MRSA (Hairil Anuar et al., 2022). To continue the previous study, this study was done to unravel the inhibition mechanism of AgNPs-K against MRSA.
2 Materials and methods
2.1 Synthesis of silver Nanoparticles-Kaempferol
The synthesis of silver nanoparticles-kaempferol was done by green synthesis methods (Hairil Anuar et al., 2022). Synthesizing silver nanoparticles-kaempferol (AgNPs-K) was done in the ratio of 9:1 based on its volume. In this research, 900 mL of 1 mM aqueous silver nitrate (AgNO3) solution was added drop by drop with 100 mL of 1 mM of aqueous kaempferol (C15H10O6) solution under vigorous stirring for 30 min. Both silver nitrate and kaempferol used ultrapure water (H2O) as their solvent. The solution was then incubated in the oven (Memmert UF 110, Germany) for 168 h at 60 ˚C. The reduction of silver nitrate to silver ions was confirmed by the color changes from colorless to reddish brown. The solution was then freeze-dried (Christ Alpha 1–4 LSC basic, Germany) to obtain the AgNPs-K powder.
2.2 Preparation and validation of bacterial strains
The bacteria that were used in this research is methicillin-resistant Staphylococcus aureus (ATCC 33,591 obtained from Hospital Tuanku Ja’afar Seremban, Malaysia). The bacteria were grown on Brain Heart Infusion (BHI) agar (Oxoid Ltd., Basingstoke, UK) and then incubated aerobically at 37 ˚C for 24 h in incubator (IN-110 Memmert, German).
Next, the bacteria were validated by susceptibility test based on Clinical and Laboratory Standard Institute (CLSI) guidelines and the bacteria was tested by using cefoxitin (Oxoid Ltd.). Firstly, the bacterial suspension was prepared by transferring the bacterial colony using a sterile inoculating loop into 10 mL BHI broth (Oxoid, Ltd.). The inoculums were mixed thoroughly and were incubated at 37 ˚C for 24 h. After incubation, the turbidity of the suspension was adjusted to 0.5 McFarland. A sterile cotton swab was dipped into the inoculum and then was swabbed evenly onto BHI agar by rotating the plate. Next, the plate was allowed to dry for 60 s before transferring the antibiotic disk gently onto it by using sterile forceps. The plate was incubated at 37 ˚C for 24 h. MRSA is confirmed when the diameter of the zone inhibition is < 21 mm. The bacteria were then further validated by using Gram staining procedures and the morphology of the MRSA was confirmed by Gram-positive cocci in grape-like clusters.
3 Microscopic observation
3.1 Scanning electron microscopy (SEM) analysis
MRSA was cultured in BHI broth at 37 ˚C for 24 h and the turbidity was standardized according to 0.5 McFarland before treated with 1 × MIC and ¾×MIC, respectively. The value of MIC was referred from a previous study which is 1.25 mg/mL (Hairil Anuar et al., 2022). It was incubated at 37 ˚C depending on the time taken for the treatments to kill most MRSA in the time kill assay. After incubation, the sample was centrifuged at 4 ˚C at 4000 rpm for 15 min. This step was repeated with untreated MRSA as a control group. The pellet was collected, and the supernatant was discarded.
For primary fixation, the samples were fixed in 2.5 % glutaraldehyde for 4–6 h at 4 °C. Then, the sample was washed with 0.1 M sodium cacodylate buffer for 3 changes of 10 min each. The samples were post fixed in 1 % osmium tetroxide for 2 h at 4 °C and followed with washing step again for 3 changes of 10 min each. Next, the samples undergo dehydration in a series of acetones with different percentages which are 35 %, 50 %, 75 %, 95 % for 10 min each and was followed with 100 % acetone for 15 min with 3 changes.
For further processing, the cells suspension was pipetted onto aluminum foil coated with albumin. The specimen was transferred onto specimen basket and put into critical point dryer for about 30 min. After that, the specimen was stuck onto the stub by using double sided tape and it was coated by gold in a sputter coater. Finally, the ultrastructural changes in MRSA were observed and photographed by using JSM-IT 100 InTouchScopeTM.
3.2 RNA isolation
RNAs isolation was carried out by using SV Total RNA Isolation System, according to the manufacturer’s protocol (Promega, Wisconsin, USA). RNA isolation was done on MRSA treated with AgNPs-K and non-treated MRSA. After purification, the RNA concentration was determined by using NanoDrop 2000c spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). A260/A280 and A260/A230 ratios were used to confirm RNA isolation is free from contaminants such as proteins, phenol, and salts. Complimentary DNA (cDNA) was synthesized using ImProm-II™ reverse transcription system (Promega, USA). Standard reaction mixture with a total volume of 20 mL for RT was prepared to contain 10 mM dNTPs, 25 mM MgCl2, RNAase inhibitor, RT 200 U, oligo-dt 0.5 µg, and 1 mg total RNA from respective samples. First strand complementary DNA (cDNA) was amplified following incubation for 90 min at 42 °C and 15 min at 72 °C. The cDNA samples were stored at − 20 °C until further use.
3.3 Next generation sequencing analysis
This step was adapted according to the previous method with slight modifications (Abd Ghafar et al., 2022). It involved library preparation for transcriptome sequencing, clustering and sequencing, quality control, reading of the mapping, quantification of gene expression level, differential expression analysis, and GO and KEGG enrichment analysis of differentially expressed genes which was processed by Apical Scientific Sdn Bhd. This analysis used three samples of RNA treated with AgNPs-K and three samples of untreated RNA as controls. All the samples were analyses in triplicate. In RNA-seq, replicates serve a dual purpose. Firstly, they demonstrate the repeatability of the experiment; secondly, they can reveal differences in gene expression between samples. The correlation between samples is a crucial indicator for testing the reliability of the experiment. The closer the correlation coefficient is to 1, the greater the similarity of the samples.
3.4 Library preparation for transcriptome sequencing
A total of 1 μg RNA per sample was used as input material for the RNA sample preparations. Sequencing libraries were generated using NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA) following manufacturer’s recommendations, and index codes were added to attribute sequences to each sample. Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature to synthesis the first strand and second strand cDNA. The rest of the overhangs and ends were converted into blunt ends. Next, PCR was performed with Phusion High-Fidelity DNA polymerase, Universal PCR Primers, and Index (X) Primer. Lastly, PCR products were purified (AMPure XP system), and library quality was assessed on the Agilent Bioanalyzer 2100 system.
3.5 Clustering and sequencing
The clustering of the index-coded samples was performed on a cBot Cluster Generation System using PE Cluster Kit cBot-HS (Illumina) according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina platform and paired-end reads were generated.
3.6 Quality control
Raw data (raw reads) of FASTQ format was first processed through fastp. In this step, clean data (clean reads) was obtained by trimming reads containing adapter and removing poly-N sequences and reads with low quality from raw data. The sequenced reads/raw reads often contain low quality reads or reads with adaptors, which will affect the quality of downstream analysis. To avoid this, it’s necessary to filter the raw reads and get the clean reads. First the adaptor of contamination was removed. Then, uncertain nucleotides constitute more than 10 percent of either read (N > 10 %) had been removed. Finally, low quality nucleotides (Base Quality less than 5) constitute more than 50 % of read were removed. At the same time, Q20, Q30, and GC content of the clean data was calculated. All the downstream analyses were based on the clean data with high quality.
3.7 Reads mapping to the reference genome
Reference genome and gene model annotation files were downloaded from genome website directly. Both building index of reference genome and aligning clean reads to reference genome were conducted by using Bowtie 2 (Langmead and Salzberg, 2012).
3.8 Quantification of gene expression level
featureCounts was used to count the reads’ numbers mapped to each gene. Then, Fragments Per Kilobase of transcript per Million (FPKM) of each gene was calculated based on the length of the gene and reads’ count mapped to this gene. FPKM, expected number of fragments per kilobase of transcript sequence per million base pairs sequenced, will consider the effect of sequencing depth and gene length for the read count at the same time and is currently the most used method for estimating gene expression levels (Trapnell et al., 2010).
3.9 Differential expression analysis
For DESeq2 with biological replicates, differential expression analysis of two conditions/groups (two biological replicates per condition) was performed using the DESeq2 R package. DESeq2 provides statistical routines for determining differential expression in digital gene expression data using a model based on the negative binomial distribution. Then, the resulting p-values were adjusted using Benjamini and Hochberg’s approach for controlling the false discovery rate. Genes with adjusted p-value less than 0.05 (p < 0.05) found by DESeq2 was assigned as differentially expressed.
3.10 GO and KEGG enrichment analysis of differentially expressed genes
Gene ontology (GO) enrichment analysis of differentially expressed genes was implemented by the clusterProfiler R package, in which gene length bias was corrected. Gene Set Enrichment Analysis (GSEA) is used to determine whether the prior gene set is significantly different between two biological states (eg, phenotype. GO terms with padj < 0.05 are significant enrichment. KEGG is a database resource for understanding high-level functions and utilities of the biological system, such as the cell, the organism, and the ecosystem, from molecular level information, especially large-scale molecular datasets generated by genome sequencing and other high-throughput experimental technologies https:// https://www.genome.jp/kegg/. ClusterProfiler R package was used to test the statistical enrichment of differential expression genes in KEGG pathways.
3.11 RT-qPCR
The extraction of total RNA was performed using RNeasy mini kit (Qiagen, German). RT-qPCR were performed by using GoTaq 2-Step RT-qPCR system which facilitates detection and quantification of RNA expression levels using GoScriptTM Reverse Transcriptase and GoTaq qPCR Master Mix (Promega,USA). The sequences of primers of the analysis are shown in Table 1 based on previous research (Atshan et al., 2013). The cycling parameter had been optimized based on the average melt temperature of all primers as shown in Table 2. The expression level of gene was normalized against MRSA 16 s rRNA as a reference gene. Next, the relative quantification model by Livak and Schmittgen was used for determination of mRNA fold change after normalization to internal control (Schmittgen and Livak, 2008). Tests were done in triplicate.
Gene
Description
Direction
Sequence (5′-3′)
Amplicon size (bp)
icaB
Intercellular adhesion gene
Forward
ATA CCG GCG ACT GGG TTT AT
140
Reverse
TTG CAA ATC GTG GGT ATG TGT
clfA
Clumping factor A
Forward
ACC CAG GTT CAG ATT CTG GCA GCG
165
Reverse
TCG CTG AGT CGG AAT CGC TTG CT
eno
Laminin binding protein
Forward
TGC CGT AGG TGA CGA AGG TGG TT
195
Reverse
GCA CCG TGT TCG CCT TCG AAC T
16S rRNA
Reference gene
Forward
GGG ACC CGC ACA AGC GGT GG
191
Reverse
GGG TTG CGC TCG TTG CGG GA
Step
Cycles
Temperatures
Time
GoTaq® DNA polymerase activation
1
95 ˚C
2 min
Denaturation
1
95 ˚C
15 s
Annealing and extension
40
57 ˚C
1 min
4 Results
4.1 Scanning electron microscope (SEM) analysis for AgNPs-K against MRSA
SEM analysis showed the morphology of non treated MRSA (negative control) was in coccus shaped and linked together (Fig. 1 A). However, the morphology of MRSA changed after exposure to AgNPs-K at ¾ xMIC, 1xMIC at 8 h incubation (Fig. 1B and C) and exposure of AgNPs-K at 1xMIC at 16 h incubation (Fig. 1D). The treated MRSA undergo lysis, disrupt their cell wall, and lose their coccal appearance, releasing their cellular contents into the surrounding environment and ultimately causing cell disruption.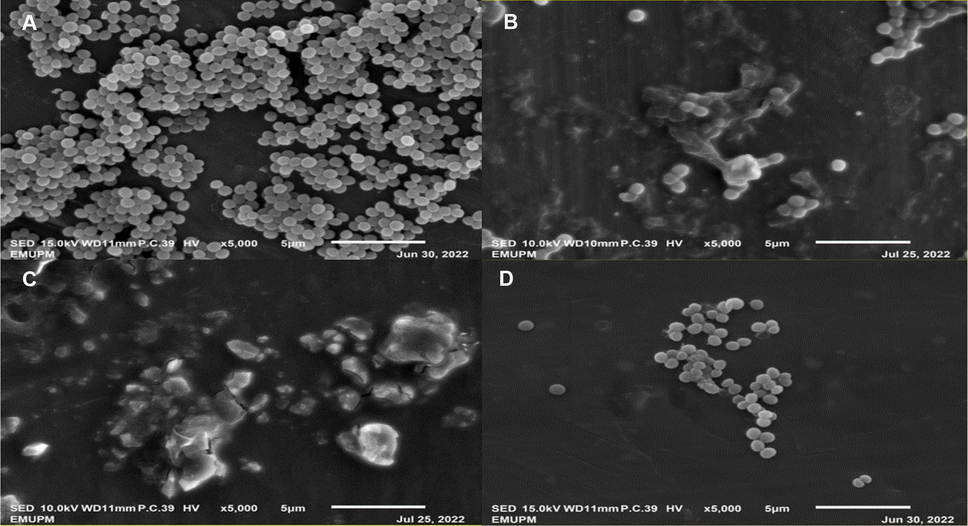
SEM images of MRSA after 16-hours incubation. (A) Negative control (untreated MRSA). (B) Treatment with ¾xMIC at 8 h. (C) Treatment with 1xMIC at 8 h. (D) treatment with 1xMIC at 16 h. Images B, C and D showed disrupted cell walls with content leakage.
4.2 Transcriptomic profile analysis
The algorithm for mapping sequences was chosen according to the characteristics of the reference genome Staphylococcus aureus subsp. aureus Rosenbach. In general, Bowtie2 was chosen for the genomes of bacteria and other species with a high gene density. The mismatch parameter is set to two, and other parameters are set to default. The total mapped rate should be more than 70 %, and the percentage of reads that can be mapped to multiple sites in the reference genome should be less than 10 %, if there is no contamination and a correct reference genome is chosen. Table 3 showed that total mapped rate is more than 70 % and the percentage of reads that can be mapped to multiple sites in the reference genome is less than 10 %, indicating no contamination of the sample and the reference genome was chosen correctly.
Sample name
A1
A2
A3
B1
B2
B3
Total reads
20,575,318
16,882,394
24,578,934
19,556,638
16,068,534
19,886,952
Total mapped reads
19,906,629
16,270,468
23,797,136
19,045,399
15,749,226
19,142,818
Uniquely mapped reads
19,592,873
16,005,372
23,416,015
18,774,781
15,528,986
18,837,398
Multiple mapped reads
313,756
265,096
381,121
270,618
220,240
305,420
Total mapping rate
96.75 %
96.38 %
96.82 %
97.39 %
98.01 %
96.26 %
Uniquely mapping rate
95.23 %
94.81 %
95.27 %
96 %
96.64 %
94.72 %
Multiple mapping rate
1.52 %
1.57 %
1.55 %
1.38 %
1.37 %
1.54 %
4.3 Differential of sequencing gene (DEG)
To understand the changes in MRSA after being treated with AgNPs-K, the transcriptome of treated MRSA was measured. In general, FPKM value of 0.1 or 1 was set as a threshold for determining whether the gene is expressed or not. The comparison for MRSA treated with AgNPs-K and untreated MRSA has been done in triplicate. RNA-sequence analysis showed that AgNPs-K has significant effect (p < 0.05) on transcriptomic profile of treated MRSA compared to control. Out of 1222 genes, 581 genes were downregulated, and 641 genes were upregulated (Fig. 2). The gene expression changes are shown in the heatmap (Fig. 3A) where it showed genes with high fold change. Red denotes genes with high expression levels, and blue denotes genes with low expression levels. The color ranges from red to blue represents the log10(FPKM + 1) value from large to small. Volcano plots are used to infer the overall distribution of differentially expressed genes (Fig. 3B). The point represents genes, blue dots indicate no significant difference in genes, red dots indicate upregulated differential expression genes, green dots indicate downregulated differential expression genes.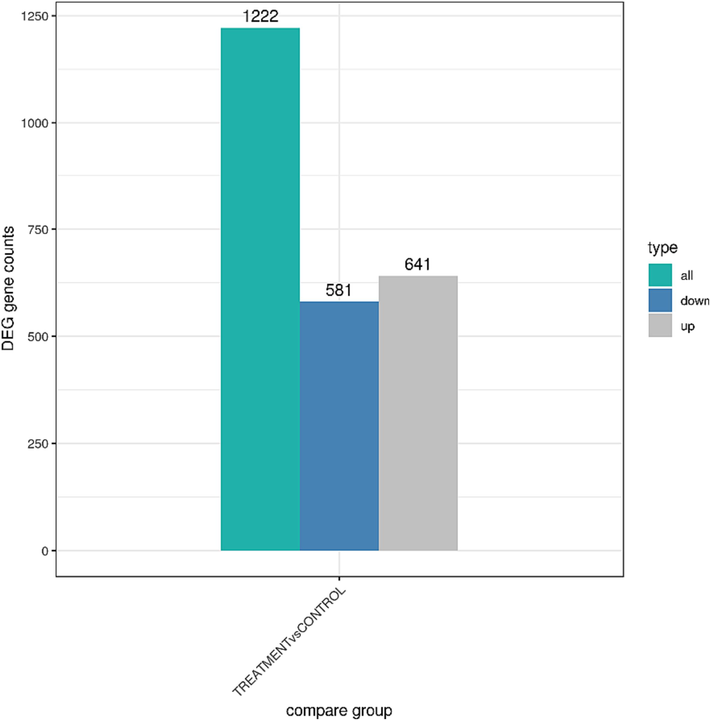
DEG gene counts of MRSA after treated with AgNPs-K (1.25 mg/mL). There are 1222 genes in total, with 581 genes were downregulated and 641 were upregulated.
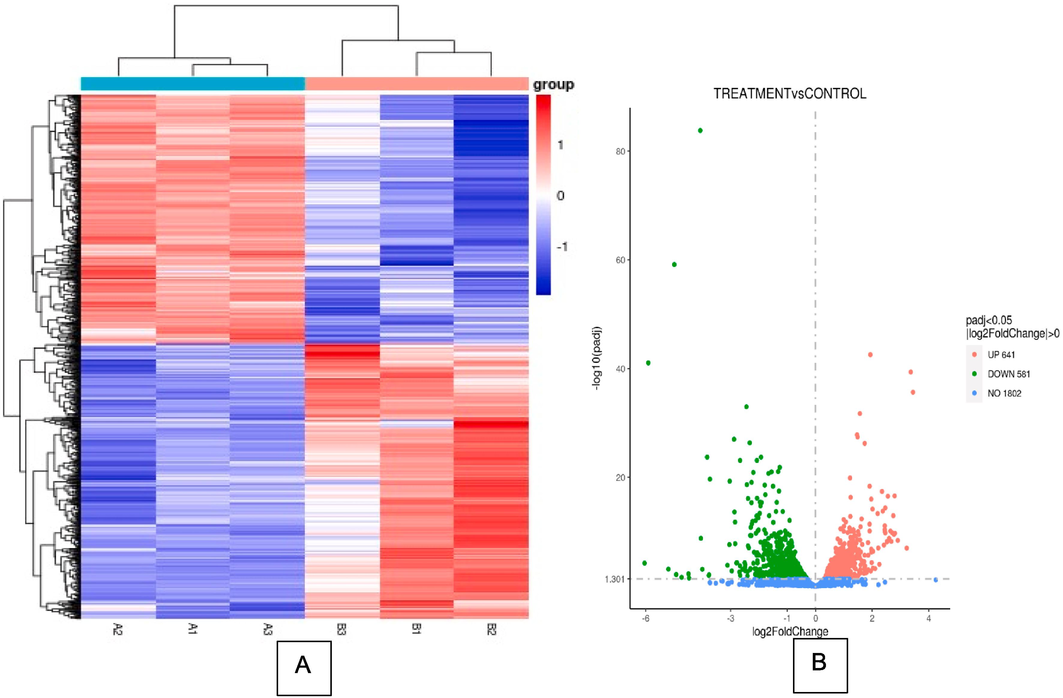
DEG analysis of MRSA treated with AgNPs-K (1.25 mg/mL). (A) Heatmap clustering of all DEGs (p value < 0.05) in MRSA treated with AgNPs-K (1.25 mg/mL). Each row represents one sample, and each column represents one gene. (B) Volcano image of gene regulation of MRSA treated with AgNPs-K (1.25 mg/mL). Green color represents downregulated genes, and red color represents upregulated genes. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
4.4 KEGG pathway analysis
The interactions of multiple genes may be involved in certain biological functions. KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis identifies significantly enriched metabolic pathways or signal transduction pathways associated with differentially expressed genes compared with the whole genome background.
KEGG Pathway analysis was used to obtain insight into the biological function of DEGs (Fig. 4A). There were some pathways different when compared to control group such as ribosomes, oxidative phosphorylation, and nitrogen metabolism were significantly enriched (p < 0.05). After being treated with AgNPs-K, the KEGG pathway was downregulated (Fig. 4B) and upregulated (Fig. 4C). Several pathways, including MRSA ribosome, Staphylococcus aureus infection, arginine biosynthesis, purine metabolism, and riboflavin metabolism were downregulated after being exposed with AgNPs-K. Meanwhile, several pathways including oxidative phosphorylation, nitrogen metabolism, microbial metabolism in diverse environments, two component system and glycolysis/gluconeogenesis were up regulated.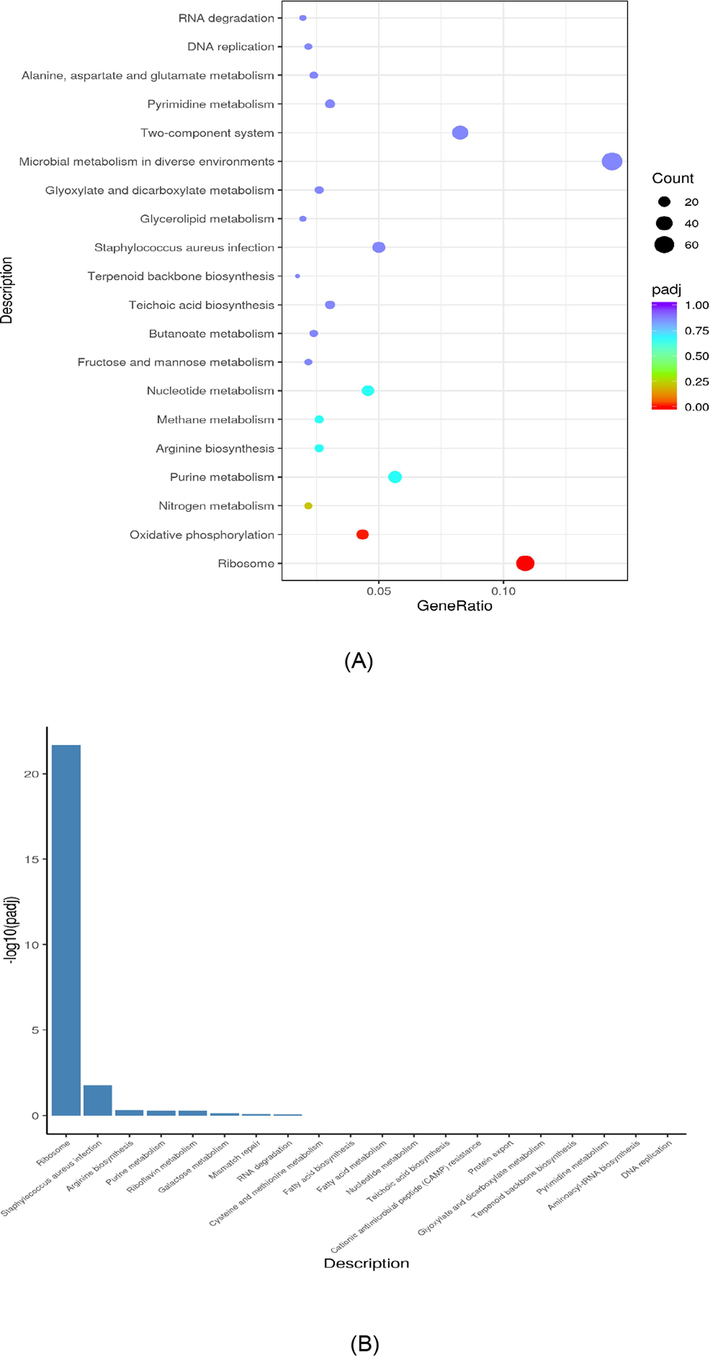
KEGG analysis of MRSA treated with AgNPs-K (1.25 mg/mL). (A) KEGG dot chart of KEGG enrichment result of trancriptome of MRSA treated with AgNPs-K (1.25 mg/mL). The size of every spot represents the number of the differential expression genes and the color of every spot represents the range of Qvalue. (B) Downregulated gene of KEGG pathway of MRSA after treated with AgNPs-K (1.25 mg/mL). (C) Upregulated gene of KEGG pathway of MRSA after treated with AgNPs-K (1.25 mg/mL).
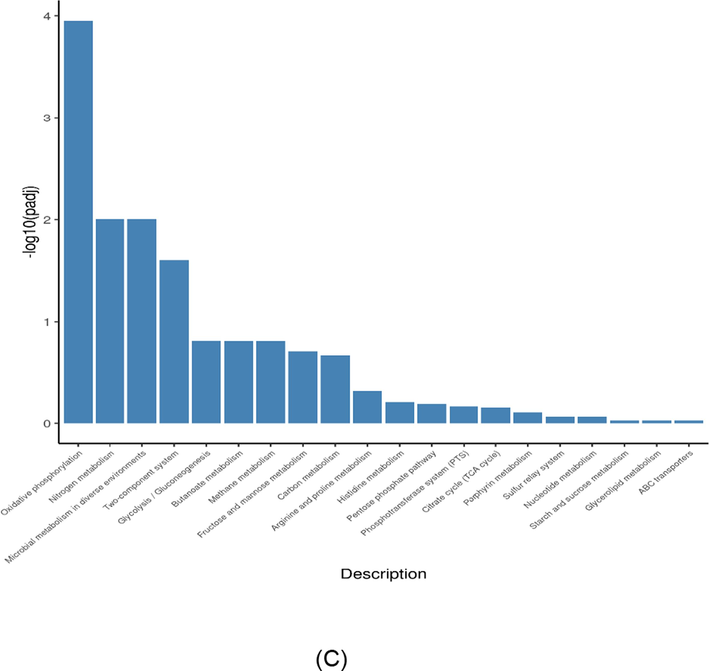
KEGG analysis of MRSA treated with AgNPs-K (1.25 mg/mL). (A) KEGG dot chart of KEGG enrichment result of trancriptome of MRSA treated with AgNPs-K (1.25 mg/mL). The size of every spot represents the number of the differential expression genes and the color of every spot represents the range of Qvalue. (B) Downregulated gene of KEGG pathway of MRSA after treated with AgNPs-K (1.25 mg/mL). (C) Upregulated gene of KEGG pathway of MRSA after treated with AgNPs-K (1.25 mg/mL).
4.5 Gene ontology (GO) enrichment analysis
GO is the abbreviation of Gene Ontology which is a major bioinformatics classification system to unify the presentation of gene properties across all species. It includes three main branches which are cellular component (CC), molecular function (MF) and biological process (BP). GO terms with p < 0.05 indicated significant enrichment (Fig. 5).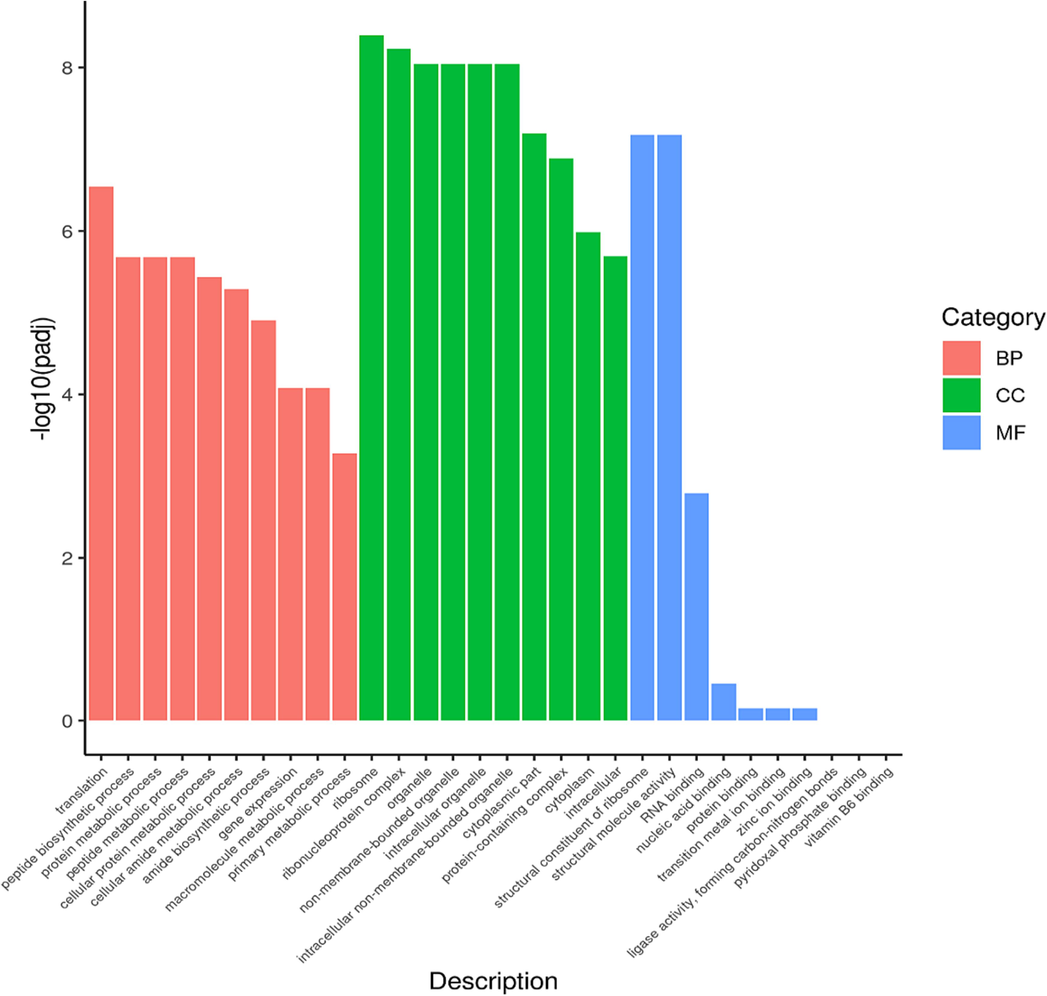
Gene ontology category of DEGs in BP, CC, and MF of MRSA after treated with AgNPs-K (1.25 mg/mL).
4.6 Novel gene prediction
To anticipate novel gene transcripts, the RNA-seq reads were assembled using Rockhopper in accordance with the reference genomes (McClure et al., 2013). By using Blastx (cutoff: evalue 1e-5), the novel transcripts were aligned to sequences in the NCBI NR database. Novel potential protein-coding transcripts were identified as novel transcripts with NR annotations (Table 4 and Table 5).
Gene_ID
Pathway_Gene_ID
Pathway_ID
Pathway_Name
Novel00006
SA0546
sau01240
Biosynthesis of cofactors
Novel00007
SA0562
sau01220
Degradation of aromatic compounds
Novel00011
SA0587
sau02010
ABC transporters
Novel00012
SA0589
sau02010
ABC transporters
Novel00013
SA0599
sau02010
ABC transporters
Novel00015
SA0660
sau02020
Two-component system
Novel00017
SA0674
sau01100
Metabolic pathways
Novel00018
SA0686
sau01232
Nucleotide metabolism
Novel00019
SA0686
sau01232
Nucleotide metabolism
Novel00022
SA0730
sau01230
Biosynthesis of amino acids
Novel00025
SA0742
sau05150
Staphylococcus aureus infection
Novel00029
SA0803
sau01100
Metabolic pathways
Gene_ID
Go_ID
Go_Ontology
Go_Term
Novel00001
GO:0016301
MF
kinase activity
Novel00001
GO:0016772
MF
transferase activity, transferring phosphorus-containing groups
Novel00001
GO:0003674
MF
molecular_function
Novel00001
GO:0003824
MF
catalytic activity
Novel00006
GO:0006464
BP
cellular protein modification process
Novel00006
GO:0008152
BP
metabolic process
Novel00006
GO:0006807
BP
nitrogen compound metabolic process
Novel00006
GO:0008150
BP
biological_process
Novel00010
GO:0016021
CC
integral component of membrane
Novel00010
GO:0005575
CC
cellular_component
Novel00010
GO:0016020
CC
membrane
Novel00010
GO:0031224
CC
intrinsic component of membrane
4.7 RT-qPCR
To validate the NGS result, RT-qPCR was performed on selected genes, which play important roles in intercellular biofilm, clumping factor A (virulent) and glycolysis pathway (Table 6). In AgNPs-K treated MRSA relative to control MRSA, the expression of icab, clfA and eno were upregulated. Those results were largely consistent with the transcriptome result that alter the biofilm, clumping factor A and glycolysis pathway.
Gene
Description
Fold Change
SD
icaB
Intercellular adhesion gene
40.42 ± 0.27
clfA
Clumping factor A
0.77 ± 0.19
eno
Glycolysis
0.27 ± 0.09
5 Discussion
5.1 Morphology of MRSA after treated with AgNPs-K under scanning electron microscope
Many researchers have utilized silver nanoparticles incorporated with plant extracts to combat MRSA or S. aureus bacteria. Table 7 shows the findings from another researcher who used silver nanoparticles incorporated with a bioactive compound against MRSA or S. aureus. However, the data on the mechanism of inhibition for silver nanoparticles incorporated with kaempferol against MRSA are still limited. Recent study demonstrated the antibacterial activities of AgNPs-K towards MRSA (Hairil Anuar et al., 2022). AgNPs-K has successfully shown bactericidal effect on MRSA with the MIC and MBC values of 1.25 mg/mL and 2.5 mg/mL, respectively and the MBC/MIC values of 2. AgNPs-K also showed bacteriostatic activity toward MRSA at 8 h with the reduction of less than 3 log10 CFU/mL (p < 0.05) compared to non-treated MRSA (Hairil Anuar et al., 2022). These properties therefore might affect the morphological changes of treated MRSA. Thus, this research was done to continue the study and observed the inhibition mechanism of MRSA treated with AgNPs-K. S. aureus (ATCC 29213) Methicillin-resistant Staphylococcus aureus (MRSA) (ATCC BAA-41) Levo–AgNPs appeared to have high antimicrobial activity against S. aureus (ATCC 29213) compared to free levofloxacin. MIC values decreased by approximately 25% against S. aureus (ATCC 29213). Levo–AgNPs appeared to have high antimicrobial activity against MRSA strains compared to free levofloxacin alone. MIC value decreased in the range of 16.7% against MRSA strains. MRSA (clinical isolates) MRSA was the most sensitive bacterial strain to D-SNPs and silver nitrate, with IZ diameters equal to 23.7 ± 0.08 and 18 ± 0.58, respectively. D-SNPs resulted in deformation of the MRSA shape, reduction in bacterial density, cell bursting, scattering of cell debris and vacuoles. S. aureus (ATCC 6538) MRSA (ATCC 33592) MRSA (ATCC 49476) The results of the kinetics of microbial growth analysis and the colony-forming capacity assay confirmed that the Ag1/80S powders have antibacterial activity against S. aureus (ATCC 6538), MRSA (ATCC 33592) and MRSA (ATCC 49476). MRSA (clinical isolates) Chitosan-Ag NPs solution showed superior antimicrobial efficacy compared to its pure forms. It reduces the minimal inhibition concentration of the substances into 2- and 4-folds (3.3 and 1.2 μg/ml), respectively. MRSA (clinical isolates) Coagulase-negative Staphylococci (CoNS) (clinical isolates) MRSA and CoNS are highly sensitive to the antimicrobial activity of AgNPs with a mean zone of inhibition (ZOI) in diameter of 19 and 17.5 mm, respectively, at 100 μg/mL concentration of AgNPs. The synthesized AgNPs have remarkable antibacterial efficacy as compared with antibiotics erythromycin and amoxicillin.
Synthesized Silver Nanoparticles
Targeted Pathogen
Antibacterial Activity
Reference
Levofloxacin-conjugated AgNps (Levo-AgNPs)
(Saadh et al., 2021)
Silver nanoparticles synthesized using Desertifilum sp. (D-SNPs)
(Hamida et al., 2020)
Ag/80S bioactive ceramic powders
(Kung et al., 2020)
Chitosan-Ag NPs solution
(Holubnycha et al., 2018)
Silver nanoparticles synthesized using Alysicarpus monilifer
(Kasithevar et al., 2017)
In this study, the morphological changes of MRSA after treated with AgNPs-K were observed under SEM to determine the effect of the treatment toward the bacteria. As mentioned before, MRSA is a Gram-positive coccus in grape-like clusters. From Fig. 1A, we have proven that the cell shape of the untreated MRSA was normal, smooth surface with a coccal shape and linked in a grape-like cluster.
The morphology was then altered after the MRSA has been treated with AgNPs-K with different concentration and time of incubation as seen in Fig. 1(B), (C) and (D). The treated MRSA undergo lysis, disrupt their cell wall, and lose their coccal appearance, releasing their cellular contents into the surrounding environment and ultimately causing cell disruption. This is possibly due to the antibacterial activities of kaempferol to inhibit the synthesis of the staphylococcal (Carruthers et al., 2020). Previous study also reported that kaempferol inhibited hemolysis at a low concentration (32 μg/mL) and exerted no effect on bacterial growth (Yin et al., 2022).
Moreover, the effect of silver nanoparticles also can be the reason behind these morphological changes. It has been reported that AgNPs cause damage to the cellular respiration process and permeability (Liao et al., 2019). AgNPs can also interact with the respiratory enzyme system which will generate reactive oxygen species (ROS) that lead to oxidative stress and this situation can cause damage to proteins and nucleic acid (Gurunathan et al., 2018). Additionally, AgNPs can also hinder cell division by binding sulfur and phosphorus present in DNA (Ahmad et al., 2020).
5.2 Transcriptomic profile analysis
Transcriptomic profile analysis was done to determine the gene regulation of MRSA treated with AgNPs-K. In this research, Next-generation sequencing (NGS) was used for transcriptome analysis profiling. NGS is a type of DNA sequencing technology that uses parallel sequencing of multiple small fragments of DNA to determine sequence. It also can expose specific biological functions affected after MRSA is treated with AgNPs-K by using differential expression analysis (DEG) and enrichment analysis which involve GO enrichment analysis and KEGG enrichment analysis. The regulation of genes was then analysed to determine whether they were down regulated or up regulated.
5.3 Down regulated gene
Downregulation is a process in which a cell reduces the production and quantity of its own components in response to external stimuli. It can either give a positive effect or negative effect towards the cell. In this research, for the KEGG pathway analysis, several pathways, including MRSA ribosome, Staphylococcus aureus infection, arginine biosynthesis, purine metabolism, and riboflavin metabolism were downregulated after being exposed with AgNPs-K.
For GO analyses, there are several genes that were also downregulated such as rplS, and rpIC, which involves in the 50S subunit proteins which function primarily to stabilize inter-domain interactions that are necessary to maintain the subunit's structural integrity. The downregulation of these genes might influence the stabilization of the inter domain and thus lead to morphological changes which can be seen by using SEM (Fig. 1B, C and D).
5.4 Upregulated gene
In molecular biology, “gene upregulation” refers to the process by which a gene is expressed at a higher level than its normal baseline expression in a particular cell or tissue under certain conditions. Interestingly, AgNPs-K significantly enriches the oxidative phosphorylation pathway of MRSA (Fig. 4 C). Oxidative phosphorylation is the process where energy is harnessed through a called the electron transport chain and ATP synthase to create ATP. However, this phenomenon might happen due to the limited energy production where they were been hindered by AgNPs-K which had altered the arginine biosynthesis, purine metabolism, and riboflavin metabolism. This limitation creates insufficient ATP production and thus MRSA needs other ways to create ATP. Therefore, this led to the up regulation of oxidative phosphorylation.
5.5 Novel gene
In this research, there are several novel genes that had been discovered such as Novel00001, Novel00006, and Novel000010. This discovery of novel genes can provide insight into the fundamental biological process of mechanism of action of AgNPs-K towards MRSA. Researchers also can obtain a wider knowledge of how cells and organisms’ function at the molecular level by discovering novel genes and their roles (Klasberg et al., 2016). In addition, it might aid in the detection and treatment of diseases caused by MRSA. Additionally, novel genes can offer crucial suggestions for the detection and treatment of disorders (Wangler et al., 2017). In this research, the novel gene provides us insight to the new gene that involves in kinase activity, transferase activity, transferring phosphorus-containing groups, molecular function, and other mechanism of action of AgNPS-K toward MRSA.
5.6 Inhibition mechanism of AgNPs-K towards MRSA
The NGS and RT-PCR analyses revealed upregulation of the icaB, eno, and clfA genes after treatment with AgNPs-K. The icaB gene is a component of the icaADBC operon, which is linked to the production of polysaccharide intercellular adhesin (PIA) in Staphylococcus species, including S. aureus. PIA constitutes a crucial element in the biofilm matrix-a slimy, protective layer of extracellular polymeric substances that bacteria, including MRSA (Abdel-Shafi et al., 2022). The observed upregulation of the icaB gene suggested a process in which the MRSA biofilm is disrupted after treatment with AgNPs-K. This study proposes that the mechanism of action (MOA) of AgNPs-K involves inhibiting the function of the biofilm pathway in MRSA.
The clfA gene in S. aureus, including MRSA encodes a surface protein known as clumping factor A (clfA). ClfA serves as a vital virulence factor, playing a crucial role in the pathogenesis of S. aureus infections by facilitating adhesion to host cells and the formation of microbial aggregates. Its involvement in the ability of S. aureus cells to clump together is attributed to the binding of clfA to fibrinogen, ultimately leading to the creation of microbial aggregates (Azizi et al., 2023).
SEM analysis demonstrated that the morphology of MRSA treated with AgNPs-K exhibited lysis and cell disruption. This finding is consistent with NGS results, indicating that the clfA gene was interrupted following treatment with AgNPs-K. Subsequent to AgNPs-K treatment, the clfA gene was upregulated, suggesting an attempt to aggregate. Understanding the roles of virulence factors like clfA is crucial for developing effective strategies to combat S. aureus infections, including those caused by MRSA. Targeting virulence factors may present alternative approaches to treating infections, especially in the face of rising antibiotic-resistant strains such as MRSA. This study implies that the mode of action (MOA) of AgNPs-K involves inhibiting the function of MRSA aggregate formation.
The eno gene found in bacteria, including MRSA encodes the enzyme enolase-a multifunctional enzyme integral to glycolysis, the cellular process through which glucose is broken down to generate energy (Krucinska et al., 2019). In the context of this study, the eno gene exhibited upregulation following treatment with AgNPs-K. This observed upregulation suggests that the glycolysis pathway was disrupted by AgNPs-K, subsequently impeding MRSA's ability to produce the necessary energy for cell division and growth.
6 Conclusion
In conclusion, AgNPs-K possesses bacterial activity and significantly inhibits the growth of MRSA through several mechanism inhibitions of MRSA. The result of transcriptomic and RT-PCR analysis suggests that AgNPs-K inhibit the expression of icab, clfa and eno genes which are responsible in clumping factor A (Virulent), biofilm and glycolysis pathway, respectively. These findings provide new insight into the mechanism of antibacterial activity of AgNPs-K towards MRSA in biofilm pathway, virulent and glycolysis activity.
Acknowledgments
This research was sponsored by Grant Scheme of FRGS from the Ministry of Higher Education in Malaysia (FRGS/1/2020/STG05/USIM/02/2).
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Transcriptomic profile analysis of Streptococcus mutans response to Acmella Paniculata flower extracts. Evidence-Based Complement. Alternat. Med. 2022
- [Google Scholar]
- The association between icaa and icab genes, antibiotic resistance and biofilm formation in clinical isolates of Staphylococci spp. Antibiotics. 2022;11(3):389.
- [Google Scholar]
- Bactericidal activity of silver nanoparticles: a mechanistic review. Mater. Sci. Energy Technol.. 2020;3:756-769.
- [Google Scholar]
- Quantitative PCR analysis of genes expressed during biofilm development of methicillin-resistant Staphylococcus aureus (MRSA) Infect. Genet. Evol.. 2013;18:106-112.
- [Google Scholar]
- The correlation between clumping factor a gene expression in biofilm formation and antibiotic resistance among Staphylococcus aureus isolated from urine samples of Imam Khomeini Hospital, Tehran. J. Adv. Biomed. Sci.. 2023
- [Google Scholar]
- The anti-MRSA compound 3-O-alpha-L-(2 ″, 3 ″-di-p-coumaroyl) rhamnoside (KCR) inhibits protein synthesis in Staphylococcus aureus. J. Proteomics. 2020;210:103539
- [Google Scholar]
- Silver nanoparticles affect the inflammatory response in a lung epithelial cell line. Open Biotechnol. J.. 2020;14(1)
- [Google Scholar]
- Antibacterial efficacy of silver nanoparticles on endometritis caused by Prevotella melaninogenica and Arcanobacterum pyogenes in dairy cattle. Int. J. Mol. Sci.. 2018;19(4):1210.
- [Google Scholar]
- Hairil Anuar, A.H., Abdul Ghafar, S.A., Hanafiah, R.M., Lim, V., 2022. Green synthesis, characterization and antibacterial activity of silver nanoparticles-kaempferol (AgNPs-K) against methicillin-resistant Staphylococcus aureus (MRSA) [Poster presentation]. In: International Applied Biology Conference, Melaka, Malaysia.
- Novel biogenic silver nanoparticle-induced reactive oxygen species inhibit the biofilm formation and virulence activities of methicillin-resistant Staphylococcus aureus (MRSA) strain. Front. Bioeng. Biotechnol.. 2020;8:433.
- [Google Scholar]
- Antibacterial activity of in situ prepared chitosan/silver nanoparticles solution against methicillin-resistant strains of Staphylococcus aureus. Nanoscale Res. Lett.. 2018;13:1-8.
- [Google Scholar]
- Synthesis, characterization and antibacterial activity of ethylene di-amine and 2-hydroxybenzadehyde Schiff base and its metal complexes. J. Chem. Environ.. 2022;1(01):5-16.
- [Google Scholar]
- Green synthesis of silver nanoparticles using Alysicarpus monilifer leaf extract and its antibacterial activity against MRSA and CoNS isolates in HIV patients. J. Interdiscip. Nanomed.. 2017;2(2):131-141.
- [Google Scholar]
- Computational identification of novel genes: current and future perspectives. Bioinf. Biol. Insights. 2016;10:BBI-S39950.
- [Google Scholar]
- Structural and functional studies of bacterial enolase, a potential target against gram-negative pathogens. Biochemistry. 2019;58(9):1188-1197.
- [Google Scholar]
- Antibacterial activity of silver nanoparticles (AgNP) confined to mesostructured, silica-based calcium phosphate against methicillin-resistant Staphylococcus aureus (MRSA) Nanomaterials. 2020;10(7):1264.
- [Google Scholar]
- Bactericidal and cytotoxic properties of silver nanoparticles. Int. J. Mol. Sci.. 2019;20(2):449.
- [Google Scholar]
- Computational analysis of bacterial RNA-Seq data. Nucleic Acids Res.. 2013;41(14):e140.
- [Google Scholar]
- Cytotoxic activity and antibiofilm efficacy of biosynthesized silver nanoparticles against methicillin-resistant Staphylococcus aureus strains colonizing cell phones. Can. J. Infect. Dis Medical Microbiol. 2022
- [Google Scholar]
- First principles study of silver argyrodites-structured compounds a8bc6 (a= ag; b= si, ge; c= te) for opto-electronic application. J. Chem. Environ. 2023:40-51.
- [Google Scholar]
- Effect of silver nanoparticles on the antibacterial activity of levofloxacin against methicillin-resistant Staphylococcus aureus. Eur. Rev. Med. Pharmacol. Sci. 2021;25(17):5507-5510.
- [Google Scholar]
- Improved antibacterial activity of water-soluble nanoformulated kaempferol and combretastatin polyphenolic compounds. Int. J. Polym. Sci. 2021
- [Google Scholar]
- Analyzing real-time PCR data by the comparative CT method. Nat. Protoc.. 2008;3(6):1101-1108.
- [Google Scholar]
- The global prevalence of Daptomycin, Tigecycline, Quinupristin/Dalfopristin, and Linezolid-resistant Staphylococcus aureus and coagulase–negative staphylococci strains: a systematic review and meta-analysis. Antimicrob. Resist. Infect. Control. 2020;9(1):1-20.
- [Google Scholar]
- Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol.. 2010;28(5):511-515.
- [Google Scholar]
- Model organisms facilitate rare disease diagnosis and therapeutic research. Genetics. 2017;207(1):9-27.
- [Google Scholar]
- Recent advances in metal decorated nanomaterials and their various biological applications: a review. Front. Chem.. 2020;8:341.
- [Google Scholar]
- Kaempferol inhibits the expression of α-hemolysin and protects mice from methicillin-resistant Staphylococcus aureus-induced lethal pneumonia. Microb. Pathog.. 2022;162:105336
- [Google Scholar]
- The antibacterial mechanism of silver nanoparticles and its application in dentistry. Int. J. Nanomed. 2020:2555-2562.
- [Google Scholar]
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105489.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1
The profile of RNA and gene regulation results can be achieved in suplementary data.







