Translate this page into:
Integration of fingerprint-activity relationship and chemometric analysis to accurately screen Q-markers for the quality control of traditional Chinese medicine compound preparation: Jinlian Qingre granules as an example
⁎Corresponding authors at: School of Pharmacy, Ningxia Medical University, Yinchuan, Ningxia 750000, China. sunlei_668@126.com (Lei Sun), zhangxia@nxmu.edu.cn (Xia Zhang)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Abstract
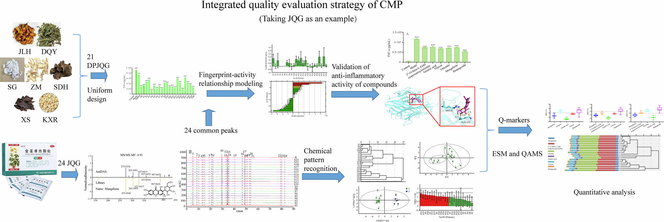
We developed a comprehensive strategy to identify and quantify quality markers of CMP by integrating multidisciplinary technology of metabolomics, chemometrics, and pharmacodynamics. We constructed a quantitative analysis approach of QAMS based on various internal reference substances. We explored the relationship between anti-inflammatory activity and the fingerprint of JQG for the first time. We clarified the main chemical compositions in JQG for the first time.
Abstract
This study aims to develop a comprehensive strategy that accurately screens and quantifies quality markers (Q-markers) by combining LC-MS metabolomics with fingerprint-effect relationships, chemometrics, and QAMS methods for the quality control in traditional Chinese medicine compound preparations (CMP). First, the chemical compositions in Jinlian Qingre granules (JQG) were analyzed and identified using ultrahigh-performance liquid chromatography-high resolution mass spectrometry (UHPLC-HRMS). Next, the HPLC fingerprint of JQG was established, and chemometrics were employed to evaluate JQG quality. Then, potential anti-inflammatory bioactive constituents were screened using a fingerprinting-effect relationship model utilizing partial least squares regression (PLSR) and a back propagation neural network optimized using a genetic algorithm (GA-BPNN). On this basis, Q-markers of JQG were determined and analyzed using quantitative analysis of multi-components by the single marker (QAMS) employing three components as reference substances (RS). 24 chemical constituents in JQG were characterized using UHPLC-HRMS. Twenty-four common peaks were assigned to the fingerprint. Six compounds, mangiferin, 2′'-O-beta-L-galactopyranosylorientin, orientin, veratric acid, vitexin, and harpagoside, identified from the common peaks associated with anti-inflammatory efficacy, could be used as Q-markers of JQG. The content determination results displayed no significant difference in the content of Q-markers in 20 batches of JQG samples measured using the external standard method and QAMS. Q-markers could be efficiently screened and determined by integrating multidisciplinary technologies. This comprehensive strategy could be beneficial for the thorough evaluation of CMP.
Keywords
Comprehensive quality evaluation
Fingerprint-activity relationship
Anti-inflammatory
Chemometrics
QAMS
Jinlian Qingre Granules
1 Introduction
Traditional Chinese medicine (TCM) has been extensively utilized in clinical practice due to its proven efficacy (Li et al., 2021, Yu et al., 2022). The multi-component, multi-targeted, and multifunctional characteristics of TCM contribute to its heritage and innovation (Han et al., 2022, Chen et al., 2023). However, the key issue restricting the development of TCM is the difficulty of quality control caused by its complex composition and unclear mechanism of action (Li et al., 2022). Especially, traditional Chinese medicine compound preparations(CMP), consisting of intricate combinations of various herbal medicines, accentuate the advantages of collaborative therapy and magnify the difficulties in ensuring quality control (Wang et al., 2023, Yao et al., 2023). Hence, integrating contemporary detection techniques and advanced analysis methods in researching CMP to regulate their quality and enhance the quality standard system poses a formidable undertaking and an inexorable trajectory.
Fingerprints have proven to be an effective method for evaluating the chemical consistency of CMP. It exhibits “integrity” and “fuzziness”, which are key characteristics that are reflected in TCM's basic theories (Zhang et al., 2023). Nevertheless, the high-performance liquid chromatography (HPLC) fingerprints may not identify unknown constituents. To overcome this challenge, ultrahigh-performance liquid chromatography-high resolution mass spectrometry (UHPLC-HRMS), a rapid and highly accurate analysis technology with strong separation capabilities, has been combined for compound identification (Xu et al., 2018, Wang et al., 2023). The fingerprint analysis can characterize the overall characteristics of the primary chemical components of CMP, whereas quantitative analysis of Q-markers can be used to perform more specific quality control (Li et al., 2019). Therefore, exploring Q-markers that can indicate the intrinsic quality of CMP is greatly crucial. Fingerprint-activity relationship modeling can characterize the correlation between chemical composition and efficacy of CMP. It can rapidly screen active compounds from complex systems and has been widely used to screen CMP's Q-markers (Chen et al., 2019).
Currently, the external standard method (ESM) and quantitative analysis of multi-components using the single marker (QAMS) are the primary approaches for simultaneously determining the multi-component content of CMP. (Zhang et al., 2022). QAMS only requires the determination of one reference substance (RS) compared to ESM. This study calculated the content of other components using relative correction factors (RCFs), providing a powerful and potentially useful solution for the lack of sufficient reference substance (Zhao et al., 2021, Wang et al., 2023). Particularly, establishing a QAMS with different substances as RS in a study can simultaneously quantify multiple components in the presence of only one of these RSs (Yang et al., 2022).
Jinlian Qingre Granules (JQG) contain seven Chinese herbal medicines: Flos Trollii (FT), Isatidis Folium (IF), Gypsum Fibrosum (GF), Anemarrhenae Rhizoma (AR), Rehmanniae Radix (RR), Scrophulariae Radix (SR), and Armeniacae Semen Amarum (ASA). JQG has the effects of producing hypothermia, detoxifying, boosting fluid production, moistening the throat, relieving cough and phlegm, and it is clinically used to treat external heat syndrome (Li et al., 2021). JQG has the characteristics of a fast antipyretic, high curative effect, and suitability for acute high fever (Zhang et al., 2012). The JQG quality is controlled only by determining the vitexin content in ChP 2020. To date, most studies on the quality standards of JQG have used one or more components as control indicators, and the correlation between the selected indicators and quality has not been confirmed (Sun et al., 2013, Li et al., 2021, Gao et al., 2021). Consequently, these methods cannot achieve an accurate and complete quality evaluation of JQG. In this study, a synthetic strategy was developed to screen and quantify Q markers accurately. UHPLC-HRMS was utilized for qualitative analysis to identify the components of JQG. HPLC fingerprints of JQG were established and combined with chemical pattern recognition to analyze quality differences in batches. Based on the efficacy of JQG, its anti-inflammatory effect was selected as pharmacodynamic index. The anti-inflammatory efficacy of JQG was studied using the RAW264.7 cell inflammatory model (Wang et al., 2023). Partial least squares regression (PLSR) and back propagation neural network optimized by genetic algorithm (GA-BPNN) were used to screen the anti-inflammatory components of JQG (Liu et al., 2021, Deng et al., 2023, Zhang et al., 2023). Based on the above research, combined with the measurability of Q-makers, the Q-makers of JQG were determined. Molecular docking technology was used to determine the binding ability of the selected compounds to inflammatory proteins, further verifying the anti-inflammatory activity of the compounds. Finally, the QAMS method was established using three components, orientin, 2″-O-beta-L-galactopyranosylorientin, and vitexin, with apparent pharmacological effects and high content in JQG as the RS, respectively, and can achieve the simultaneous determination of six Q-makers in JQG when there is any one of the RS. This study offers an empirical foundation for enhancing the quality standards of JQG and serves as a valuable resource for developing quality standards of CMP.
2 Materials and methods
2.1 Chemicals and reagents
Twenty batches of JQG were purchased from Ningxia Qiyuan Chinese Medicine Co., Ltd. (Yinchuan, China) and labeled as S1-S20; detailed information is provided in Tab. S1. FT, IF, GF, AR, RR, SR, and ASA were procured from Ningxia Mingde Healthy Pharmaceuti Co., Ltd. (Yinchuan, China). The corresponding author authenticated the samples and kept them in the hall of traditional Chinese medicine (TCM) specimens of Ningxia Medical University at room temperature.
Reference standards of harpagide (≥98 %; batch no. PS011469) and harpagoside (≥98 %; batch no. PS000404) were purchased from the Chengdu Push Bio-technology Co., Ltd. (Sichuan, China). Mangiferin (98.4 %; batch no. 111607–201704), indirubin (99.6 %; batch no. 110717–201805), indigo (98.7 %; batch no. 110716–201612), anemarrhena saponin B II (94.4 %; batch no. 111839–201505), catalpol (98.8 %; batch no. 110808–202112) and amygdalin (88.2 %; batch no. 110820–201808) were obtained from National institute for food and drug control (Beijing, China). Orientin (≥98 %; batch no. S19GB161663), 2″-O-beta-L-galactopyranosylorientin (≥98 %; batch no. P01O11F126428), veratric acid (≥98 %; batch no. DN1127BA13), and vitexin (≥98 %; batch no. D29GB172870) were purchased from the Shanghai Yuanye Bio-Technology Co., Ltd. (Shanghai, China). Chromatographic methanol and acetonitrile were bought from Thermo Fisher (Massachusetts, USA). Phosphoric acid was obtained from Kermel Chemistry Reagent Co., Ltd. (Tianjin, China). Ultrapure water was obtained from Watson (Guangzhou, China). Lipopolysaccharide (LPS) was obtained from Sigma (MO, USA).
RAW 264.7 cells were purchased from Zhongqiao Xinzhou Biological Technology Co., Ltd. (Shanghai, China) and cultured in Dulbecco’s modified Eagle’s medium (DMEM) containing penicillin, streptomycin, and fetal bovine serum in a humid atmosphere at 37 ℃ and 5 % CO2. Cell counting kit 8 (CCK8) was purchased from APExBIO Technology LLC (St. Louis, MO, USA). Mouse TNF-α and IL-6 ELISA kits were obtained from Elabscience Biotechnology Co., Ltd (Wuhan, China). Typically, 10 % fetal bovine serum, 100 units/mL penicillin, and 100 g/mL streptomycin sulfate were added to prepare the complete medium. RAW 264.7 cells were cultured in the complete medium.
2.2 Sample and standard solution preparation
2.2.1 Sample solution preparation
The sample was crushed into powder using a mortar. The sample powder (0.50 g) was accurately weighed, and 50 % (v/v) methanol aqueous solution (25 mL) was accurately measured in a 50 mL conical flask with cover ultrasound in a water bath (120 W, 40 k HZ) at room temperature for 30 min. Methanol aqueous solution (50 %) was used to compensate for reduced weight. The appropriate sample solution was placed in an EP tube and centrifuged at 10,000 rpm for 15 min. Then, the supernatant was injected into the instrument for further analysis.
2.2.2 Negative sample solution preparation
According to the prescription ratio and the process conditions of extraction, drying, and molding of JQG, negative samples lacking FT, AR, and SR were prepared. Negative sample solutions were prepared using the same method described above.
2.2.3 Standard solution preparation
Harpagide, amygdalin, mangiferin, 2″-O-beta-L-galactopyranosylorientin, orientin, veratric acid, vitexin, indirubin, indigo, anemarrhena saponin B II, catalpol, and harpagoside were accurately weighed and separately dissolved in methanol to get individual stock standard solutions. The stock standard solutions were appropriately diluted by methanol to prepare the mixed standard solution. A mixed standard solution consisting of all twelve reference standards above was prepared for UHPLC-HRMS analysis. In addition, a mixed standard solution composed of 31.8 μg/mL mangiferin, 118 μg/mL 2′′-O-beta-L-Galactopyranosylorientin, 188 μg/mL orientin, 28.6 μg/mL veratric acid, 55.6 μg/mL vitexin, and 12.0 μg/mL harpagoside was prepared for quantitative analysis. Then, it was diluted by methanol to obtain the serial working standard solutions.
2.3 UHPLC-HRMS condition
All JQG samples were mixed equally for quality control (QC). UHPLC-HRMS qualitative analysis was performed using the QC samples. The sample was analyzed using a Waters Acquity UPLC I-Class Plus UHPLC system (Waters, USA) and an Agilent Eclipse Plus C18 RRHD columns (2.1 × 100 mm, 1.8 μm) (Du et al., 2023). The mobile phase was water containing 0.1 % (V/V) formic acid (A) and acetonitrile containing 0.1 % (V/V) formic acid (B). The mobile phase velocity was 0.2 mL/min, and the injection volume was 2 μL. The gradient changes of the mobile phase were as follows: the initial composition of the mobile phase was 5 % B, which was gradually increased to 15 % B at 1 min, 20 % B at 7 min, 25 % B at 13 min, 35 % B at 19 min, 58 % B at 27 min, 75 % B at 32 min, 85 % B at 35 min, and 100 % B from 36 to 39 min.
An ABSciex TOF 5600 + mass spectrometer (Massachusetts, USA) with an electric spray ion source (ESI) in the positive ion mode was employed to obtain the MS data. The IDA scan type covered a mass range from 100 to 1000 Da. Nitrogen gas (N2) served as both the collision and auxiliary gas. The ion source pressure of GS1 and GS2 was set to 50 psi. Collision energy (CUR) was 30 psi, the temperature (TEM) was 500 °C, the ion spray voltage floating (ISVF) was 5500 V, the decluttering potential (DP) was 80 V, and the collision energy (CE) was 10 V. The MS/MS scanning mode was product-ion scanning. A collision energy of 35 eV was employed, with a collision energy range of 15 V. The ion release delay (IRD) was set to 67 ms, whereas the ion release width (IRW) was adjusted to 25 ms.
2.4 HPLC fingerprint analysis
2.4.1 HPLC conditions
HPLC analysis was conducted using a SHIMADZU HPLC system (SHIMADZU, Japan) consisting of an LC-20AT pump and an SPD-20A DAD detector. An Agilent Eclipse Plus C18 liquid chromatography column (4.6 × 250 mm, 5 µm) was utilized. The injection volume, flow rate, and column temperature were 10 µL, 1 mL/min, and 30 ℃, respectively. The mobile phases were acetonitrile (A) and a 0.1 % phosphoric acid aqueous solution(pH = 2.5) (B). The volume change of the mobile phase was as follows: 0–5 min, 97 % B; 5–15 min, 97–95 % B; 15–25 min, 95–90 % B; 25–91 min, and 90–68 % B. The detection wavelength was 270 nm in 0–18 min, 210 nm in 18–36 min, 270 nm in 36–70 min, and 210 nm at 70–91 min.
2.4.2 Methodological verification
Fingerprinting, stability, precision, and repeatability of the analysis method were examined to verify the applicability of HPLC. The precision was evaluated by conducting six replicates of the same sample solution and calculating the relative standard deviation (RSD) derived from the relative retention time (RRT), relative peak area (RPA) and relative peak height (RPH) of the common peak. Repeatability was assessed by injecting six solutions from the same batch prepared in parallel. Sample stability was determined by re-analyzing the identical sample solution at various time intervals within a 24-h period (0, 2, 4, 6, 8, 10, 12, and 24 h).
2.5 Chemical pattern recognition
Hierarchical clustering analysis (HCA) was conducted using SPSS Statistics 22.0 (Armonk, NY, USA). Import the common peaks of JQG into SIMCAP 14.1 (Umetrics, Umeå, Sweden) to perform principal component analysis (PCA) and orthogonal partial least squares discriminant analysis (OPLS-DA) to Cluster JQG samples.
2.6 Evaluation of the anti-inflammatory effects
2.6.1 Sample preparation
Different proportions of JQG (DPJQGs, labeled N0 ∼ N20) samples were prepared to ensure variations in the chemical composition. The proportion of herbs in DPJQGs was determined using a uniform design and the Data Processing System (Table S2). The ratio of N0 was consistent with that of JQG. The extraction method employed for DPJQG preparation followed the guidelines outlined in ChP 2020. This involved fire-boiling the herbs twice with water, followed by filtration. Then, the resulting twice-combined-extraction decoction was concentrated under reduced pressure and spray-dried. Negative control samples were prepared using the same method. The sample solution of each DPJQG was prepared using the same method as in section “2.2.1”, and the common peak content was analyzed using the HPLC conditions described in section “2.4.1”. These samples were also accurately weighed separately and dissolved in DMEM to obtain samples for cell experiments.
2.6.2 Cell viability assay
The CCK8 assay was used to evaluate the RAW 264.7 cell viability. RAW 264.7 cells (3 × 104 wells/100 μL) were inoculated into a 96-well plate. After 24 h, the old culture medium was removed, and different concentrations of JQG sample solution were added to continue cultivation for 24 h. After removing the old culture medium, 10 μL CCK8 and 90 μL DMEM were added and incubated at 37 ℃ for 2 h. Each well's optical density (OD) was measured at 450 nm using a microplate absorbance meter. Each drug concentration was repeated for threefold repetition, and all measurements were conducted in three independent experiments.
2.6.3 Cytokine measurement
LPS stimulates an inflammatory response in RAW 264.7 cells, and JQG suppresses the inflammatory response. Inflammatory cytokines were quantitatively analyzed using a reagent kit to evaluate the anti-inflammatory efficacy of JQG. The cells were inoculated into 96 well plates (2 × 104 wells/100 μL) and treated with DPJQG in the presence of LPS. The cells treated with DMEM complete basic medium were used as the blank control. The cells treated with LPS alone were used as the model control group. The cells treated with LPS and aspirin (ASP) were used as the positive control group. Following a 24-h incubation period, TNF-α and IL-6 levels were analyzed in the cell supernatant per the protocols provided by the manufacturer.
2.7 Establishment of fingerprint-effect relationship
2.7.1 PLSR
PLSR was established to screen anti-inflammatory active compounds, with common peak areas set as independent variables and inhibition rates of TNF-α and IL-6 as dependent variables. All modeling and analysis procedures were executed using SIMCA 14.1 software.
2.7.2 GA-BPNN
A GA-BPNN model was developed to depict the intricate and nonlinear correlation between 24 independent variables (24 common peak areas) and two dependent variables (inhibition rates of TNF-α and IL-6). Firstly, normalize the common peak areas of 21 samples and inhibition rates of TNF-α and IL-6. Then, use the common peak area as the input layer neuron, the inhibition rate as the output neuron, and select one layer from the middle layer neuron to establish a BP neural network. Through repeated experiments and optimization, the number of hidden layer neurons was ultimately determined to be 6, and the algorithm was trainlm. Calculate the MIV values of the inhibition rates of TNF-α and IL-6 for each common peak using the optimal GA-BP model, clarify the relative importance of each variable's impact on the network output, and identify potential active components.The GA-BPNN fitness was evaluated using statistical metrics, including root mean square error (MSE). All modeling and analysis procedures were performed using MATLAB 2019b.
2.8 Verification of anti-inflammatory activities of six reference compounds
The anti-inflammatory effectiveness of the active compounds was validated using cell experiments, molecular docking techniques, and a spectral effect relationship model.
2.8.1 Inhibition of TNF-α and IL-6
Six reference compounds (mangiferin, 2″-O-beta-L-galactopyranosylorientin, orientin, veratric acid, vitexin, and harpagoside) were dissolved in a 0.1 % DMSO solution and diluted with DMEM to an appropriate concentration to conduct anti-inflammatory experiments (measuring TNF-α and IL-6 release inhibition rates) and assess the rationality of determining the efficacy of compounds, following the procedure outlined in section “2.6.3”.
2.8.2 Molecular docking verification
Protein structure was downloaded from the RCSB PDB database (https://www.rcsb.org). The 3D structures of the chemical compounds were determined using PubChem (https://pubchem.ncbi.nlm.nih.gov). Finally, molecular docking was conducted using the potential targets and their respective constituents.
2.9 Quantitative analysis of the Q-marker of JQG
2.9.1 Chromatographic conditions, preparation of sample solutions, and standard solutions
The content analysis of chromatographic conditions was the same as in section ‘‘2.4.1″. Meanwhile, the sample and standard solution preparation was the same as in section ‘‘2.2.1″ and “2.2.3”.
2.9.2 Validation of the method
Method validation, including linearity, precision, repeatability, stability, and recovery rate, was performed to ascertain the suitability of the HPLC analysis method. The standard solutions were diluted with methanol at six different concentrations to establish standard curves. The linearity of the calibration curves was evaluated by plotting the peak area against the corresponding concentration of each standard. Subsequently, the regression equation, correlation coefficient, and linear range were determined based on the obtained curves. Limit of detection (LODs) and quantification (LOQs) were determined at signal-to-noise (S/N) ratios of 3:1 and 10:1, respectively. The precision was evaluated by conducting six replicates of the same sample solution and calculating the RSD derived from the peak area (PA) of the compound being measured. Both intra-day and inter-day precision were evaluated. The repeatability was assessed by calculating the RSD derived from the content of the compound measured in six parallel samples. The stability was assessed by calculating the RSD derived from the PA measured and re-analyzing the identical sample solution at various time intervals within a 24-h period (0, 2, 4, 6, 8, 10, 12, and 24 h). The recoveries were calculated using the concentration (100 %, equivalent to 1.0 times the original amounts in samples), and the concentrations were analyzed six times.
3 Results and discussion
3.1 UHPLC-HRMS
UHPLC-HRMS was applied to identify the chemical constituents of JQG using QC samples. Fig. 1A presents the total ion currents (TIC) of JQG. UHPLC-HRMS data files and extracted ion chromatograms (EIC) were analyzed using AntDAS, a software platform built by our research group. Compounds were identified by extracting EIC, resolving EIC peaks, and identifying chemical compositions. The fragment ion identification results obtained from AntDAS were used to select features corresponding to the same compound; subsequently, MS1 and MS2 spectra were constructed. Then, these spectra were compared with MS-DIAL, a publicly available compound library imported into AntDAS for compound identification (Du et al., 2023). The MS1 and MS2 of mangiferin are shown in Fig. 1B. Twenty-four reliable compounds were identified using the above methods and referring to existing literature, 12 of which were compared with retention time and the characteristic ion fragments of the corresponding reference standards. Table 1 reveals that these compounds have detailed mass data, retention times, and chemical formulas. Fig. 1C illustrates the cleavage pathways of typical compounds (Lee et al., 2023, Zhen et al., 2023). Note: “*” compounds were verified by standards.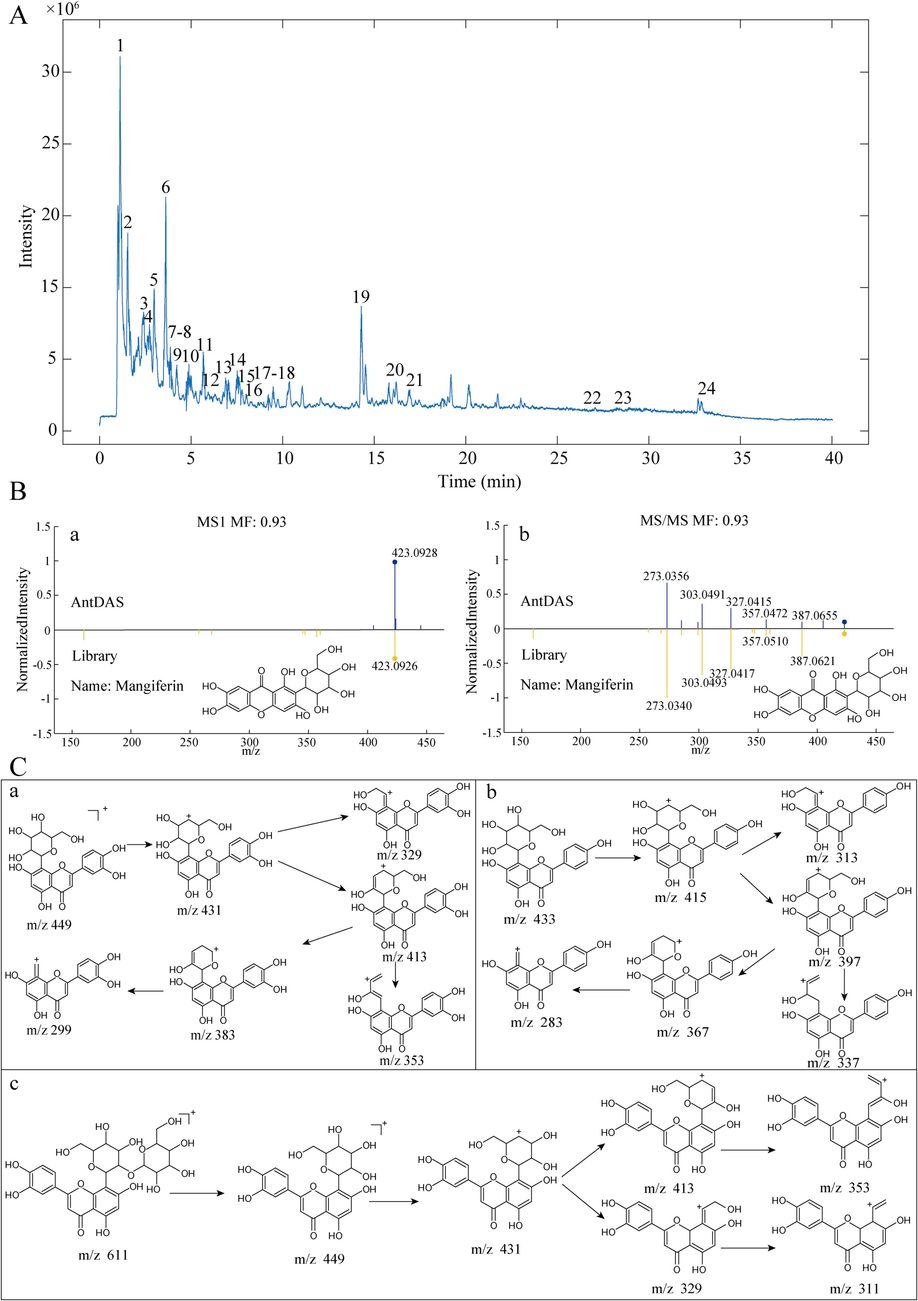
A: The TIC of JQG in a positive ion mode. B: (a) The compound identification based on the MS1 spectrum. (b) MS2 spectrum of the compound. C: The hypothesized fragmentation pathway of some typical compounds: orientin (a), vitexin (b), and 2′′-O-beta-L-galactopyranosylorientin (c).
NO.
tR/ min
Mass ([M + H+], m/ z)
Error (ppm)
Fragment Ions (m/z)
CAS. No
Formula
Identification
1
1.2836
244.0925
−1.23
244.0925, 112.0472, 95.0224
147–94-4
C9H13N3O5
Cytarabine
2
1.6882
180.1012
6.66
180.0983, 163.0749, 145.0609, 127.0519, 117.0683, 115.0528
27740–96-1
C10H13NO2
Salsolinol
3
2.3312
365.1408
1.64
365.1068, 203.0503, 185.0400
6926–8-5
C15H24O10
Harpagide*
4
3.0808
390.1755
−0.77
390.1332, 193.0843, 161.0593, 133.0630
55598–67-9
C17H24O9
Syringin
5
3.6163
458.1656
0.00
458.1656, 325,1133, 145.0485, 115.0375, 97.0268, 85.0273
29883–15-6
C20H27NO11
Amygdalin*
6
3.8686
423.0928
0.95
423.0848, 387.0655, 357.0472, 327.0415, 303.0491, 273.0365
4773–96-0
C19H18O11
Mangiferin*
7
3.98
183.0658
3.28
183.0658, 165.0530, 139.0706, 124.0483
93–07-2
C9H10O4
Veratric acid*
8
4.62
611.1601
1.15
611.1608, 449.1080, 431.0970, 413.0865, 353.0633, 329.0846, 311.0542
861691–37-4
C27H30O16
2′'-O-β-L-Galactopyranosylorientin*
9
5.0845
627.1569
2.07
627.1561, 465.1128, 303.0499
29125–80-2
C27H30O17
Quercetin-3,4′-O-di-beta-glucoside
10
5.2340
449.1082
0.00
449.1018, 431.0999, 383.0906, 353.0627, 329.0676, 299.0545
4261–42-1
C21H20O11
Isoorientin
11
5.6663
449.1076
−0.67
449.1075, 431.0992, 413.0715, 383.0906, 353.0627, 329.0698, 299.0487
28608–75-5
C21H20O11
Orientin*
12
5.9058
619.1285
2.42
619.1311, 487.0932, 317.0833
23284–18-6
C26H28O16
Peltatoside
13
6.2795
565.1567
8.32
565.1530, 433.1116, 415.0988, 337.0856, 313.0698, 283.0708
53382–71-1
C26H28O14
Isovitexin 2′'-O-arabinoside
14
6.9086
433.1136
1.62
433.1106, 397.0963, 379.0803, 337.0705, 313.0700, 283.0617, 165.0206, 121.0721
38953–85-4
C21H20O10
Isovitexin
15
7.0886
551.1041
1.63
551.1185, 303.0469
96862–01-0
C24H22O15
Quercetin 3-O-malonylglucoside
16
7.7932
465.1030
0.43
465.1265, 303.0495, 257.0476, 153.0196, 85.0280
482–35-9
C21H20O12
Isoquercitrin
17
9.7524
579.1717
1.90
579.1684, 271.0617
17306–46-6
C27H30O14
Rhoifolin
18
12.7604
433.1133
0.92
433.1133, 415.0984, 397.1017, 337.0703, 313.0691, 367.0825, 283.0556
3681–93-4
C21H20O10
Vitexin*
19
14.2787
943.4894
2.23
943.4838, 925.4894, 797.4337, 779.4197
136656–07-0
C45H76O19
Timosaponin BII*
20
15.7829
593.1875
1.69
593.1844, 447.1259, 285.0744, 242.0612, 153.0190
480–36-4
C28H32O14
Acacetin-7-O-rutinoside
21
16.0427
495.1857
−3.84
495.1473, 333.0981, 315.0826, 203.0539
19210–12-9
C24H30O11
Harpagoside*
22
25.2912
263.0820
0.38
263.0926, 235.0860, 219.0897, 217.0739, 165.0702
482–89-3
C16H10N2O2
Indigo*
23
26.4665
263.0816
−1.52
263.0811, 235.0839, 219.0946, 132.0401
479–41-4
C16H10N2O2
Indirubin*
24
33.1514
687.5230
5.08
687.2151, 524.1624, 346.1158
C27H42O20
81720–08-3
Rehmannioside D*
3.2 HPLC fingerprint of JQG
3.2.1 Optimization of HPLC conditions
The optimization of HPLC conditions, encompassing the selection of the chromatographic column, mobile phase, flow rate, detection wavelength, and column temperature, was conducted systematically to attain a desirable separation and improved analytical efficiency. Section “2.4.1” provides detailed information regarding chromatographic conditions. During the establishment of a fingerprint, the characteristic peak distributions at various wavelengths and time intervals are compared to identify distinctive peaks, leading to the selection of different detection wavelengths at different time intervals (Chen et al., 2023).
3.2.2 Methodology validation
P17 (orientin) was selected as the reference peak due to its superior peak shape and large peak area. Furthermore, the RSD of the RRT, RPA and RPH for the other 23 common peaks were calculated to assess precision, stability, and repeatability. The precision test results indicated that the RSD values for RRT, RPA, and RPH were below 1.12 %, 1.94 %, and 1.57 %, respectively. Similarly, the repeatability test results demonstrated that RRT, RPA, and RPH were less than 1.05 %, 2.98 %, and 1.38 %, respectively. The stability test results demonstrated that the RSD values for RRT, RPA, and RPH of each chromatographic peak were below 1.41 %, 2.84 %, and 2.14 %, respectively. These findings suggest that the method is accurate and reliable, making it suitable for establishing HPLC fingerprints for sample analysis.
3.2.3 Establishment of HPLC fingerprint
HPLC analysis of 20 (S1–S20) batches of JQG samples yielded the fingerprints of JQG, and HPLC data were collected. The peak matching was performed using the “TCM Chromatographic Fingerprint Similarity Evaluation System (Version 2012)” software. The reference fingerprint was the S1 sample, and the “time width” was 0.1 min. Twenty-four peaks (P1-P24) were identified as common peaks. JQG quality was assessed by evaluating similarity. Twenty-four common peaks were calibrated, and their similarity was calculated. Table 2 presents the results. The obtained similarity values between the 20 batches of JQG and the reference fingerprint exceeded 0.97, indicating that the JQG preparation exhibited uniformity, stability, and controllability. However, the RSD of RPA for each common peak ranged from 5.44 % to 29.39 %, indicating that despite the significant consistency observed among various batches of JQG, discernible differences persisted. Integrating fingerprint analysis with chemometric techniques is imperative for identifying the compounds responsible for these variations.
No.
Similarity
No.
Similarity
S1
0.998
S11
0.997
S2
0.996
S12
0.995
S3
0.997
S13
0.999
S4
0.995
S14
0.988
S5
0.998
S15
0.994
S6
0.998
S16
0.997
S7
0.998
S17
0.995
S8
0.999
S28
0.997
S9
0.999
S19
0.995
S10
0.999
S20
0.995
Based on the RT of the peak and the photodiode array detector (PDA) spectrogram, 6 of the 11 compounds identified by reference substance comparison in UHPLC-HRMS analysis, could be detected under optimized HPLC chromatographic conditions, namely P15 mangiferin from the minister drug AR, P16 2′'-O-beta-L-galactopyranosylorientin, P17 orientin, P19 veratridic acid, P20 vitexin from monarch drug FT, and P23 harpagoside from the adjuvant drug SR. Fig. 2A displays the HPLC chromatograms of references, JQG, and negative samples. Fig. 2B indicates the fingerprints of 20 batches of JQG samples and their reference fingerprints. Fig. 2C depicts the UV spectrogram of the six references and the corresponding peaks in the sample.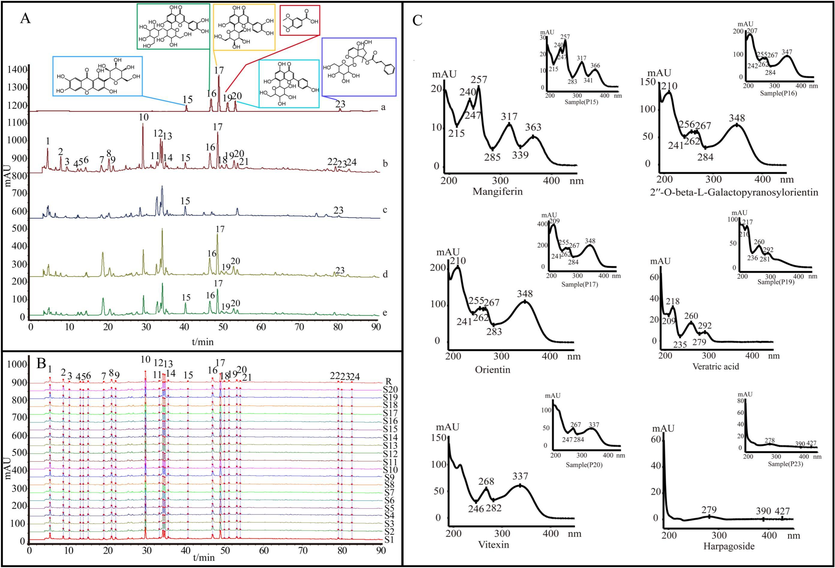
A: Chromatograms: references (a), JQG (b), the negative samples that lack FT (c), the negative samples that lack AR (d), and the negative samples that lack SR (e). B: The fingerprints of 20 batches of JQG samples. C: Comparison of the spectrogram of six compounds. 15. mangiferin, 16. 2′′-O-beta-L-galactopyranosylorientin, 17. orientin, 19. veratric acid, 20. vitexin, 23. harpagoside.
3.3 Chemical pattern recognition
3.3.1 HCA
Chromatographic common peaks were investigated using HCA to classify samples of JQG (Shi et al., 2023). Fig. 3A indicates that the 20 batches of JQG can be classified into different categories. At an Euclidean distance of 25, various batches of JQG can be classified into two categories: S1–S13 and S14–S20. Table S1 displays that each product category was prepared from a different batch number of extractum. The classification results were related to raw material production. The presence of different categories may lead to variations in quality due to disparities in raw materials. The results also suggest that products of similar quality can be grouped based on 24 common peaks.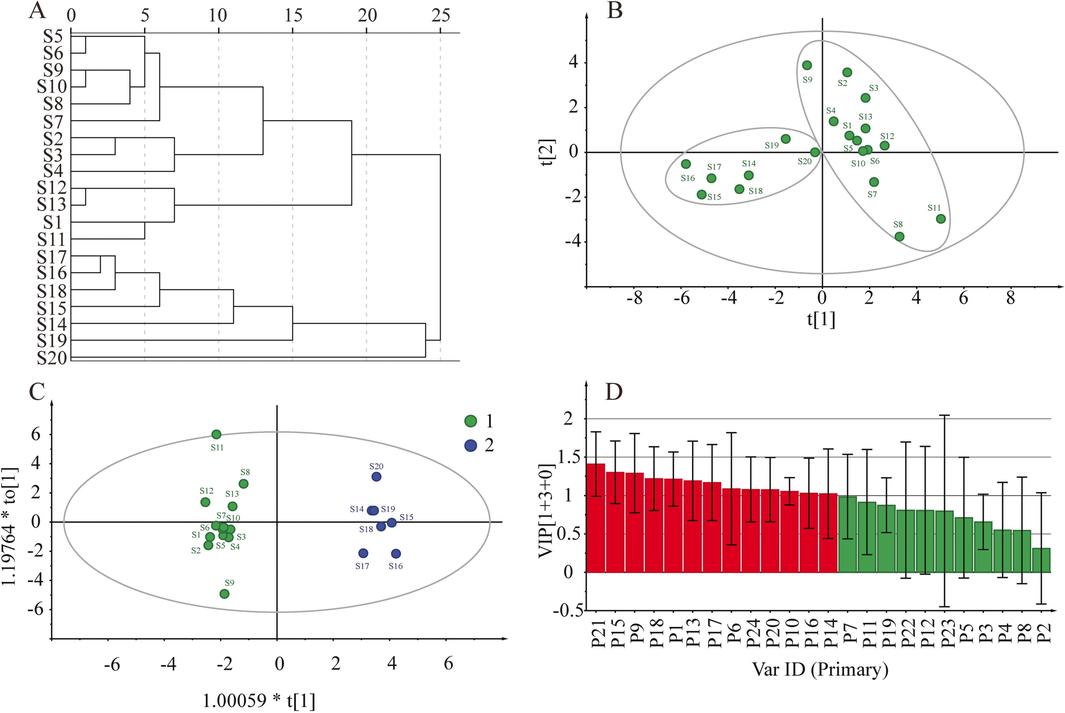
HCA results (A). PCA score scatters plot (B). OPLS-DA score scatter plot (C). The OPLS-DA VIP results (D).
3.3.2 PCA
PCA is currently the prevailing technique for unsupervised classification visualization. PCA effectively represents the data characteristics of the original variables while minimizing information loss by employing dimensionally reduced variables (Fan et al., 2022). For PCA model construction, 24 common peak areas from 20 batches of JQG samples were imported into SIMCA 14.1 software, and the score scatters plot of PCA was acquired. To predict discrimination in the model, the parameters R2 and Q2 were calculated. The result indicated the R2 X and Q2 were 0.828 and 0.513, respectively, indicating that the model had good prediction ability. Fig. 3B depicts establishing a two-dimensional PCA model in which all data points are contained within 95 % confidence intervals, and 20 batches of JQG can be separated into two distinct categories consistent with HCA. This suggests the presence of quality disparities among different batches of JQG.
3.3.3 OPLS-DA
OPLS-DA can promote sample separation and explore group differences while reducing sample dispersion within categories (Wang et al., 2023). Fig. 3C indicates that 20 batches of JQG are divided into two groups, and the results are consistent with those of PCA. The results indicate that JQG varies between different batches. The model underwent 200 substitution tests to assess its performance, resulting in interpretation rate parameters for the model (R2X and R2Y) and prediction parameters (Q2). The parameters were 0.569, 0.937, and 0.729, respectively, proving that the model was reliable. Furthermore, the VIP plot (Fig. 3D) effectively demonstrates the significant contribution of nine specific chemical constituents (P21, P15, P9, P18, P1, P13, P17, P6, P24, P20, P10, P16, and P14) to the categorization process. These are the key chemical ingredients responsible for the differences in JQG quality.
According to the above analysis, unsupervised and supervised pattern recognition approaches based on 24 common peaks could effectively distinguish JQG samples with quality differences, and 24 common peaks are of great significance for evaluating JQG quality.
3.4 Q-marker discovery based on the fingerprint-effect relationship
Q-markers should be associated with the biological activity of CMP. Anti-inflammatory is one of the primary pharmacological effects of JQG. However, the material basis of its anti-inflammatory activity remains unclear. The anti-inflammatory efficacy of DPJQG was evaluated via cell experiments. Furthermore, fingerprints-activity relationship modeling was constructed to screen for potentially effective compounds.
3.4.1 Research on anti-inflammatory effects
3.4.1.1 Cell viability assay
DPJQG concentration was determined with N0 and LPS concentrations using the CCK8 experiment. The results indicate that N0 had no cytotoxic effect on RAW264.7 cells at concentrations of 62.5, 125, 250, and 500 μg/mL, but 1000 μg/mL significantly impacts cell viability. Meanwhile, when LPS concentrations of 0.1 and 1 μg/mL had no significant inhibitory, the 10 μg/mL concentration had a significant inhibitory effect. Therefore, 500 μg/mL DPJQG and 1 μg/mL LPS were chosen as the optimal therapeutic and induction concentrations, respectively. The cell viability of all batches treated with 500 μg/mL DPJQG and 1 μg/mL LPS added to cells (Fig. S2).
3.4.1.2 Inhibition of inflammatory cytokines
The anti-inflammatory efficacy of DPJQG was assessed by quantifying TNF-α and IL-6 in cell supernatants. Fig. 4A and 5B illustrate that DPJQG can inhibit inflammatory cytokine production. The inflammatory cytokine content was significantly increased in the model group than in the control group (p < 0.01), whereas inflammatory cytokines were significantly suppressed in the aspirin group than in the model group (p < 0.01). Furthermore, different DPJQG samples have different inhibitory efficacies, indicating that their ratio composition influences the anti-inflammatory activity of DPJQGs. The dissimilarities in the chemical compositions of DPJQGs may account for the variations in their anti-inflammatory properties.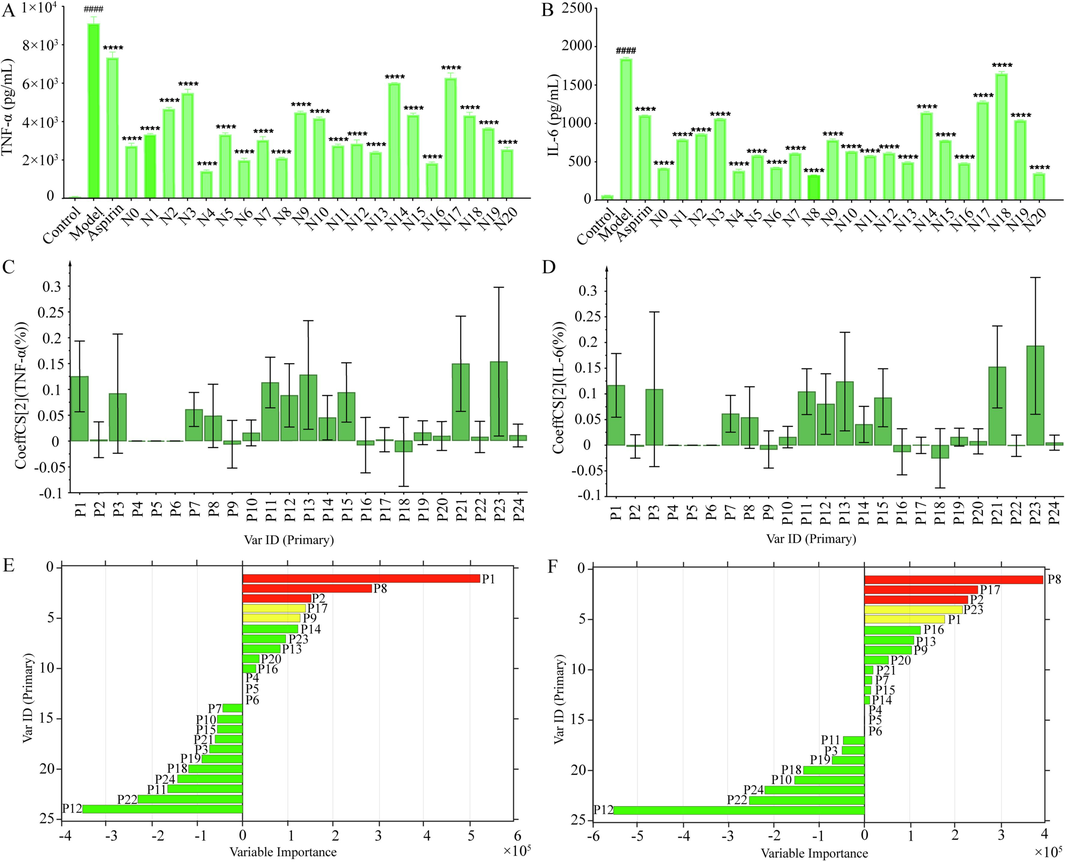
Effect of DPJQG samples on TNF-α levels (A) and IL-6 (B) in the supernatant of LPS-induced RAW 264.7 cells inflammation model (n = 3. data are expressed as the mean ± SEM. ####P < 0.001 contrast with the control group, ****P < 0.001 contrast with the model group). Regression coefficients of 24 common peaks of JQG in the PLS models of TNF-α (C) and IL-6 (D). Variable Importance of 24 common peaks of JQG in the GA-BPNN models of TNF-α (E) and IL-6 (F).
3.4.2 Determination of anti-inflammatory active ingredients in JQG
PLSR and GA-BPNN were utilized to analyze the association between common peaks and anti-inflammatory activity in DPJQG and screen anti-inflammatory active compounds.
3.4.2.1 PLSR
PLSR was utilized to build an association between common peaks and biological activity and identify the potential anti-inflammatory chemical composition in JQG (Zhang et al., 2023). The stronger the inhibitory effect on the release of inflammatory cytokines, the stronger the anti-inflammatory activity. PLSR was established using the common peak areas as the independent variable and inhibition rate as the dependent variable (Y). A positive regression coefficient in PLSR indicates a positive correlation between the two variables. Fig. 4C reveals that P1, P2, P3, P7, P8, P10, P11, P12, P13, P14, P15, P17, P19, P20, P21, P22, P23, and P24 were positively correlated with TNF-α (the values of R2 X, R2 Y and Q2 were 0.891, 0.941, and 0.902). P1, P3, P7, P8, P10, P11, P12, P13, P14, P15, P16, P17, P18, P19, P20, P21, P22, P23, and P24 had positive relations with IL-6 (the values of R2 X, R2 Y and Q2 were 0.888, 0.927, and 0.896, Fig. 4D). This indicated that P1, P3, P7, P8, P10, P11, P12, P13, P14, P15, P17, P19, P20, P21, P22, P23, and P24 contributed to the primary anti-inflammatory activity of JQG. These results indicate that the anti-inflammatory effects of JQG result from the comprehensive effect of many components.
3.4.2.2 GA-BPNN
According to the data in Table S3, all error parameters of GA-BPNN model are lower than those of the BPNN, indicating the GA-BPNN model's fitting and generalization capabilities. The MIV algorithm was used to screen the independent variables that significantly impacted drug efficacy. A positive MIV value indicates a positive correlation between two variables. The larger the MIV value, the stronger the relevancy (Yang et al., 2021). Fig. 4 indicates the MIV of TNF-α and IL-6 for 24 common peaks. Fig. 4E displays that P1, P2, P8, P9, P13, P14, P16, P17, P20, and P23 were positively correlated with TNF-α. P1, P2, P7, P8, P9, P13, P14, P15, P16, P17, P20, P21, and P23 were positively correlated with IL-6 (Fig. 4F). This indicated that P1, P2, P8, P9, P13, P14, P16, P17, P20, and P23 contributed to the main anti-inflammatory activity of JQG.
3.5 Validation of anti-inflammatory activity of compounds
Six anti-inflammatory compounds identified by HPLC analysis (mangiferin, orientin, 2″-O-beta-L-galactopyranosylorientin, veratridic acid, vitexin, and harpagoside) were validated using cell anti-inflammatory and molecular docking assays to verify the analytical results of the fingerprint-activity relationship (Deng et al., 2023).
3.5.1 Inhibition of inflammatory cytokines
The cytotoxicity of six compounds on RAW 264.7 cells was evaluated to mitigate the influence of diminished activity on inflammatory factor production. Fig. S3 displays the observed cytotoxicity. TNF-α and IL-6 were quantitatively measured to evaluate the anti-inflammatory properties of each compound using the maximum non-toxic concentration of each compound as the therapeutic concentration. Fig. S4 indicates that each compound significantly inhibits inflammatory cytokine production. Therefore, six selected chemical components exhibited good anti-inflammatory activity.
3.5.2 Molecular docking studies
The docking analysis involved the examination of Q-markers with TNF-α (PDB ID: 1A8M) and IL-6 (PDB ID: 1ALU) using data from the PDB database, providing comprehensive information on proteins and their 3D structures (Li et al., 2023). PyRx software was used to minimize the energy of the isolated and identified compound molecular structure. When combined with AutoDock Tools software, the predicted target proteins are dehydrated, hydrogenated, added with atomic charges, set with atomic types, and stored as a backup for receptors. Then, AutoDock Vina in PyRx software was used to dock the identified compounds with receptor proteins, and the results are shown in Fig. 5A. Fig. 5A indicates that the absolute values of the docking scores are all greater than 5. The active ingredients obtained by screening have a strong binding capacity with anti-inflammatory-related proteins, thereby proving the reliability of the selected quality markers. Fig. 5B and 5C exhibit the binding diagrams of orientin with TNF-α and IL-6. This was consistent with previous report that mangiferin(Kumar et al., 2023), orientin, vitexin(Yu et al., 2022), veratric acid, 2′'-O-beta-L-galactopyranosylorientin(Wang et al., 2015; Liu et al., 2018) and amygdalin(Zeng et al., 2023) independently exhibited stronger anti-inflammatory effects and further confirmed the reliablility of these compounds as the main material basis for anti-inflammatory activity of JQG.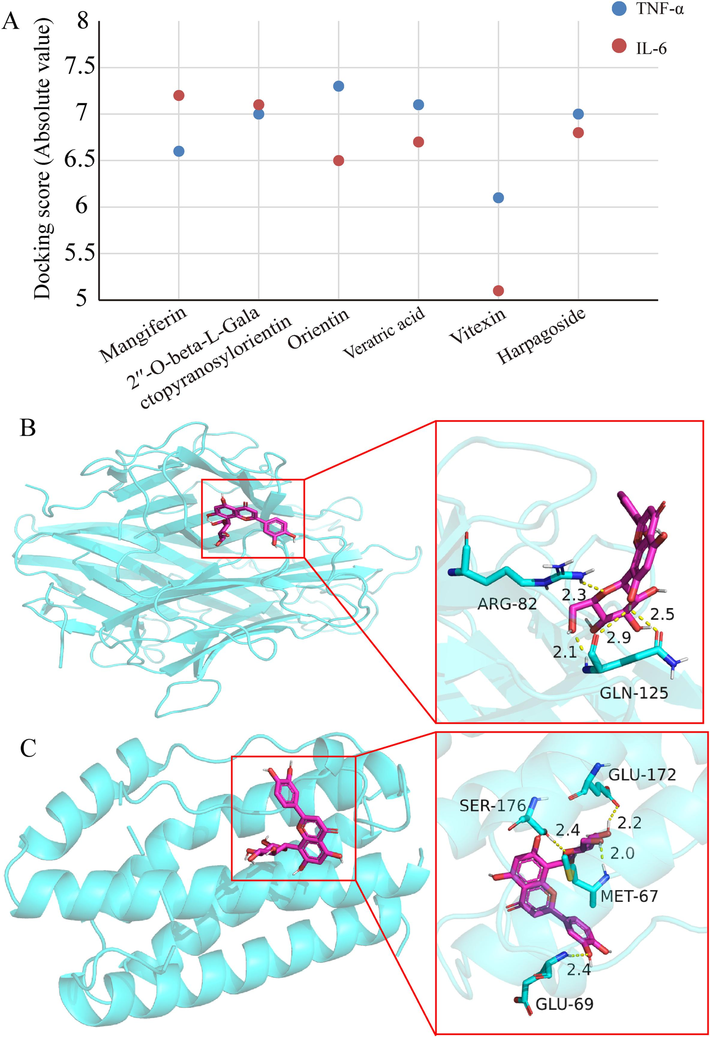
Docking scores of six compounds with TNF-α and IL-6 (A). Molecular docking analysis of orientin with TNF-α (B) and IL-6 (C).
3.6 Quantitative analysis of Q-marker in JQG
P15 (mangiferin), P16 (2′'-O-beta-L-galactopyranosylorientin), P17 (orientin), P19 (veratric acid), P20 (vitexin), and P23 (harpagoside) were identified as key components responsible for JQG quality variations using chemical pattern recognition and fingerprints-activity relationship modeling. Six Q-markers were quantitatively analyzed using QAMS (Wang et al., 2023).
3.6.1 Methodology validation of quantitative analysis
Validation tests were conducted using JQG (sample S1) on HPLC. Table S4 depicts the specific results of precision, repeatability, and stability. The range of spike-recoveries was 95.69 %∼101.42 %, and Table S5 presents specific information. Table 3 illustrates the regression equations. The LOD and LOQ values are 0.063 to 0.125 and 0.250 to 0.500 μg/mL, respectively. The r values of the established linear equations were greater than 0.9998 within the corresponding concentration range. These results indicate that this method is highly accurate and can meet the requirements for content determination.
Compounds
Regression equation
Linearity range (μg/mL)
r
LOD (μg/mL)
LOQ (μg/mL)
Mangiferin
y = 17273x + 1484.3
0.995 ∼ 31.8
0.9998
0.1250
0.500
2′′-O-beta-L-Galactopyranosylorientin
y = 14528x + 8325.3
3.70 ∼ 118
0.9998
0.125
0.500
Orientin
y = 20878x + 16042
5.88 ∼ 188
0.9998
0.063
0.250
Veratric acid
y = 44973x + 4764.9
0.895 ∼ 28.6
0.9999
0.063
0.250
Vitexin
y = 21430x + 3953.9
1.74 ∼ 55.6
0.9998
0.125
0.500
Harpagoside
y = 27712x + 1930.4
0.375 ∼ 12.0
0.9999
0.125
0.500
3.6.2 Calculation and robustness of RCF
Determining and calculating the RCF are key issues that must be addressed to promote and apply QAMS. RCF is a relatively fixed constant that must be established by determining all reference solutions for the testing components. However, redundant research in the field of QAMS is present, where identical research objects, components to be measured, and analysis methods are employed to repeatedly establish RCFs with different RS in different studies. This practice leads to the squandering of resources and goes against the original purpose of establishing the QAMS. Establishing a QAMS with different substances, such as RS, can quantify multiple components in the presence of only one type of RS.
RCF can be determined using a linear model, that incorporates six distinct concentrations of standard substances and their corresponding peak areas. RCF values were calculated employing one of the orientin, 2′'-O-beta-L-galactopyranosylorientin, and vitexin as the RS. Table S6 displays the calculation results. Subsequently, the established RCF was employed to calculate the content of the other substances to be tested (ST). ESM and QAMS results were compared by calculating the standard method difference (SMD) (Wu et al., 2021). The calculation formulas for RCF, content, and SMD are as follows:
where fRS/ST refers to the RCF value of the analyte, ARS and CRS are the peak area and concentration of the RS (one of the 2′'-O-beta-L-galactopyranosylorientin, orientin, and vitexin), respectively; AST and CST are the peak area and concentration of the ST, respectively; CESM and CQAMS represent the substance concentrations to be measured using ESM and QAMS methods, respectively.
3.6.3 Ruggedness test
The robustness of the RCF was investigated by changing the column temperature and flow rate, as well as using different apparatus and chromatographic columns. Tables S7–9 indicate that the established RCF has good robustness within the linear range.
3.6.4 Quantitative analysis of Q-marker in JQG
Table 4 indicates the specific contents of the six Q-markers in 20 batches of JQG, calculated using ESM and QAMS. Fig. 6A compares the two method results and calculates the SMD values, indicating that the difference between ESM and QAMS was not statistically significant. This indicates that the RCF value can be used to quantify the compounds in JQG. Note: A represents using 2′′-O-beta-L-galactopyranosylorientin as RS, B represents using orientin as RS, and C represents using vitexin as RS.
No.
Mangiferin
2′′-O-beta-L-Galactopyranosylorientin
Orientin
ESM
QAMS
ESM
QAMS
ESM
QAMS
A
B
C
AVE
RSD
B
C
AVE
RSD
A
C
AVE
RSD
S1
0.281
0.283
0.284
0.283
0.283
0.20 %
2.03
2.03
2.02
2.03
0.35 %
2.28
2.27
2.26
2.27
0.31 %
S2
0.278
0.28
0.28
0.279
0.280
0.21 %
2.05
2.06
2.05
2.06
0.34 %
2.18
2.17
2.17
2.17
0.00 %
S3
0.279
0.281
0.282
0.281
0.281
0.21 %
2.07
2.08
2.07
2.08
0.34 %
2.21
2.20
2.20
2.20
0.00 %
S4
0.266
0.268
0.269
0.268
0.268
0.22 %
2.00
2.01
2.00
2.01
0.35 %
2.31
2.30
2.29
2.30
0.31 %
S5
0.282
0.284
0.286
0.284
0.285
0.41 %
2.02
2.03
2.02
2.03
0.35 %
2.34
2.33
2.33
2.33
0.00 %
S6
0.278
0.28
0.281
0.28
0.280
0.21 %
2.02
2.03
2.02
2.03
0.35 %
2.29
2.28
2.28
2.28
0.00 %
S7
0.293
0.295
0.296
0.294
0.295
0.34 %
2.05
2.06
2.05
2.06
0.34 %
2.10
2.09
2.09
2.09
0.00 %
S8
0.305
0.307
0.308
0.306
0.307
0.33 %
1.90
1.91
1.90
1.91
0.37 %
2.14
2.14
2.13
2.14
0.33 %
S9
0.325
0.327
0.328
0.327
0.327
0.18 %
2.08
2.09
2.08
2.09
0.34 %
2.39
2.38
2.38
2.38
0.00 %
S10
0.300
0.302
0.303
0.301
0.302
0.33 %
1.99
2.00
1.99
2.00
0.35 %
2.33
2.32
2.32
2.32
0.00 %
S11
0.232
0.234
0.235
0.234
0.234
0.25 %
1.77
1.77
1.76
1.77
0.40 %
2.12
2.11
2.11
2.11
0.00 %
S12
0.257
0.259
0.26
0.259
0.259
0.22 %
1.92
1.93
1.92
1.93
0.37 %
2.27
2.26
2.26
2.26
0.00 %
S13
0.254
0.256
0.257
0.256
0.256
0.23 %
1.97
1.98
1.97
1.98
0.36 %
2.28
2.27
2.26
2.27
0.31 %
S14
0.315
0.316
0.318
0.316
0.317
0.36 %
2.03
2.04
2.03
2.04
0.35 %
2.47
2.45
2.45
2.45
0.00 %
S15
0.365
0.367
0.369
0.366
0.367
0.42 %
2.05
2.07
2.05
2.06
0.69 %
2.58
2.56
2.56
2.56
0.00 %
S16
0.365
0.366
0.368
0.366
0.367
0.31 %
2.24
2.25
2.24
2.25
0.31 %
2.51
2.50
2.50
2.50
0.00 %
S17
0.357
0.359
0.36
0.358
0.359
0.28 %
2.15
2.16
2.14
2.15
0.66 %
2.43
2.42
2.42
2.42
0.00 %
S18
0.354
0.356
0.357
0.355
0.356
0.28 %
2.06
2.07
2.06
2.07
0.34 %
2.32
2.31
2.31
2.31
0.00 %
S19
0.346
0.347
0.349
0.347
0.348
0.33 %
2.09
2.10
2.09
2.10
0.34 %
2.39
2.38
2.38
2.38
0.00 %
S20
0.304
0.306
0.307
0.305
0.306
0.33 %
2.02
2.03
2.02
2.03
0.35 %
2.31
2.30
2.30
2.30
0.00 %
No.
Veratric acid
Vitexin
Harpagoside
ESM
QAMS
ESM
QAMS
ESM
QAMS
A
B
C
AVE
RSD
A
B
AVE
RSD
A
B
C
AVE
RSD
S1
0.210
0.213
0.214
0.212
0.213
0.47 %
0.643
0.644
0.646
0.645
0.22 %
0.115
0.115
0.115
0.115
0.115
0.00 %
S2
0.212
0.215
0.216
0.215
0.215
0.27 %
0.629
0.630
0.632
0.631
0.22 %
0.124
0.123
0.124
0.123
0.123
0.47 %
S3
0.206
0.209
0.210
0.208
0.209
0.48 %
0.638
0.639
0.641
0.640
0.22 %
0.118
0.118
0.118
0.118
0.118
0.00 %
S4
0.212
0.215
0.216
0.214
0.215
0.47 %
0.648
0.649
0.652
0.651
0.33 %
0.123
0.123
0.123
0.123
0.123
0.00 %
S5
0.202
0.205
0.206
0.205
0.205
0.28 %
0.643
0.644
0.647
0.646
0.33 %
0.125
0.124
0.124
0.124
0.124
0.00 %
S6
0.200
0.202
0.203
0.202
0.202
0.29 %
0.633
0.634
0.637
0.636
0.33 %
0.123
0.123
0.123
0.122
0.123
0.47 %
S7
0.202
0.205
0.205
0.204
0.205
0.28 %
0.631
0.632
0.634
0.633
0.22 %
0.145
0.144
0.144
0.144
0.144
0.00 %
S8
0.195
0.197
0.198
0.197
0.197
0.29 %
0.614
0.615
0.617
0.616
0.23 %
0.110
0.110
0.110
0.110
0.110
0.00 %
S9
0.207
0.210
0.211
0.209
0.210
0.48 %
0.675
0.675
0.678
0.677
0.31 %
0.123
0.123
0.123
0.123
0.123
0.00 %
S10
0.203
0.205
0.206
0.205
0.205
0.28 %
0.632
0.633
0.636
0.635
0.33 %
0.113
0.113
0.113
0.112
0.113
0.51 %
S11
0.194
0.197
0.198
0.197
0.197
0.29 %
0.597
0.598
0.600
0.599
0.24 %
0.106
0.106
0.107
0.106
0.106
0.54 %
S12
0.212
0.215
0.216
0.214
0.215
0.47 %
0.633
0.634
0.637
0.636
0.33 %
0.103
0.103
0.104
0.103
0.103
0.56 %
S13
0.214
0.216
0.217
0.216
0.216
0.27 %
0.635
0.636
0.639
0.638
0.33 %
0.104
0.104
0.105
0.104
0.104
0.56 %
S14
0.210
0.213
0.214
0.213
0.213
0.27 %
0.662
0.663
0.667
0.665
0.43 %
0.109
0.109
0.109
0.109
0.109
0.00 %
S15
0.212
0.215
0.216
0.214
0.215
0.47 %
0.647
0.647
0.651
0.649
0.44 %
0.117
0.117
0.117
0.116
0.117
0.49 %
S16
0.222
0.225
0.226
0.224
0.225
0.44 %
0.699
0.699
0.703
0.701
0.40 %
0.108
0.108
0.108
0.108
0.108
0.00 %
S17
0.210
0.212
0.214
0.212
0.213
0.54 %
0.669
0.670
0.673
0.672
0.32 %
0.105
0.105
0.106
0.105
0.105
0.55 %
S18
0.205
0.208
0.209
0.207
0.208
0.48 %
0.646
0.647
0.649
0.648
0.22 %
0.081
0.082
0.082
0.082
0.082
0.00 %
S19
0.211
0.213
0.214
0.213
0.213
0.27 %
0.664
0.665
0.668
0.667
0.32 %
0.122
0.122
0.122
0.121
0.122
0.47 %
S20
0.211
0.214
0.215
0.214
0.214
0.27 %
0.647
0.648
0.651
0.650
0.33 %
0.112
0.112
0.112
0.112
0.112
0.00 %
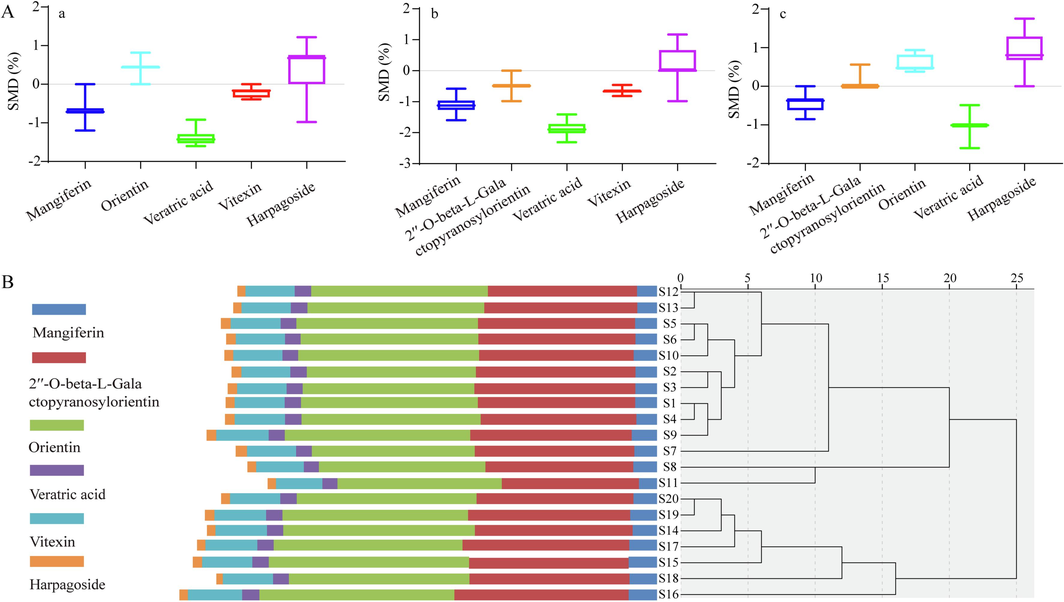
A: SMD between QAMS using 2′′-O-beta-L-Galactopyranosylorientin as RS and ESM (a). SMD between QAMS using orientin as RS and ESM (b). SMD of QAMS using vitexin as RS and ESM (c). B: Cluster analysis of six Q-marker contents in JQG.
The comparison was performed that the 4 batches sample results observed in this study with standard method in ChP 2020. The content of vitexin in S16-S19 JQG samples using the method in ChP 2020 was 0.685 mg/g, 0.656 mg/g, 0.626 mg/g, 0.648 mg/g, respectively. The difference was not statistically significant.
3.6.4 HCA based on the contents of six Q-markers
The HCA of 20 batches of JQG was conducted based on the contents of six Q-markers using SPSS Statistics 22.0 (Armonk, NY, USA). Fig. 6B indicates that the clustering results are the same as those in Fig. 3A. These results also suggest that products of similar quality can be grouped. Consequently, mangiferin, 2′'-O-beta-L-galactopyranosylorientin, orientin, veratric acid, vitexin, and harpagoside were identified. It can be used to distinguish between different JQG quality levels and assess JQG quality.
4 Conclusion
This study developed a comprehensive strategy to accurately screen and quantify quality markers (Q-markers) by combining LC-MS metabolomics with fingerprinting-effect relationships, chemometrics, and QAMS methods for quality control of CMP. Initially, 24 compounds constituting the chemical components of JQG were identified by UHPLC-HRMS. Then, the HPLC fingerprints of 20 batches of JQG were established, 24 common peaks were extracted, and the similarity was above 0.97, indicating that the JQG quality was relatively stable. The HCA, PCA, and OPLS analysis results based on a common peak can cluster the samples according to different raw materials, indicating that raw materials are the major factor causing quality differences. Twenty-four common peaks assigned are of great significance for evaluating JQG quality. Subsequently, the anti-inflammatory efficiency of DPJQG was explored in cell experiments. The fingerprint-effect relationship of JQG was studied for the first time using PLS and GA-BPNN analysis by combining fingerprints and pharmacodynamics. These results presented that the anti-inflammatory activity of JQG may be a multi-component interaction. Six identified compounds, mangiferin, 2′'-O-beta-L-galactopyranosylorientin, orientin, veratric acid, vitexin, and harpagoside, were selected as Q-markers of JQG based on the common peaks associated with anti-inflammatory efficacy. The anti-inflammatory activities of these markers were confirmed by cell experiments and macromolecular docking. Additionally, a QAMS was established to quantify the Q-markers, enabling a thorough evaluation of JQG quality. This study can provide a reference for JQG material basic research, quality standard improvement, and clinical applications. This comprehensive strategy could be beneficial for the thorough evaluation of CMP. However, some chromatographic peaks with anti-inflammatory effects were not identified in the fingerprints established in this study. The chemical composition of JQG should be studied further in the future.
Funding sources
This work was supported by National Natural Science Foundation of China (Nos. 81960711, 22378214), and Natural Science Foundation of Ningxia (No. 2023AAC03185).
CRediT authorship contribution statement
Min He: Methodology, Writing – original draft. Shan Mao: Writing – review & editing. Qingyu Du: Methodology. Xin Gao: Supervision, Validation. Jie Shi: Data curation. Xin Zhou: Writing – review & editing. Fang Zhang: Writing – review & editing. Youyuan Lu: Writing – review & editing. Hanqing Wang: Writing – review & editing. Yongjie Yu: Writing – review & editing, Methodology, Funding acquisition. Lei Sun: Writing – review & editing, Methodology, Funding acquisition. Xia Zhang: Writing – review & editing, Funding acquisition.
Acknowledgments
We thank Home for Researchers editorial team (www.home-for-researchers.com) for language editing service.
Declaration of Competing Interest
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Xia Zhang reports financial support was provided by the National Natural Science Foundation of China (Grant No. 81960711). Lei Sun reports financial support was provided by the Key Research and Development Project of Ningxia province (No. 2023AAC03185). Yong-Jie Yu reports financial support was provided by the National Natural Science Foundation of China (Grant No. 22378214).
References
- A strategy for quality evaluation of salt-treated Apocyni Veneti Folium and discovery of efficacy-associated markers by fingerprint-activity relationship modeling. Sci. Rep.. 2019;9(1):16666.
- [CrossRef] [Google Scholar]
- Quality control of Hugan capsule based on four-wavelength fusion profiling and electrochemical fingerprint combined with antioxidant activity and chemometric analysis. Anal Chim Acta.. 2023;1251:341015
- [CrossRef] [Google Scholar]
- A novel strategy for active compound efficacy status identification in multi-tropism Chinese herbal medicine (Scutellaria baicalensis Georgi) based on multi-indexes spectrum-effect gray correlation analysis. J Ethnopharmacol.. 2023;300:115677
- [CrossRef] [Google Scholar]
- Geographical discrimination of Flos Trollii by GC-MS and UHPLC-HRMS-based untargeted metabolomics combined with chemometrics. J Pharm Biomed Anal.. 2023;234:115550
- [CrossRef] [Google Scholar]
- Quality assessment of fried licorice based on fingerprints and chemometrics. Food Chem.. 2022;378:132121
- [CrossRef] [Google Scholar]
- Determination of vitexin and orientin in Jinlian qingre granules by HPLC. Ningxia Medical Journal.. 2021;43(12):1130-1132.
- [CrossRef] [Google Scholar]
- Qualitative and quantitative evaluation of Flos Puerariae by using chemical fingerprint in combination with chemometrics method. J Pharm Anal.. 2022;12(13):489-499.
- [CrossRef] [Google Scholar]
- Mangiferin from Enicostemma littorale Blume with in silico and in vitro anti-inflammatory potential. J Biomol Struct Dyn.. 2023;5:1-10.
- [CrossRef] [Google Scholar]
- Elucidation of phenolic metabolites in wheat seedlings (Triticum aestivum L.) by NMR and HPLC-Q-Orbitrap-MS/MS: Changes in isolated phenolics and antioxidant effects through diverse growth times. Food Chem X.. 2023;17:100557
- [CrossRef] [Google Scholar]
- Quality consistency evaluation between dispensing granules and traditional decoction of Gardeniae Fructus based on chemical similarity and bioequivalence. J Pharm Biomed Anal.. 2022;213:114708
- [CrossRef] [Google Scholar]
- Research progress of Shegan Mahuang Decoction and predictive analysis on its Q-markers. China Journal of Chinese Materia Medica.. 2023;48(08):2068-2076.
- [CrossRef] [Google Scholar]
- Quality Evaluation of Gastrodia Elata Tubers Based on HPLC Fingerprint Analyses and Quantitative Analysis of Multi-Components by Single Marker. Molecules.. 2019;24(8):1521.
- [CrossRef] [Google Scholar]
- An ultra-robust fingerprinting method for quality assessment of traditional Chinese medicine using multiple reaction monitoring mass spectrometry. J Pharm Anal.. 2021;11(1):88-95.
- [CrossRef] [Google Scholar]
- Bergamotane-type sesquiterpenes from the leaves and twigs of Illicium oligandrum and their anti-inflammatory activities. Phytochemistry.. 2023;209:113617
- [CrossRef] [Google Scholar]
- Anti-inflammatory effect of the compounds from the flowers of Trollius chinensis. Pak. J. Pharm. Sci.. 2018;31(5):1951-1957.
- [Google Scholar]
- A new strategy for the preparation of total iridoids from Radix Gentianae Macrophyllae and anti-inflammatory profile digesting by UPLC-Q-TOF-MS characterization coupled with PLS analysis. Industrial Crops and Products.. 2021;168:113586
- [CrossRef] [Google Scholar]
- A comprehensive quality control evaluation for standard decoction of Smilax glabra Roxb based on HPLC-MS-UV/CAD methods combined with chemometrics analysis and network pharmacology. Food Chem.. 2023;410:135371
- [CrossRef] [Google Scholar]
- Sun, Y., Du, Y., Sheng, N., Yuan, L., Yu, X., Liu, M., Zhang, L., 2013. Simultaneous Determination of Seven Components in Jinlianqingre Granules by HPLC-MS. Chinese Pharmaceutical Journal. 48(16), 1407-1411. Beijing, China.
- Study on quality control of Zuojin pill by HPLC fingerprint with quantitative analysis of multi-components by single marker method and antioxidant activity analysis. J Pharm Biomed Anal.. 2023;225:115075
- [CrossRef] [Google Scholar]
- Undescribed sesquiterpene coumarins from the aerial parts of Ferula sinkiangensis and their anti-inflammatory activities in lipopolysaccharide-stimulated RAW 264.7 macrophages. Phytochemistry.. 2023;210:113664
- [CrossRef] [Google Scholar]
- Veratric Acid Inhibits LPS-Induced IL-6 and IL-8 Production in Human Gingival Fibroblasts. Inflammation.. 2015;39(1):237-242.
- [CrossRef] [Google Scholar]
- A Q-marker screening strategy based on ADME studies and systems biology for Chinese herbal medicine, taking Qianghuo Shengshi decoction in treating rheumatoid arthritis as an example. Mol Omics 2023
- [CrossRef] [Google Scholar]
- Comparative antioxidant activity and untargeted metabolomic analyses of cherry extracts of two Chinese cherry species based on UPLC-QTOF/MS and machine learning algorithms. Food Res Int.. 2023;171:113059
- [CrossRef] [Google Scholar]
- Establishment and validation of the quantitative analysis of multi-components by single marker for the quality control of Qishen Yiqi dripping pills by high-performance liquid chromatography with charged aerosol detection. Phytochem Anal.. 2021;32(6):942-956.
- [CrossRef] [Google Scholar]
- Fingerprint and multi-ingredient quantitative analyses for quality evaluation of hawthorn leaves and Guang hawthorn leaves by UPLC–MS. Revista Brasileira De Farmacognosia.. 2018;28(3):369-373.
- [CrossRef] [Google Scholar]
- Identification of anti-inflammatory components in Dioscorea nipponica Makino based on HPLC-MS/MS, quantitative analysis of multiple components by single marker and chemometric methods. J Chromatogr B Analyt Technol Biomed Life Sci.. 2022;1213(15):123531
- [CrossRef] [Google Scholar]
- Integrating metabolomic data with machine learning approach for discovery of Q-markers from Jinqi Jiangtang preparation against type 2 diabetes. Chin Med.. 2021;16(1):30.
- [CrossRef] [Google Scholar]
- Guizhi Shaoyao Zhimu granules attenuate bone destruction in mice with collagen-induced arthritis by promoting mitophagy of osteoclast precursors to inhibit osteoclastogenesis. Phytomedicine.. 2023;118:154967
- [CrossRef] [Google Scholar]
- Novel assays for quality evaluation of XueBiJing: Quality variability of a Chinese herbal injection for sepsis management. Pharm Anal.. 2022;12(4):664-682.
- [CrossRef] [Google Scholar]
- Orientin and vitexin attenuate lipopolysaccharide-induced inflammatory responses in RAW264.7 cells: a molecular docking study, biochemical characterization, and mechanism analysis. Food Science and Human Wellness.. 2022;11(5):1273-1281.
- [CrossRef] [Google Scholar]
- Amygdalin Delays Cartilage Endplate Degeneration and Improves Intervertebral Disc Degeneration by Inhibiting NF-κB Signaling Pathway and Inflammatory Response. J Inflamm Res.. 2023;16:3455-3468.
- [CrossRef] [Google Scholar]
- Zhang, J., Song. G., Li. J., 2012. Observation on the therapeutic effect of Jinlian Qingre Granules on mild type A H1N1 influenza. Shaanxi Journal of Traditional Chinese Medicine. 33(08), 957-959. Shaanxi, China.
- A strategy for antioxidant quality evaluation of Aster yunnanensis based on fingerprint-activity relationship modeling and chemometric analysis. Arabian Journal of Chemistry.. 2023;16:104755
- [CrossRef] [Google Scholar]
- E-eye, flash GC E-nose and HS-GC-MS combined with chemometrics to identify the adulterants and geographical origins of Ziziphi Spinosae Semen. Food Chem.. 2023;424:136270
- [CrossRef] [Google Scholar]
- Analysis of Bitter Almonds and Processed Products Based on HPLC-Fingerprints and Chemometry. Chem Biodivers.. 2023;20(3):e202200989.
- [Google Scholar]
- Chromatographic fingerprint combined with quantitative analysis of multi-components by single-marker for quality control of total lignans from Fructus arctii by high-performance liquid chromatography. Phytochem Anal.. 2022;33(8):1214-1224.
- [CrossRef] [Google Scholar]
- Quality evaluation of Acanthopanax senticosus via quantitative analysis of multiple components by single marker and multivariate data analysis. J Pharm Biomed Anal.. 2021;201:114090
- [CrossRef] [Google Scholar]
- Influence of Ultra-High-Pressure Pretreatment Method on Chemical Constituents, Antioxidant and Cytoprotective Activities of Free, Esterified, and Bound Phenolics from Anneslea Fragrans Wall. Leaves. Plant Foods Hum Nutr.. 2023;78:407-418.
- [CrossRef] [Google Scholar]
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105481.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







