Translate this page into:
Molecular pathways of generation and detoxification of reactive oxygen species and induction of apoptosis in yeasts
⁎Corresponding author. rehman.mmg@pu.edu.pk (Abdul Rehman)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Reactive oxygen species (ROS) can trigger cell death in Saccharomyces cerevisiae and other lower eukaryotic organisms. Protein structure and function may be altered by ROS produced by cellular metabolism or by exposure to environmental oxidants. This happens regardless of whether peroxidases, superoxide dismutases, or catalases are active. Depending on the redox state of cysteine residues, oxidative alteration of sulfhydryl groups in proteins may inhibit protein activity or begin alternative pathways that affect cell function. Redox control is made possible by conserved, interdependent thioredoxin and glutaredoxin systems, which may play a comparable function in metal homeostasis in eukaryotic organisms. In response to oxidative stress, gene expression in S. cerevisiae is regulated by several transcription factors, including Yap1, and Gpx3 peroxidase, and one function of Yap1 is to regulate gene expression. Notwithstanding this information, many issues about the link between ROS production and apoptosis remain unsolved. ROS may disrupt cells or activate certain molecular and cellular pathways during cell death, promoting apoptosis. This review aims to elucidate ROS’s essence, their harm to cells, how S. cerevisiae reacts to ROS, and how ROS might affect cell integrity and cause cell death.
Keywords
Yeasts
Reactive oxygen species
Oxidative stress
ROS detoxification via enzymes
Glutathione
Apoptosis
1 Introduction
Reactive oxygen species (ROS) are acknowledged as unstable, partially reduced, and highly reactive derivatives of oxygen with unpaired electrons that are generated in various cellular physiological pathways as a by-product (Qamer et al., 2021). Oxygen (O2) is relatively inert despite having two outer-shell unpaired electrons. This is because the usual criteria for higher reactivity are not met since each of these electrons has the same spin. When an unpaired electron becomes excited and changes its spin, the resultant species, what we call singlet O2, sets off an extremely strong oxidant. The result of the reactive oxygen species (ROS) includes a range of molecules acting as either reductants or oxidants, such as superoxide anion (O2−) and hydrogen peroxide (H2O2). These molecules can impact the cellular redox homeostasis (Nazir et al., 2024).
ROS may produce beneficial and/or detrimental effects depending on the cell’s internal conditions (Zhang et al., 2018). Secondary ROS molecules and radicals with varying degrees of reactivity are produced when primary ROS interact with different parts of cells. Therefore, it has been proven that the mechanism responsible for cellular damage is associated with ROS. ROS plays an important physiological role of ROS in the normal functioning of the cells. Various diseases may arise where the concentration of ROS exceeds and overwhelms the body’s natural defense against them. Moreover, ROS may induce genomic alterations that affect cellular homeostasis and result in diseases including cancer. The present review describes toxicity generated by ROS and protective systems including antioxidant enzymes and antioxidant molecules to combat ROS toxicity in yeast cells (Fig. 1).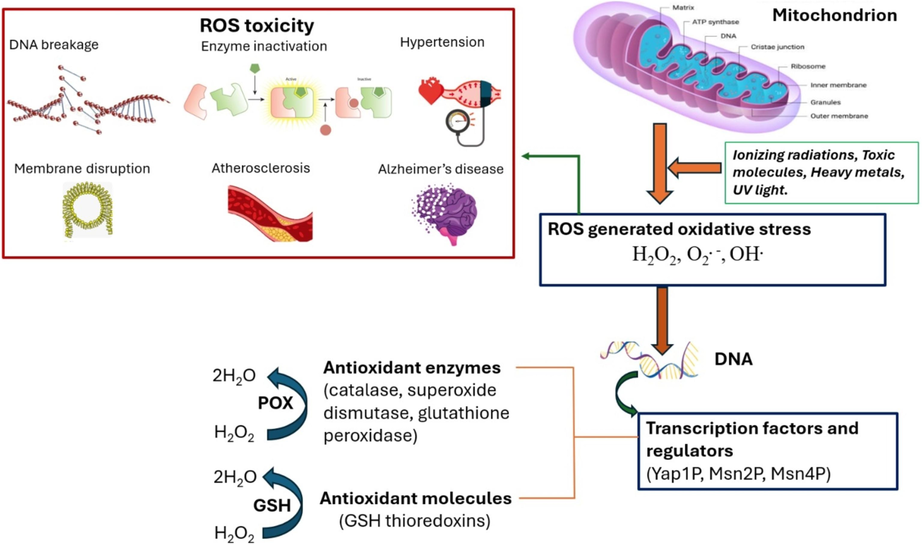
Schematic diagram showing ROS toxicity and protective systems to combat ROS toxic effects.
2 Reactive nitrogen species
Cells initiate the synthesis of reactive nitrogen species (RNS) and generate ROS via the interaction of oxygen with the nitric oxide radical (NO). Byproducts of this process include nitrogen dioxide radical (NO2) and reactive peroxynitrite (ONOO−). Peroxynitrator activation of aromatic amino acid residues can (Taysi et al., 2019; Gong et al., 2024), result in DNA lesion formation and thiol group oxidation. Peroxynitrite has higher reactivity than H2O2 when it comes to protein damage and thiol oxidation. Peroxynitrite has higher reactivity than H2O2 when it comes to protein damage and thiol oxidation (Mandal et al., 2022). ONOO− is a very active compound that forms when NO reacts with superoxide anions (O-2). It can modify thiol groups, resulting in the production of S-nitrosothiols, as well as nitrate tyrosine residues (Pérez de la Lastra et al., 2022). S. cerevisiae appears to generate NO in a hypoxic situation in mitochondria (Sunyer-Figueres et al., 2020). A reduction of nitric oxide generation lowers the risk of damage to the S-nitrosylation of glyceraldehyde-3-phosphate dehydrogenase (GAPDH), reduces ROS generation, and increases cell existence (Muronetz et al., 2017). NO can also be produced via reducing nitrite, in addition to NO synthase. These kinds of operations would be facilitated through mitochondrial nitrite reductase or cytochrome c reductase (Lancellotti et al., 2020; Zhan et al., 2024). Yeast hemoglobin (YHb) is an important enzyme that helps control levels of NO and protects the cell from harmful quantities of nitric oxide, especially when it comes from outside the cell. This provides further evidence that NO production is going on inside yeast cells as well (Poole, 2020).
3 Yeast cell-mediated production of reactive oxygen species
3.1 Mitochondria
The evolutionary history of oxygen’s emergence has ushered in the rage of stresses due to the various reactive species as the result of metabolism or sensitive metabolites' reaction with oxygen. Throughout the redox reaction, the ROS production is directly coupled with the concentration of oxygen (Sachdev et al., 2021). In yeast, the substantial intracellular source of ROS involves the extraction of electrons from the mitochondrial electron transport chain (ETC) during aerobic respiration (Fig. 2) under typical physiological circumstances (Payen et al., 2019). Despite numerous valid debates regarding the precise measurement of ROS production at the alternate site of the respiratory chain, there is a consensus on the controversy (Nair et al., 2018).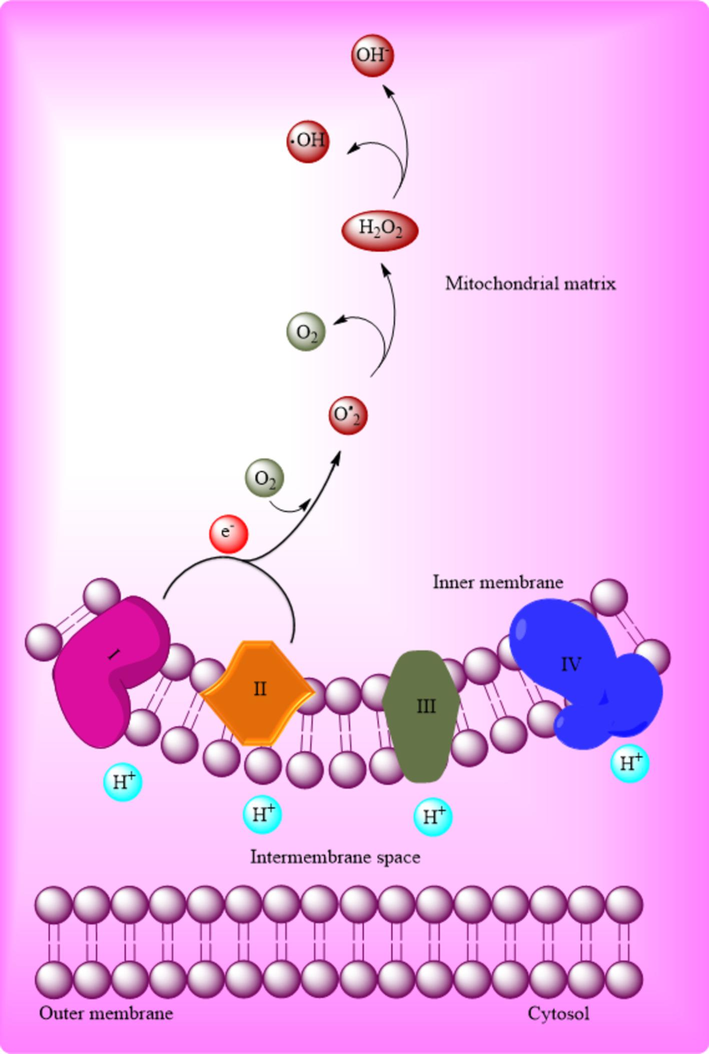
ROS formation in the mitochondrion (Noreen et al., 2018).
The inner membrane space of mitochondria in yeast cells contains three nicotinamide adenine dinucleotide dehydrogenases that are not sensitive to rotenone, regardless of whether complex 1 is present or not (Fig. 3) (Brambilla et al., 2020). NDI1 also known as internal NADH ubiquinone oxidoreductase, oxidized NADH which is generated in the mitochondrial matrix. Ndi1p, in addition to an oxidizing Nicotinamide adenine dinucleotide which is generated through ethanol oxidation via ethanol/acetaldehyde shuttle, has been hypothesized to also play a part in modulating the redox balance at the extent of mitochondrial NADHA, which is generated through the citric acid cycle (Signorile et al., 2019). Contrary to mammalian mitochondria yeast mitochondria are like plant mitochondria which oxidize cytosolic NADH directly (Zhao et al., 2020). Nde1 and Nde2 are two external NADH dehydrogenases found on the inner membrane of mitochondria, showing five active sites opposite the mitochondrial intermembrane gap. They facilitate the oxidation of cytosolic NADH, generated in the course of glycolysis because of the mitochondrial inner membrane's inability to transport NADH, but some scientists hypothesized that they are predominantly jumbled in respiration when cells were vegetated on ethanol (Mancebo et al., 2022). Due to the rotenone-insensitive status of Nde1p and Nde2P, they cannot pump protons (Fedoseeva et al., 2017). On the other hand, the glycerol-3-phosphate de-hydrogenase shuttle comprises cytosolic Nicotinamide adenine dinucleotide link G3PDH (Gpd1/2), and membrane-bound G3P; Ubiquinone, oxidoreductase can oxidize cytosolic NADH which is produced through glycolysis (Gut2) (Costa et al., 2021). In the ETC, the succinate dehydrogenase also called succinate dehydrogenase or complex II through FADH2 is liable for shifting the electron from succinate to coenzyme Q (ubiquinone). In S. cerevisiae all enduring pathways of respiratory FADH2 or NADH coincide at the ubiquinone pool. Through the cytochrome bc1 complex (complex III), ubiquinone transfers its electron to cytochrome c. Cytochrome c’s terminal oxidation is mediated by cytochrome c oxidase through molecular oxygen (cox or complex 1V) (Watson and McStay, 2020).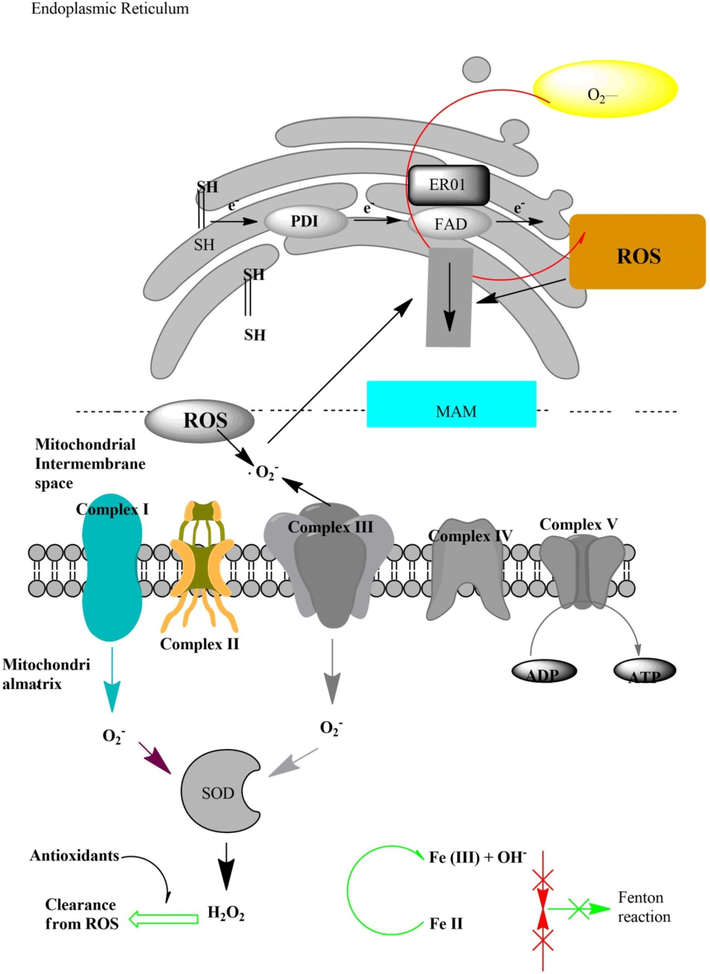
Generation of reactive oxygen species (ROS) in the ER and mitochondria in response to ER stress (Park et al., 2018).
3.2 Endoplasmic reticulum
The endoplasmic reticulum (ER) stands as one of the most extensive membrane-bound organelles, undertaking a diverse range of functions that include protein and lipid synthesis, folding, calcium storage and release, ER maturation, and protein secretory pathways (Schwarz and Blower, 2016).
The functioning of the protein disulfide bond-forming machinery (PDBFM) entails several components, including multiple proteins such as protein disulfide isomerase pdi1p, flavin-containing oxidoreductase, Ero1p, Eug1p, and Mdi1p. These components collaboratively participate in the process of protein folding in the ER under aerobic circumstances (Narayan, 2020). In aerobic environments, two electrons transfer to Ero1p/Pdi1p, where oxygen acts as a terminal electron acceptor, as a consequence of correct disulfide bond formation, which is critical for a protein to achieve its natural shape. (Fu et al., 2020). Fig. 3 shows that PDBFM produces H2O2 as a result of oxidative protein folding in the ER. H2O2 is a byproduct of the disulfide bonds formed in vitro by thioredoxin oxidation mediated by Ero1p (Peixoto et al., 2018; Mennerich et al., 2019). Consequently, it is anticipated that under normal physiological situations, ER set up a source of ROS in living cells, and the flow of electrons from reduced sulfhydryl to Pdi1p/Ero1p would be reflected in which the amount of ROS generated (Park et al., 2018) (Fig. 3).
ETC of mitochondria consists of four protein complexes (1–4) and an ATP synthetase resides inside the inner mitochondrial membrane. The electron transfer from NADH+ and FADH+ to NAD and FAD by complexes 1 and 4 produce O2− radicals which are then converted into H2O2 enzymatically by the catalytic activity of SOD. To suppress the generation of H2O2 various antioxidants such as catalase reacts with them and neutralize their effect. OH− can react with metals like Fe (already present in the cytosol) to start a chain of oxidation reaction and the subsequent generation of ROS, resulting from the Fenton reaction, leads to cellular damage (Hasan et al., 2024).
3.3 Peroxisomes
Peroxisome, previously known as a microbody, is a membrane-bounded organelle that is very important for the metabolism of D-amino acids, methanol, and fatty acids in yeasts. These influence the size, number, and enzyme content of peroxisomes and initiate their bio-genesis (Mast et al., 2020). It is reported that a signal called peroxisomal targeting signals (PTS) directs proteins to these microbodies' milieu. On the other hand, independent proteins importing pathways of PTS have also been reported (Kim and Hettema, 2015). The genesis of ROS, in most cases which is H2O2, is carried out by various peroxisomal enzymes, for example, the degradation of fatty acid during the participation of acyl-CoA oxidase in the β-oxidation pathway. Subsequently, yeast cells produce large amounts of H2O2 when they consume fatty acids, which in turn produce a large amount of catalase (Sen and Heerspink, 2021). Methanol may be chemically converted into formaldehyde and hydrogen peroxide with the help of alcohol oxidase, which is the first enzyme in the methanol metabolism process. H2O2 is also generated by urate oxidation to allantoin using urate oxidase (Kurylenko et al., 2019).
4 Types of ROS
4.1 Superoxide anion (O2•−)
ROS produces superoxide anion (O2−) as its main product. Oxygen is very abundant inside the living cell and relatively stable when there are two electrons in the outermost shell upon losing or reducing by an electron it yields a dangerous byproduct which is a product of oxygen metabolism.
For the most part, it is the result of the withdrawal of electrons from the respiratory chain of mitochondria, which is an end product of aerobic respiration (Wong et al., 2017). It is said to be hew through phagocytes during respiratory burst and microsomal metabolism. The superoxide anion has been reported to be directly involved in the reaction with some proteins but is not potentially reactive (Castro et al., 2019).
4.2 Hydrogen peroxide (H2O2)
Enzyme superoxide dismutase (SOD) catalyzes the transformation of superoxide ion into hydrogen peroxide. This conversion of H2O2 to water takes place as a result of several peroxidase reactions. The enzyme is a byproduct of fatty acid oxidation in the peroxisome and protein folding in the ER. Yeast produces H2O2 as a byproduct of O2-detoxification and several oxidase processes (Sachdev et al., 2021). H2O2 is reasonably unreactive and can eagerly pass through many biological membranes but has a detrimental outcome upon converting into tremendously reactive hydroxyl radical (OH−). A very reactive hypochlorite (CLO−) is formed from chloride ions and H2O2 through myeloperoxidase action by neutrophils during phagocytosis. CLO− which is generated through this reaction will now form the chloramines by reacting with free amines, which are extremely toxic to living cells (Maiorino et al., 2018).
4.3 Hydroxyl radical
Hydroxyl radical (OH−) is found to be very strongest oxidant in nature. H2O2 and O2 are combined in the Fenton and Haber-Weiss reaction, which is catalyzed through transition metal ions like Fe2+ which yield hydroxyl radicals (Walter et al., 2020). H2O2 upon reacting with Fe2+ yields the OH and OH− and leads to the production of Fe3+ which is then degraded again to Fe2+ through the action of O2− to relaunch the process. Several metabolites and bio-macromolecules react with this radical and yield secondary radicals (Kim et al., 2019a,b). The homeostasis process involved in Cu and Fe is vital for the cellular defense mechanism through Lassen down the damage spawned by ROS. Unsaturated fatty acids represent a primary target for the protonated O2−. This interaction leads to the formation of lipid radicals and the subsequent generation of toxic lipid hydroperoxides (Su et al., 2019). Carbonyls and methylglyoxal formed through the breakdown of lipid hydroperoxide and non-enzymatically by glycolysis respectively, which can both lead to the generation of carbonylation of protein and glycation. Several amino acids, predominantly arginine, and proline are attacked with metal-catalyzed forces to yield glutamic semialdehyde, whereas lysine generates aminoadipic semialdehyde (Bellahcène et al., 2018). These changed proteins assemble with chronological age resulting in apoptosis (Arlia Ciommo, 2018).
5 ROS detoxification via enzymes
5.1 Superoxide dismutase
The catalysts for the disproportionation of superoxide ions to hydrogen peroxide in S. cerevisiae have been identified as manganese-dependent superoxide dismutase (Mn-SOD) and copper, zinc-dependent superoxide dismutase (Cu, Zn-SOD). The major superoxide scavenger in mitochondria is Mn-SOD, Superoxide dismutase-2 gene encodes it and is responsible for 5–15 % of all SOD activity (Gangwar et al., 2020; Wang et al., 2021). Its activity in fermentative cells is fairly modest and can be increased by respiration, starvation, or ethanol exposure (Bagamery et al., 2020). The gene responsible for superoxide dismutase-1 encodes a SOD which contains copper and zinc. This enzyme is believed to be primarily located in the cytosol, with some presence in the mitochondrial intermembrane gap (Warris and Ballou, 2019). During respiration and fermentation, Cu, Zn-SOD accounts for up to 90 % of total SOD1 activity and around one percent of soluble protein in the cell (Montllor Albalate, 2021). It prevents oxidation of cytosolic and mitochondrial components and ensures their survival for the long term. After heat shock, a defect in cytosolic superoxide dismutase causes a rise in intracellular oxidation and rapid mortality in the stationary phase (Sunthonkun et al., 2019) (Table 1).
Mechanism
Gene(s)
Target Location*
Role
Metabolites of antioxidants
Glutathione
General
Redox Aqueous buffer, xenobiotic excretion
Ubiquinone
Mit, ER and Mem
Antioxidant (lipid soluble), Respiratory Chain components
D-erythroascorbate
Cyt?
Antioxidant (water-soluble)
Enzymes
Catalases
CTT1
Cyt
Dis-mutation of H2O2 to H2O + O2
CTA1
Per
Superoxide dismutases
SOD1
Cyt, Mit, and Nuc?
Dis-mutation of O2− to H2O + O2
SOD2
Mit
SOD chaperone
CCS1
Cyt
Needed for stacking copper in toSod1p
Thioredoxins
TRX1, TRX2
Cyt
Redox control, sulphate assimilation, Deoxyribonucleotide synthesis, a cofactor for some peroxidase.
Dithiol glutaredoxins
GRX1, GRX2
Cyt, Nuc
Enzymatic activities overlap with thioredoxins, and contain glutathione peroxidase activity.
Monothiol glutaredoxins
GRX3, GRX4, GRX5
Nuc, Mit, Matrix
Defend cells against redox damage Assembling Fe/S clusters
Peroxiredoxins
AHP1, PRX1
Cyt Mit
Reduction of alkyl peroxidase, ROOH + R’SH = R’SSR’+ROH
Thioredoxin peroxidases
TSA1
Cyt
H2O2 + RSH = H2O + RSSR peroxiredoxin activity, thioredoxin peroxidase activity Yap1p nuclear localization is also mediated by this protein.
Sulfiredoxin
SRX1
Cyt, Nuc
Reduced cysteine sulfinic acid residue in Ahp1p,Tas1p
Glutathione peroxidases
GPX1, GPX2
Cyt/Mem
Reduction of oxidized lipid hydroperoxides
Phospholipid hydroperoxide
GSH peroxidaseGPX3/ORP1
Cyt
Thioredoxin peroxidase activity is present. Yap1p receives a signal from the hydroperoxide receptor
cytochrome c peroxidase
CCP1
Mit inner membrane space
Hydrogen peroxide reduction in the mitochondrion
Glutathione synthesis
GSH1, GSH2
Cyt
GSH synthesis in stages
Glutathione reductase
GLR1
Cyt
GSSG to GSH reduction
Pentose phosphate pathway enzymes
ZWF1, GND1, TAL1, TKL1,2, RPE1,
ALO1, ARA2
Cyt
Mit Outer mem?NADPH is produced in order to recycle oxidized glutathione, glutaredoxins, and thioredoxins
D-erythroascorbate synthesis
Metal-binding proteins
CUP1, CRS5
Cyt
Multiple genes at this locus code for a copper-binding protein that also binds Cd2+.
Transcription factors, regulators
Yap1p
YAP1
Cyt/Nuc
oxidative stress, xenobiotic resistance, and cadmium
Skn7p
SKN7
Nuc
Auxiliary transcription factor that interacts with Yap1p to help with oxidative stress and also has a role in osmoregulation.
Msn2/4
MSN2, MSN4
Cyt/Nuc
Heat, hunger, osmotic, and oxidative stress all cause this transcription factor to react to the PKA pathway.
Yap1-binding protein
YBP1
Cyt
Yap1p nuclear localization is mediated in part by a protein.
Haem activated protein
HAP1
Nuc
HAP2,3,4,5
Nuc
Regulation of respiratory functions
Metal-binding activator
MAC1
Nuc
H2O2-induction of CTT1 is regulated by a gene that regulates genes involved in copper ion homeostasis
Cup2p
ACE1/CUP2
Nuc
CUP1 is activated by a copper-binding transcription factor when copper levels are high.
S. cerevisiae SOD1 and SOD2 mutations are oxygen hypersensitive. SOD deficiency results in poor aerobic development, a high amount of mutations, and a high susceptibility to redox-cycling medicines (Lalève et al., 2016; Rosa et al., 2021). SOD1 mutants have a range of symptoms, including poor respiratory growth, hypersensitivity to exogenous oxidants (like paraquat or menadione), stationary phase instability, and defects in methionine and lysine biosynthesis (Funada et al., 2022). Under internal oxidative stress circumstances, the pentose phosphate pathway is inhibited consequently, the NADPH pool is depleted, which is essential for methionine biosynthesis (Bertels et al., 2021). Lysine auxotrophy might be a result of damaged mitochondria's retrograde reaction, which affects the lysine biosynthesis process (Parkhitko et al., 2019). Even though superoxide dismutase is an important antioxidant enzyme, several studies have found that high dosages of superoxide dismutase promote lipid peroxidation, produce OH, and cause cell death (Fig. 4) (Case, 2017; Palma et al., 2020).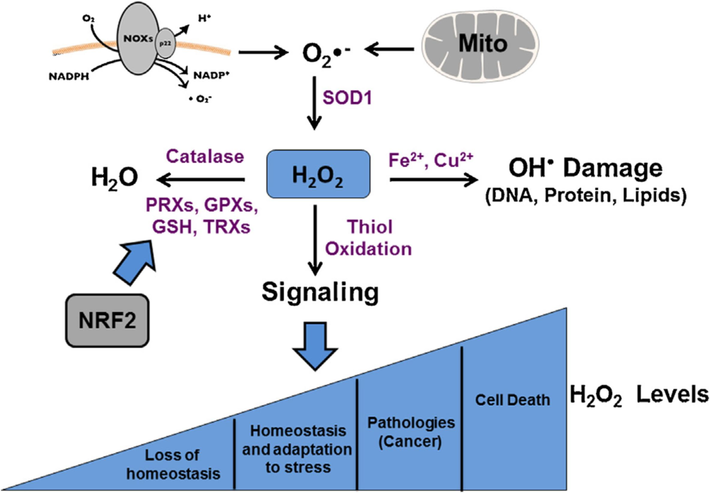
ROS regulation and cellular effects (Noreen et al., 2018).
5.2 Chaperone Ccs1
For SOD1 activity the intermolecular disulfide bond is very vital. The copper chaperone Ccs1 is important and its requirement is essential for the maturation of SOD1 and its inter-compartmentalization between mitochondria and cytosol (Barchiesi, 2018; Horn and Wittung-Stafshede, 2021). In addition to assisting in the delivery of copper to superoxide dismutase-1, the principal functions of the Cs1 chaperone include protein maturation and the synthesis of disulfide bonds. The absence of an intermolecular disulfide bond in sod1 facilitates protein aggregation (McAlary et al., 2019; Wright et al., 2019; Cox et al., 2020).
5.3 Catalase
S. cerevisiae has two enzymes and both contain heme. cytosolic catalase T is encoded by CTT1, while peroxisomal catalase is encoded by CTA1 (Dzanaeva et al., 2020; Yuan et al., 2021). Catalase T (CTT1) has been reported non-essential one for yeast in normal conditions perhaps it plays a crucial role in protection during severe stress situations (Guaragnella et al., 2019). CTA1 is proposed to have a function in the process of neutralizing H2O2 produced through fatty acid oxidation due to its coordinated regulation with peroxisomal structure and fatty acid metabolism (Deori et al., 2018; Fransen and Lismont, 2019). CTT1 is considered to have a broader function as an antioxidant during oxidative stress, several stresses may trigger its activation, including high temperatures, osmotic stress, hunger, and hydrogen peroxide (Miles et al., 2018). Furthermore, CTT1 lacking yeast strain was susceptible to heat stress and demonstrated low cross-adaption to hydrogen peroxide after mild heat stress (Bayliak et al., 2017; Galkina et al., 2020). The double cta1 ctt1 mutant grew normally under aerobic conditions and was insensitive to H2O2 raising issues regarding the actual role of catalases in yeast (Eleutherio et al., 2018). Nevertheless, after being pre-treated with a mild dosage of H2O2 and entering the stationary phase, catalase is shown to be important for peroxide resistance acquisition, suggesting its involvement in the adaptive response (Agashe and Kuzminov, 2021). The double cta1 ctt1 mutant exhibits hypersensitivity to peroxide during the stationary phase, indicating that both catalases work together to defend against external hydrogen peroxide (Xiao et al., 2018). Nonetheless, during oxidative, osmotic carbon deprivation, or freeze-thawing stressors, CTT1 expression is increased (Fig. 5) (Guaragnella et al., 2021).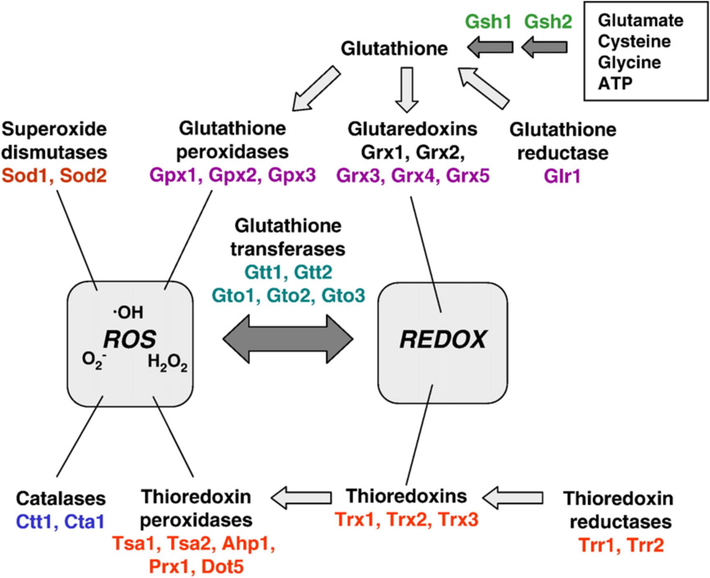
In S. cerevisiae, the enzyme systems involved in detoxifying reactive oxygen species and controlling the redox state of protein sulphydryl groups and their interrelationships (Herrero et al., 2008).
6 Glutathione pathway
The enzymes γ-glutamyl cysteine synthetase (GSH1) and glutathione synthase (GSH2) in yeast are responsible for producing glutathione (GSH), which is made up of glutamate, cysteine, and glycine (Fig. S1). Two glutaredoxin classes and one glutathione reductase have also been found involved in GSH genesis (Fig. 5). One well-known antioxidant in many species is the tripeptide GSH and is generally available as a most frequent non-protein thiol of low molecular weight and plays a postulated detoxification role for xenobiotics, ROS, and carcinogens (Fig. S2) (Sao Emani et al., 2018).
GSH may be used in yeast S. cerevisiae as an endogenous sulfur source (Chen et al., 2018). In the presence of sulfate, much of the sulfur is incorporated into GSH. Under the starved condition, GSH functioned as an endogenous sulfur supply until it reached a crucial residual level of about 10 % of its typical concentration. This led to a rise in GSH turnover and a decrease in γ-glutamyl transpeptidase (γ-GT: GSH + amino acid + H2O) and 3L γ-glutamyl-amino acid (L-glutamate). A single generation of S. cerevisiae on a minimal medium deficient in (NH4)2SO4 resulted in an increase in the GSH pool from 7 to 17 nM/mg dry weight (Chen et al., 2021). The yeast cytoplasm contained about 50 % of the GSH while the rest stood up in the central vacuole on nitrogen-sufficient media. The central vacuole accumulates GSH in response to nitrogen deprivation. In eukaryotic cells, in addition to being the major redox buffer, GSH is also an important marker of the redox status of a cell, in which the GSH/GSSG a redox potential couple is also considered. In wild-type S. cerevisiae cells, GSH concentration can range from 1–10 nM or even higher (Hasanuzzaman et al., 2019). In the TRX system, a GSH, substrate for GSTs, GRXs, and GPXs is also involved (Lyall et al., 2020). Since mutants devoid of GSH (gsh1) or with an altered GSH redox ratio (glr1) were susceptible to H2O2, GSH is a crucial antioxidant found in yeast cells that plays a vital role in protecting against H2O2 damage (Shao et al., 2019; Vázquez et al., 2017). Furthermore, H2O2 was discovered to change both the extent of GSH and the redox state in connection with a more oxidized state. As in typical aerobic situations yeast cells in the exponential phase exhibit a high redox ratio (GSH/GSSG), showing that more than 99.5 % of intracellular GSH is kept in reduced form (Picazo and Molin, 2021).
In eukaryotic cells, GSH is very important, in yeast cells needed as a reductant, which may serve to eliminate endogenously produced toxic metabolites (Cooper and Hanigan, 2018; Fabarius et al., 2021). Toxic lipid peroxidation products and oxidative stress caused by peroxides are problems for yeast strains that are either GSH-deficient or have changes to their GSH redox status. The redox-active sulfhydryl moiety in GSH's cysteine residue performs an inescapable role in its biological importance by acting as a free radical scavenger (Kim et al., 2019a,b). Thorsen et al. (2012) reported that GSH biosynthesis was boosted in S. cerevisiae when exposed to arsenite. A 128 % rise in GSH levels and a 66 % rise in the GSH/GSSG ratio were noted. It was also discovered that a 118.6 % rise in non-protein thiols was caused by Cd+2 stress (Hasanuzzaman et al., 2017). Furthermore, as reported by Meunier et al. (2004) ROS undergo a crucial oxidative change through the cysteine-to-heme enzyme of reticular membranes, found in the endoplasmic reticulum. One reason yeast cells are more resistant to metal-induced ROS is because they include cytochrome P450, cytoplasmic GSH, and non-protein thiols (Fig. S2) (Liu and Ma, 2020).
6.1 Monothiol glutaredoxins
In S. cerevisiae, monothiol GRX molecules were originally discovered, composed of three monothiol enzymes; Grx3, Grx4, and Grx5. They all have CGFS as their catalytic core (Chi et al., 2018; Mondal et al., 2020; Berndt et al., 2021;). Aft1, a transcription factor that senses iron in the nucleus, interacts with Grx3 and Grx4. In this case, when there is an iron shortage, Aft1 causes the transcription of genes belonging to an iron regulon (Gupta and Outten, 2020; Pujol-Carrion et al., 2021).
Aft1 was constitutively localized in Grx3 and Grx4, iron homeostasis was disrupted, and they became hypersensitive to oxidative stress. It is not known how Grx3 and Grx4 identify iron in yeast cells. The mitochondrial matrix contains Grx5, an enzyme involved in the last stages of iron/sulfur cluster synthesis (Ramos-Alonso et al., 2018; Wang et al., 2019). In eukaryotic cells, Grx5 is an inevitable component of the mitochondrial production of Fe/S clusters (Braymer et al., 2021; Pandey et al., 2019; Tsaousis, 2019). When it is absent, iron builds up and inhibits Fe/S enzymes in the respiratory electron chain. Enhanced sensitivity to oxidants is also shown by Grx5 mutants same as shown by Grx3 and Grx4 (Talib and Outten, 2021). Iron metabolism is regulated by Grx3, Grx4, and Grx5 (Chi et al., 2018; Mühlenhoff et al., 2020) (Fig. 2).
6.2 Dithiol glutaredoxins
The glutaredoxin system consists of glutathione reductase (GR), glutathione peroxidase (GPx), and glutathione-S-transferase (GST), which allows electrons to be transferred from NADPH to glutaredoxin via GSH (Miller et al., 2018; Zuo et al., 2022). Glutaredoxin is a tiny heat-stable oxidoreductase that is well preserved in the active sites during the development period (Kumar et al., 2020). Besides reducing dehydroascorbate and protecting cells from ROS, glutaredoxins are important for protein folding and regulatory activities in sulfur metabolism (Hasanuzzaman et al., 2019; Kim et al., 2020). Glutaredoxin may act as a catalyst for the dissolution of mixed disulfide complexes when exposed to oxidative stress (Fig. S3). Several proteins are rendered inactive by thiol oxidation, which may also restore the functionalities of proteins that have been oxidized (Barinova et al., 2017; Xiao et al., 2019). Glutaredoxin from bacterial and mammalian species share 4–50 % and 61–76 % identity with Grx1 and Grx2, respectively. Both enzymes are found in the cytoplasm, however ATG sites in the frame are responsible for alternative translation initiation, and a portion of Grx2 is also found in the mitochondria (Menon, 2015). The -C-P-Y-C- motif is found in the catalytic core of Grx1 and Grx2 (Liedgens et al., 2020; Roret et al., 2021). There was a concentration-dependent rise in Grx1 expression in response to Cd+2 stress. Evidence suggests that Candida tropicalis 3Aer Grx/GSH system directly reduces ROS damage caused by Cd+2.
Grx1 protects superoxide anion and hydroperoxide, based on a relevant mutation, but exclusively Grx2 provides hydroperoxide protection suggesting partial overlap functions (Auesukaree, 2017; Ulrich and Jakob, 2019). Although the double mutant strains were viable, they exhibited heat-sensitive oxidoreductase function when L-hydro-oxyethylene disulfide was used as the disulfide substrate. Surprisingly, both Grx1 and Grx2 have shown 64 % similarity, Grx1 mutants were not impacted whereas 5 % wild-type activity affected Grx2 mutants, which indicated that Grx2 played the leading role. Furthermore, Grx1 mutants were susceptible to superoxide anion-induced oxidative stress, but Grx2 mutants were selectively sensitive to H2O2, showing that Grx1 and Grx2 operate differently in a yeast cell. The stress response element (STRE) and Hog1 mitogen-activated protein kinase (MAPK) pathway control Grx1 and Grx2. Unfortunately, we still don't know what these proteins do or what they bind to (Gutin et al., 2015).
6.3 Thioredoxins (Trx) system
Three thioredoxins designated Trx1, Trx2, and Trx3, are found in S. cerevisiae. The cytoplasm contains two of them, while the mitochondria house the third (Dickson-Murray et al., 2021). The TRX reductase mutations show that the mitochondrial and cytosolic TRX systems work separately, based on the redox state of the TRX proteins. (Matsuzawa, 2017). S. cerevisiae's cytosolic TRX system is crucial for defense against externally introduced hydroperoxides. Individual and double mutant experiments with cytosolic TRXs indicated that their activities do not exactly coincide. Trx2 has been demonstrated to have a more essential function in the defensive system than Trx1 (Meyer et al., 2021). Similarly, when oxidants are supplied externally, Trx2 expresses more induction levels than Trx1. Trx2 metabolizes Tsa1 Trx peroxidase cytosolically, providing protection (Garrigós et al., 2020). It seems that the cytosolic Trx system does not play an important protective function against singlet oxygen (Gutiérrez-Escobedo et al., 2020). A set of proteins i.e., Tsa1, Tsa2, Ahp1, and Prx1, together with Gpx Prx1, belong to a small family of proteins that may hyperoxidize cysteine groups when Trx1 and Trx2 are not present. The involvement of cytoplasmic thioredoxins is vital in safeguarding against hydroperoxides (Curtis, 2019). During reductive stress, Trx2 expression is also increased by dithiothreitol as a disulphide-reducing agent which causes the development of unfolded protein response (UPR) (Zhang, 2019). While this is not due to the activation of UPR, the dithiothreitol is hypersensitive to two Trx1 and Trx2 proteins. The chaperone function of Tsa1 prevents the accumulation of misfolded ribosomal proteins under reductive stress, which occurs when Tsa1 is present in the cytoplasm (Dauplais et al., 2021; Picazo and Molin, 2021) (Fig. 2).
7 Transcription factors and regulators
7.1 Yap1p
Yap1p, a leucine zipper (bZip DNA-binding protein) AP-1 transcriptional component is known to be the foremost designated element of ROS sensing in S. cerevisiae, which also acts as a key regulator in response to oxidants. It regulates the production of cellular antioxidants and components involved in thiol reduction (Zhang et al., 2020). Yap1 binds to the Yap1 recognition element (YRE) in different gene promoters. Nevertheless, the discovery of several Yap1 targets that do not include YRE boxes suggests the presence of other recognition sites (Rodrigues-Pousada et al., 2019). Alongside its important part in oxidative stress response, Yap1 is activated by a variety of agents including methyl methane sulphonate, electrophile diethyl maleate, thiol oxidant diamide benomyl, cadmium, mercury, selenite quinone menadione, N-ethylmaleimide, methylglyoxal ionizing radiation, and drugs (Ansarypour and Shahpiri, 2017; Gast et al., 2021; Yilmaz, 2021). Because of its interaction with hydroperoxides and H2O2, Yap1p accumulates in the nucleus and influences the transcription of over 70 genes. Thioredoxin reductase, cytosolic superoxide dismutase, cytosolic catalase, and cytochrome c peroxidase are all encoded by the same set of genes. GSH metabolism and synthesis genes are also impacted by this process (Dawes and Perrone, 2020; Sen and Imlay, 2021). The C-CRD of Yap1 consists of a leucine-rich export signal (NES) and two cysteine-rich domains (CRD) positioned at the N- and C-termini. All the cysteine residue is provided through N- and C-CRD which is found in the Yap1 sequence and controls the Yap1 function under oxidative stress (Netto and Antunes, 2016; So et al., 2019). Because of the rapid nuclear export set off through nuclear export receptor (Crm1) in unstressed cells, Yap1 constrain in the cytoplasm (Kritsiligkou, 2016). Yap1 nuclear export is inhibited through diamide and hydroperoxides, which impair the connection between Yap1 and Crm1, resulting in Yap1 nuclear accumulation (So et al., 2019). A redox-dependent Yap1 NES, which is activated through two distinct pathways, depends on the kind of oxidant that disrupts the Yap1-Crm1 complex.
The same regulator is triggered YaP2 by two molecular redox centers, each in a different way. The occurrence of a ROS-induced reaction requires the presence of both C- and N-CRD redox centers, involving superoxide anions and hyperoxides. Conversely, a chemical reaction that involves the interaction of thiol groups with divalent heavy metal cations and electrophiles may be achieved only via the presence of C-CRD redox centers. Since an intramolecular disulfide bond forms between the N- and C-terminal CRDs (Cys310-Cys629 and Cys303-Cys598), the higher amounts of reactive oxygen species cause Yap1 to undergo transitory redox-induced conformational changes that hide the NES. As a result, the Yap1-Crms1 connection is disrupted and spurned, resulting in Yap1 nuclear accumulation and activation. The disulfide bond is reduced, causing structural changes that contribute to NES exposure (Mendoza-Martínez et al., 2020). Yap1 does not immediately start the oxidation of hydroperoxides.
The Gpx3 peroxidase, also known as Orp1, acts as H2O2 detector and transmits the signal to Yap1. Ybp1 is an essential component that forms a complex with Yap1 by specifically attaching to its C-terminus. It also plays a critical role in the activation of Yap1 caused by H2O2 (Yaakoub et al., 2022). Although the specific molecular mechanisms of Ybp1 are not yet fully understood, it is hypothesized that Ybp1 plays a role in relaying signals from Gpx3 to Yap1. The basis for this presumption is the fact that the synthesis of the Gpx3-Yap1 intermediate relies on the existence of Ybp1 (Tomalin, 2015; Curtis, 2019). Sulfonic acid (Cys36-SOH) is produced when peroxides directly oxidize Gpx3′s catalytic Cys36. A temporary Yap1-Gpx3 disulfide intermediate is formed when this sulfonic acid reacts with Yap1 Cys598. The Yap1 intramolecular disulfide bond is formed once this intermediate is resolved. Since Gpx3 is responsible for detecting and relaying an increase in ROS levels to Yap1, Yap1 becomes a vital member of a two-component system. Afterward, Yap1 promotes the activation of genes (Montibus et al., 2015; Mendoza-Martínez et al., 2017).
7.2 Msn2p and Msn4p and others
Msn2 and Msn4 (Zn-fingers) have been discovered as transcription factors that mediate various stress responses (Gupta, 2020). As they transition towards diauxic growth, face challenges like high temperature, changes in osmotic pressure, acidity, and exposure to hydrogen peroxide. These stressors attach themselves to the DNA at the CCCCT (a component of the stress response element) (Święciło, 2016). Both compounds endure a reversible translocation into the core as a result of oxidative stress. The molecular mechanisms that regulate this phenomenon, nevertheless, necessitate further explanation. An unexpected finding emerged suggesting that a single instance of the STRE component (CCCCT) was sufficient to increase the expression of a reporter construct in response to thermal stress. On the contrary, the stress-induced expression was significantly enhanced by the additive effect of multiple copies (Alford, 2020). HSP12, TPS2 (GSY2), SOD2, CTT1, and DDR2 are genes identified as being regulated by STRE. SKn7, alternatively referred to as Pos9 (peroxide sensitivity), controls the susceptibility of S. cerevisiae to H2O2, methyl viologen, and elevated oxygen pressure. Previously, this factor was thought to be insignificant in typical conditions (Ruta et al., 2020). The sequencing of Pos9 has unveiled its association with Skn7. There are notable resemblances between it and a bacterial protein that is linked to the two-component signal transduction system. Skn7 is susceptible to phosphorylation in response to hyperosmotic stress; this modification inhibits the oxidative stress response. Nevertheless, the specific molecular mechanisms that govern their operation continue to be mysterious. Skn7 proteins are implicated in yeast pathogens, including C. albicans and Cryptococcus neoformans, by constitutively protecting against oxidation, thereby boosting their pathogenic potential (Pais et al., 2016). One protein that helps fission yeast deal with oxidative stress is Prr1, which is similar to Skn7 (Papadakis and Workman, 2015; Shehzad et al., 2024).
CRediT authorship contribution statement
Sidra Gull: Methodology, Investigation. Chaudhry Ahmed Shabbir: Methodology, Investigation, Data curation. Zaman Khan: Writing – original draft, Investigation. Dilara A. Bukhari: Writing – original draft, Supervision, Investigation. Abdul Rehman: Writing – review & editing, Supervision, Conceptualization.
Funding
The present study was not granted any specified financial support from public, commercial, or not-for-profit funding agencies.
Conflict of interest
None of them have any declared conflicts of interest. Each author made a separate contribution to the manuscript's conception, composition, and final revision.
References
- Catalase inhibition by nitric oxide potentiates hydrogen peroxide to trigger catastrophic chromosome fragmentation in Escherichia coli. Genetics. 2021;218:iyab057.
- [Google Scholar]
- Identifying Mechanisms of Regulation and Signal Integration of the Heat Shock Response. Stanford University; 2020.
- Heterologous expression of a rice metallothionein isoform (Osmti-1b) in Saccharomyces cerevisiae enhances cadmium, hydrogen peroxide and ethanol tolerance. Braz. J. Microbiol.. 2017;48:537-543.
- [Google Scholar]
- Mechanisms of Longevity Extension by Caloric Restriction And Lithocholic Acid IN The Yeast Saccharomyces cerevisiae. Concordia University; 2018.
- Molecular mechanisms of the yeast adaptive response and tolerance to stresses encountered during ethanol fermentation. J. Biosci. Bioeng.. 2017;124:133-142.
- [Google Scholar]
- A putative Bet-Hedging strategy buffers budding yeast against environmental instability. Curr. Biol.. 2020;30(4563–4578):e4564.
- [Google Scholar]
- Barchiesi, A., 2018. The Mitochondrial Nature of the DNA Repair Protein Ape1.
- S-glutathionylation of glyceraldehyde-3-phosphate dehydrogenase induces formation of C150-C154 intrasubunit disulfide bond in the active site of the enzyme. Biochim. Biophys. Acta (BBA)-Gen. Subj.. 2017;1861:3167-3177.
- [Google Scholar]
- Growth on alpha-ketoglutarate increases oxidative stress resistance in the yeast Saccharomyces cerevisiae. Int. J. Microbiol.. 2017;2017:5792192.
- [Google Scholar]
- Methylglyoxal-derived stress: An emerging biological factor involved in the onset and progression of cancer. Semin. Cancer Biol.. 2018;49:64-74.
- [Google Scholar]
- Berndt, C., Christ, L., Rouhier, N., Mühlenhoff, U., 2021. Glutaredoxins with iron-sulphur clusters in eukaryotes-structure, function and impact on disease. Biochim. Biophys. Acta (BBA)-Bioenergetics 1862, 148317.
- The pentose phosphate pathway in yeasts–more than a poor cousin of glycolysis. Biomolecules. 2021;11:725.
- [Google Scholar]
- Stat3 inhibitor Opb-51602 is cytotoxic to tumor cells through inhibition of complex I and ROS induction. Iscience. 2020;23:101822
- [Google Scholar]
- Mechanistic concepts of iron-sulfur protein biogenesis in biology. Biochim. Biophys. Acta (BBA)-Mol. Cell Res.. 2021;1868:118863
- [Google Scholar]
- On the origin of superoxide dismutase: an evolutionary perspective of superoxide-mediated redox signaling. Antioxidants. 2017;6:82.
- [Google Scholar]
- Aconitases: non-redox iron–sulfur proteins sensitive to reactive species. Acc. Chem. Res.. 2019;52:2609-2619.
- [Google Scholar]
- Cross-kingdom comparative transcriptomics reveals conserved genetic modules in response to cadmium stress. Msystems. 2021;6 e01189-01121
- [Google Scholar]
- The complete pathway for thiosulfate utilization in Saccharomyces cerevisiae. Appl. Environ. Microbiol.. 2018;84 e01241-1218
- [Google Scholar]
- Structural and biochemical insights into the multiple functions of yeast Grx3. J. Mol. Biol.. 2018;430:1235-1248.
- [Google Scholar]
- Metabolism of glutathione S-conjugates: Multiple pathways. Compreh. Toxicol.. 2018;2018:363-406.
- [Google Scholar]
- Unraveling the conformational dynamics of glycerol 3-phosphate dehydrogenase, a nicotinamide adenine dinucleotide-dependent enzyme of Leishmania mexicana. J. Biomol. Struct. Dyn.. 2021;39:2044-2055.
- [Google Scholar]
- Protein aggregation in cell biology: An aggregomics perspective of health and disease. Semin. Cell Dev. Biol.. 2020;99:40-54.
- [Google Scholar]
- The Regulation of Deubiquitinases by Oxidative Stress in Saccharomyces cerevisiae. Newcastle University; 2019.
- Exposure to the methylselenol precursor dimethyldiselenide induces a reductive endoplasmic reticulum stress in Saccharomyces cerevisiae. Int. J. Mol. Sci.. 2021;22:5467.
- [Google Scholar]
- Peroxisomes: role in cellular ageing and age related disorders. Biogerontology. 2018;19:303-324.
- [Google Scholar]
- The Mia40/Chchd4 oxidative folding system: redox regulation and signaling in the mitochondrial intermembrane space. Antioxidants. 2021;10:592.
- [Google Scholar]
- The role of peroxisomes in xylose alcoholic fermentation in the engineered Saccharomyces cerevisiae. Cell Biol. Int.. 2020;44:1606-1615.
- [Google Scholar]
- Oxidative stress and aging: learning from yeast lessons. Fungal Biol.. 2018;122:514-525.
- [Google Scholar]
- Synthetic methylotrophy in yeasts: towards a circular bioeconomy. Trends Biotechnol.. 2021;39:348-358.
- [Google Scholar]
- The role of flavin-containing enzymes in mitochondrial membrane hyperpolarization and ROS production in respiring Saccharomyces cerevisiae cells under heat-shock conditions. Sci. Rep.. 2017;7:1-14.
- [Google Scholar]
- Redox signaling from and to peroxisomes: progress, challenges, and prospects. Antioxid. Redox Signal.. 2019;30:95-112.
- [Google Scholar]
- Pdi-regulated disulfide bond formation in protein folding and biomolecular assembly. Molecules. 2020;26:171.
- [Google Scholar]
- Funada, C., Tanino, N., Fukaya, M., Mikajiri, Y., Nishiguchi, M., Otake, M., et al. 2022. Sod1 mutations cause hypersensitivity to high-pressure-induced oxidative stress in Saccharomyces cerevisiae. Biochim. Biophys. Acta (BBA)-Gen. Subj. 1866, 130049.
- Deletion of Cdr1 reveals redox regulation of pleiotropic drug resistance in Candida glabrata. Biochimie. 2020;170:49-56.
- [Google Scholar]
- Characterisation of recombinant thermostable manganese-superoxide dismutase (Nemnsod) from Nerium oleander. Mol. Biol. Rep.. 2020;47:3251-3270.
- [Google Scholar]
- Wine yeast peroxiredoxin Tsa1 plays a role in growth, stress response and trehalose metabolism in biomass propagation. Microorganisms. 2020;8:1537.
- [Google Scholar]
- The yeast Eif2 kinase Gcn2 facilitates H2O2-mediated feedback inhibition of both protein synthesis and endoplasmic reticulum oxidative folding during recombinant protein production. Appl. Environ. Microbiol.. 2021;87 e00301-e321
- [Google Scholar]
- Novel electrochemical sensor strategy: Si-69 modified β-cyclodextrin/nitrogen-doped reduced graphene synthesis and characterization for fipronil detection. J. Environ. Chem. Eng.. 2024;12(5):113876
- [Google Scholar]
- Acid stress triggers resistance to acetic acid-induced regulated cell death through Hog1 activation which requires RTG2 in yeast. Oxidat. Med. Cell. Longev.. 2019;2019:4651062
- [Google Scholar]
- Rtg signaling sustains mitochondrial respiratory capacity in Hog1-dependent osmoadaptation. Microorganisms. 2021;9:1894
- [Google Scholar]
- Iron–sulfur cluster signaling: The common thread in fungal iron regulation. Curr. Opinion Chem. Biol.. 2020;55:189-201.
- [Google Scholar]
- Gupta, M. 2020. Investigating the Role of Iron-Sulfur Signaling in Yeast Transcriptional Regulation.
- Candida glabrata peroxiredoxins, Tsa1 and Tsa2, and sulfiredoxin, Srx1, protect against oxidative damage and are necessary for virulence. Fungal Genet. Biol.. 2020;135:103287
- [Google Scholar]
- Condition‐specific genetic interaction MAPS reveal crosstalk between the Camp/Pka and the Hog MAPK pathways in the activation of the general stress response. Mol. Syst. Biol.. 2015;11:829.
- [Google Scholar]
- Biogenic zinc oxide nanoregulator determines the quantitative analysis of morpho-anatomical and antioxidant capacity in Lactuca sativa L. Food Sci. Nutr.. 2024;00:1-14.
- [Google Scholar]
- Glutathione in plants: biosynthesis and physiological role in environmental stress tolerance. Physiol. Mol. Biol. Plants. 2017;23:249-268.
- [Google Scholar]
- Regulation of ascorbate-glutathione pathway in mitigating oxidative damage in plants under abiotic stress. Antioxidants. 2019;8:384.
- [Google Scholar]
- Redox control and oxidative stress in yeast cells. Biochim. Biophys. Acta (BBA)-Gen. Subj.. 2008;1780:1217-1235.
- [Google Scholar]
- ATP7A-regulated enzyme metalation and trafficking in the menkes disease puzzle. Biomedicines. 2021;9:391.
- [Google Scholar]
- Multiple pathways for protein transport to peroxisomes. J. Mol. Biol.. 2015;427:1176-1190.
- [Google Scholar]
- Heavy metal toxicity: an update of chelating therapeutic strategies. J. Trace Elem. Med. Biol.. 2019;54:226-231.
- [Google Scholar]
- Homogeneous photocatalytic Fe3+/Fe2+ redox cycle for simultaneous Cr(VI) reduction and organic pollutant oxidation: roles of hydroxyl radical and degradation intermediates. J. Hazard. Mater.. 2019;372:121-128.
- [Google Scholar]
- Molecular characterization, redox regulation, and immune responses of monothiol and dithiol glutaredoxins from disk abalone (Haliotis discus discus) Fish Shellfish Immunol.. 2020;107:385-394.
- [Google Scholar]
- Peroxiredoxins: Yeast Redox Switches That Regulate Multiple Cellular Pathways. UK: University of Manchester; 2016.
- Over-expression of chickpea glutaredoxin (CaGrx) provides tolerance to heavy metals by reducing metal accumulation and improved physiological and antioxidant defence system. Ecotoxicol. Environ. Saf.. 2020;192:110252
- [Google Scholar]
- Kurylenko, O.O., Dmytruk, K.V., Sibirny, A., 2019. Glutathione metabolism in yeasts and construction of the advanced producers of this tripeptide. In: Non-Conventional Yeasts: From Basic Research to Application. Springer. p. 153-196.
- The antimalarial drug primaquine targets Fe–S cluster proteins and yeast respiratory growth. Redox Biol.. 2016;7:21-29.
- [Google Scholar]
- Adsorbing surface strongly influences the pseudoperoxidase and nitrite reductase activity of electrode-bound yeast cytochrome C. The effect of hydrophobic immobilization. Bioelectrochemistry. 2020;136:107628
- [Google Scholar]
- Quantitative assessment of the determinant structural differences between redox-active and inactive glutaredoxins. Nat. Commun.. 2020;11:1-18.
- [Google Scholar]
- Pathway-based signature transcriptional profiles as tolerance phenotypes for the adapted industrial yeast Saccharomyces cerevisiae resistant to furfural and Hmf. Appl. Microbiol. Biotechnol.. 2020;104:3473-3492.
- [Google Scholar]
- Comparative analysis of ROS network genes in extremophile eukaryotes. Int. J. Mol. Sci.. 2020;21:9131.
- [Google Scholar]
- Gpx4, Lipid peroxidation, and cell death: discoveries, rediscoveries, and open issues. Antioxid. Redox Signal.. 2018;29:61-74.
- [Google Scholar]
- Fungal patterns induce cytokine expression through fluxes of metabolic intermediates that support glycolysis and oxidative phosphorylation. J. Immunol.. 2022;208:2779-2794.
- [Google Scholar]
- Reactive oxygen species (ROS) and reactive nitrogen species (RNS) in plants–maintenance of structural individuality and functional blend. Adv. Redox Res.. 2022;5:100039
- [Google Scholar]
- Peroxisome prognostications: Exploring the birth, life, and death of an organelle. J. Cell Biol.. 2020;219:e201912100
- [Google Scholar]
- Thioredoxin and redox signaling: roles of the thioredoxin system in control of cell fate. Arch. Biochem. Biophys.. 2017;617:101-105.
- [Google Scholar]
- Prion-like propagation of protein misfolding and aggregation in amyotrophic lateral sclerosis. Front. Mol. Neurosci.. 2019;12:262.
- [Google Scholar]
- NapA mediates a redox regulation of the antioxidant response, carbon utilization and development in Aspergillus nidulans. Front. Microbiol.. 2017;8:516.
- [Google Scholar]
- Yap1 Homologs mediate more than the redox regulation of the antioxidant response in filamentous fungi. Fungal Biol.. 2020;124:253-262.
- [Google Scholar]
- Hypoxia and reactive oxygen species as modulators of endoplasmic reticulum and Golgi homeostasis. Antioxid. Redox Signal.. 2019;30:113-137.
- [Google Scholar]
- Menon, D., 2015. The biological roles of glutathione transferase omega 1. PhD thesis.
- Mechanism of oxidation reactions catalyzed by cytochrome P450 enzymes. Chem. Rev.. 2004;104:3947-3980.
- [Google Scholar]
- Shifting paradigms and novel players in cys-based redox regulation and ROS signaling in plants-and where to go Next. Biol. Chem.. 2021;402:399-423.
- [Google Scholar]
- The role of protein chaperones in the survival from anthracycline-induced oxidative stress in Saccharomyces cerevisiae. Int. J. Adv. Res.. 2018;6:144.
- [Google Scholar]
- NADPH-dependent and-independent disulfide reductase systems. Free Radic. Biol. Med.. 2018;127:248-261.
- [Google Scholar]
- Phylogenetic distribution and structural analyses of cyanobacterial glutaredoxins (Grxs) Computat. Biol. Chem.. 2020;84:107141
- [Google Scholar]
- Coupling of transcriptional response to oxidative stress and secondary metabolism regulation in filamentous fungi. Crit. Rev. Microbiol.. 2015;41:295-308.
- [Google Scholar]
- Montllor Albalate, C. 2021. Parsing the Dual Roles of Cu/Zn Superoxide Dismutase (Sod1) in Oxidative Stress Protection and Redox Signaling. Georgia Institute of Technology.
- Glutaredoxins and iron-sulfur protein biogenesis at the interface of redox biology and iron metabolism. Biol. Chem.. 2020;401:1407-1428.
- [Google Scholar]
- Glyceraldehyde-3-phosphate dehydrogenase: aggregation mechanisms and impact on amyloid neurodegenerative diseases. Int. J. Biol. Macromol.. 2017;100:55-66.
- [Google Scholar]
- Impaired mitochondrial fatty acid synthesis leads to neurodegeneration in mice. J. Neurosci.. 2018;38:9781-9800.
- [Google Scholar]
- Revisiting the formation of a native disulfide bond: consequences for protein regeneration and beyond. Molecules. 2020;25:5337.
- [Google Scholar]
- Zinc oxide nano-fertilizer differentially effect on morphological and physiological identity of redox-enzymes and biochemical attributes in wheat (Triticum aestivum L.) Sci. Rep.. 2024;14:13091.
- [Google Scholar]
- The roles of peroxiredoxin and thioredoxin in hydrogen peroxide sensing and in signal transduction. Mol. Cells. 2016;39:65.
- [Google Scholar]
- Reactive oxygen species: Synthesis and their relationship with cancer-A review. Pak. J. Zool.. 2018;50(5):1951-1963.
- [Google Scholar]
- Transcriptional control of drug resistance, virulence and immune system evasion in pathogenic fungi: A cross-species comparison. Front. Cell. Infect. Microbiol.. 2016;6:131.
- [Google Scholar]
- Mitochondrial superoxide dismutase: what the established, the intriguing, and the novel reveal about a key cellular redox switch. Antioxid. Redox Signal.. 2020;32:701-714.
- [Google Scholar]
- Mitochondria export iron–sulfur and sulfur intermediates to the cytoplasm for iron–sulfur cluster assembly and Trna thiolation in yeast. J. Biol. Chem.. 2019;294:9489-9502.
- [Google Scholar]
- Oxidative stress response pathways: fission yeast as archetype. Crit. Rev. Microbiol.. 2015;41:520-535.
- [Google Scholar]
- Enzymatic activity of endoplasmic reticulum oxidoreductin 1 from Bombyx mori. Int. J. Indust. Entomol.. 2018;37:15-20.
- [Google Scholar]
- Methionine metabolism and methyltransferases in the regulation of aging and lifespan extension across species. Aging Cell. 2019;18:e13034.
- [Google Scholar]
- Pro-and antitumor effects of mitochondrial reactive oxygen species. Cancer Metastasis Rev.. 2019;38:189-203.
- [Google Scholar]
- Peroxynitrite preferentially oxidizes the dithiol redox motifs of protein-disulfide isomerase. J. Biol. Chem.. 2018;293:1450-1465.
- [Google Scholar]
- The nitration of proteins, lipids and DNA by peroxynitrite derivatives-chemistry involved and biological relevance. Stresses. 2022;2:53-64.
- [Google Scholar]
- Impact of hydrogen peroxide on protein synthesis in yeast. Antioxidants. 2021;10:952.
- [Google Scholar]
- Poole, R.K., 2020. Flavohaemoglobin: The pre-eminent nitric oxide–detoxifying machine of microorganisms. F1000Res. 9:F1000 Faculty Rev-7.
- The MAPK Slt2/Mpk1 plays a role in iron homeostasis through direct regulation of the transcription factor Aft1. Biochim. Biophys. Acta (BBA)-Mol. Cell Res.. 2021;1868:118974
- [Google Scholar]
- Review of oxidative stress and antioxidative defense mechanisms in Gossypium hirsutum L. in response to extreme abiotic conditions. J. Cotton Res.. 2021;4:1-9.
- [Google Scholar]
- Molecular strategies to increase yeast iron accumulation and resistance. Metallomics. 2018;10:1245-1256.
- [Google Scholar]
- Yeast Ap-1 like transcription factors (Yap) and stress response: a current overview. Microbial Cell. 2019;6:267.
- [Google Scholar]
- Atypical iron-sulfur cluster binding, redox activity and structural properties of Chlamydomonas reinhardtii glutaredoxin 2. Antioxidants. 2021;10:803.
- [Google Scholar]
- Superoxide dismutase administration: a review of proposed human uses. Molecules. 2021;26:1844.
- [Google Scholar]
- Saccharomyces cerevisiae cells lacking transcription factors Skn7 or Yap1 exhibit different susceptibility to cyanidin. Heliyon. 2020;6:e05352
- [Google Scholar]
- Abiotic stress and reactive oxygen species: generation, signaling, and defense mechanisms. Antioxidants. 2021;10:277.
- [Google Scholar]
- Gamma-glutamylcysteine protects ergothioneine-deficient Mycobacterium tuberculosis mutants against oxidative and nitrosative stress. Biochem. Biophys. Res. Commun.. 2018;495:174-178.
- [Google Scholar]
- The endoplasmic reticulum: structure, function and response to cellular signaling. Cell. Mol.life Sci.. 2016;73:79-94.
- [Google Scholar]
- A kidney perspective on the mechanism of action of sodium glucose co-transporter 2 inhibitors. Cell Metab.. 2021;33:732-739.
- [Google Scholar]
- How microbes defend themselves from incoming hydrogen peroxide. Front. Immunol. 2021:1104.
- [Google Scholar]
- Oxidative stress response of Aspergillus oryzae induced by hydrogen peroxide and menadione sodium bisulfite. Microorganisms. 2019;7:225.
- [Google Scholar]
- Lead toxicity regulation via protein degradation and tetrapyrrole biosynthesis pathways in Brassica species: A comparative quantitative analysis of proteomic study. Plant Physiol. Biochem.. 2024;213:108867
- [Google Scholar]
- Prohibitins: a critical role in mitochondrial functions and implication in diseases. Cells. 2019;8:71.
- [Google Scholar]
- Regulatory mechanism of the atypical Ap-1-like transcription factor Yap1 in Cryptococcus neoformans. mSphere. 2019;4 e00785-00719
- [Google Scholar]
- Reactive oxygen species-induced lipid peroxidation in apoptosis, autophagy, and ferroptosis. Oxid. Med. Cell. Longev.. 2019;2019:5080843
- [Google Scholar]
- Life-span extension by pigmented rice bran in the model yeast Saccharomyces cerevisiae. Sci. Rep.. 2019;9:1-16.
- [Google Scholar]
- Transcriptomic insights into the effect of melatonin in Saccharomyces cerevisiae in the presence and absence of oxidative stress. Antioxidants. 2020;9:947.
- [Google Scholar]
- Cross-stress resistance in Saccharomyces cerevisiae yeast-new insight into an old phenomenon. Cell Stress Chaperon.. 2016;21:187-200.
- [Google Scholar]
- Talib, E.A., Outten, C.E., 2021. Iron-sulfur cluster biogenesis, trafficking, and signaling: Roles for Cgfs glutaredoxins and bola proteins. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 1868,118847.
- Radicals, oxidative/nitrosative stress and preeclampsia. Mini-Rev. Med. Chem.. 2019;19:178-193.
- [Google Scholar]
- Glutathione serves an extracellular defence function to decrease arsenite accumulation and toxicity in yeast. Mol. Microbiol.. 2012;84(6):1177-1188.
- [Google Scholar]
- A Systems Biology Approach to Investigate the Role of Peroxiredoxins in Responses to Hydrogen Peroxide. Newcastle University; 2015.
- On the origin of iron/sulfur cluster biosynthesis in eukaryotes. Front. Microbiol.. 2019;10:2478.
- [Google Scholar]
- Melatonin reduces oxidative stress damage induced by hydrogen peroxide in Saccharomyces cerevisiae. Front. Microbiol.. 2017;8:1066.
- [Google Scholar]
- Identifying the reactive sites of hydrogen peroxide decomposition and hydroxyl radical formation on chrysotile asbestos surfaces. Part. Fibre Toxicol.. 2020;17:1-15.
- [Google Scholar]
- A fungal ABC transporter FgAtm1 regulates iron homeostasis via the transcription factor cascade FgAreA-HapX. PLoS Pathog.. 2019;15:e1007791
- [Google Scholar]
- The extracellular superoxide dismutase Sod5 from Fusarium oxysporum is localized in response to external stimuli and contributes to fungal pathogenicity. Front. Plant Sci.. 2021;12:608861
- [Google Scholar]
- Oxidative responses and fungal infection biology. Semin Cell Dev Biol.. 2019;89:34-46.
- [Google Scholar]
- Functions of cytochrome C oxidase assembly factors. Int. J. Mol. Sci.. 2020;21:7254.
- [Google Scholar]
- Production of superoxide and hydrogen peroxide from specific mitochondrial sites under different bioenergetic conditions. J. Biol. Chem.. 2017;292:16804-16809.
- [Google Scholar]
- The biophysics of superoxide dismutase-1 and amyotrophic lateral sclerosis. Q. Rev. Biophys.. 2019;52:e12.
- [Google Scholar]
- Distinct proteome remodeling of industrial Saccharomyces cerevisiae in response to prolonged thermal stress or transient heat shock. J. Proteome Res.. 2018;17:1812-1825.
- [Google Scholar]
- Molecular mechanisms of glutaredoxin enzymes: Versatile hubs for thiol–disulfide exchange between protein thiols and glutathione. J. Mol. Biol.. 2019;431:158-177.
- [Google Scholar]
- The concept of oxidative stress by the yeast Saccharomyces cerevisiae perspective. In: Oxidative Stress and Antioxidant Defense System. 2021. p. :71.
- [Google Scholar]
- A secreted catalase contributes to Puccinia striiformis resistance to host-derived oxidative stress. Stress Biol.. 2021;1:1-15.
- [Google Scholar]
- The role of reduced graphene oxide on mitigation of lead phytotoxicity in Triticum aestivum L. plants at morphological and physiological levels. Plant Physiol Biochem.. 2024;211:108719
- [Google Scholar]
- ROS-responsive microneedle patch for acne vulgaris treatment. Adv. Ther.. 2018;1(3):1800035
- [Google Scholar]
- Role of Selenoproteins in redox regulation of signaling and the antioxidant system: a review. Antioxidants. 2020;9:383.
- [Google Scholar]
- Zhang, Y., 2019. The Relationship between P-glycoprotein, carbonic anhydrase Xii, and the thioredoxin system in Bortezomib Resistant Multiple Myeloma Cells.
- Malate circulation: linking chloroplast metabolism to mitochondrial ROS. Trends Plant Sci.. 2020;25:446-454.
- [Google Scholar]
- A fluorescent probe for specifically measuring the overall thioredoxin and glutaredoxin reducing activity in bacterial cells. Analyst. 2022;147:834-840.
- [Google Scholar]
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2024.106069.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary Data 1
Supplementary Data 1







