Translate this page into:
NMR-based metabolomics and UHPLC-ESI-MS/MS profiling of Syzygium jambos in relation to their antioxidant and anti-hyperglycemic activities
⁎Corresponding author. faridah_abas@upm.edu.my (Faridah ABAS)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
NMR and UHPLC-MS/MS identified 30 and 59 metabolites respectively in S. jambos leaf. S. jambos leaf extract showed antioxidant and anti-hyperglycemic properties. 70% ethanolic extract is the most active in all bioactivities tested. Tannins, triterpenoids and flavonoids contributed to the bioactivities. NMR metabolomics reveals the potential of S. jambos as functional ingredients.
Abstract
Search for natural sources in disease management especially diabetes has surged recently for its potential health benefits. However, several fruit tree species remained unpopular despite their folkloric usage due to insufficient scientific data. Syzygium jambos (L.) Alston (Myrtaceae) is a fruit-bearing plant that is used as a traditional dietary supplement to treat various diseases including diabetes. This study investigated the correlation between S. jambos leaves’ metabolites and the bioactivities using proton-nuclear magnetic resonance (1H NMR)-based metabolomics approach. In-vitro total phenolic content (TPC), 2,2-diphenyl-1-picrylhydrazyl (DPPH•) radicals scavenging, nitric oxide (NO•) radicals scavenging, anti-α-amylase and anti-α-glucosidase assays were performed. The findings indicated that 70 % ethanolic extract demonstrated the highest potential in overall bioactivities. A total of 59 and 30 metabolites were tentatively identified using ultrahigh-performance liquid chromatography electrospray ionization Quadrupole-Orbitrap tandem mass spectrometry (UHPLC/ESI Q-Orbitrap MS/MS) and NMR, respectively. The partial least square (PLS) model revealed tannins, triterpenoids, and flavonoids significantly contributed to the separation and bioactivities. This study provides comprehensive insights into S. jambos metabolome and reveals the potential as a reliable source of antioxidant and anti-hyperglycemic compounds.
Keywords
Syzygium jambos
1H NMR metabolomics
Antioxidant
Anti-hyperglycemic
UHPLC-ESI-MS/MS metabolites profiling
- AA
-
anti-α-amylase
- AG
-
anti-α-glucosidase
- CID
-
collision-induced dissociation
- DNS
-
3,5-Dinitrosalicylic acid
- DPPH•
-
2,2-diphenyl-1-picrylhydrazyl
- FA
-
formic acid
- HHDP
-
hexahydroxydiphenoyl
- HMBC
-
heteronuclear multiple bond correlation
- HSQC
-
heteronuclear single quantum coherence
- IDF
-
International Diabetes Federation
- MS
-
mass spectrometry
- NMR
-
nuclear magnetic resonance
- NO•
-
nitric oxide
- PCA
-
principal component analysis
- PLS
-
partial least square
- PNPG
-
p-nitrophenyl-α-D-glucopyranose
- SNP
-
sodium nitroprusside
- T2DM
-
type 2 diabetes mellitus
- TOCSY
-
total correlation spectroscopy
- TPC
-
total phenolic content
- UHPLC-ESI-MS
-
ultrahigh-performance liquid chromatography-electrospray ionization-mass spectrometry
- VIP
-
variable importance in the projection
Abbreviations
1 Introduction
Diabetes has appeared as one of the most common chronic diseases with life-threatening complications and reduction of life expectancy. According to International Diabetes Federation (IDF), the global prevalence of diabetes in adults aged 20–79 years old is estimated to rocket to 12.2 % in 2045 from 10.5 % in 2021 (Sun et al., 2022). Generally, type 2 diabetes mellitus (T2DM) accounted for most of the diabetes events which are characterized by its ineffective utilization of insulin and progressive insulin resistance. Over decades, hyperglycemia and impaired glucose tolerance in T2DM have been effectively controlled in several medical approaches including insulin and oral hypoglycemic drugs as antioxidant agents and carbohydrate hydrolyzing enzyme inhibitors (WHO, 2016). One of the strategic approaches in diabetes treatment is to mitigate postprandial hyperglycemia by delaying and reducing glucose absorption, which responsible for starch hydrolyzing enzymes, α-amylase and α-glucosidase (Alqahtani et al., 2020). In recent years, herbal therapy has been a leading field in managing diabetes together with conventional treatment procedures to enhance patients’ wellness and health systems. Therefore, scientific investigation is essential to reveal the natural therapeutic characteristics and to support the traditional claims of medicinal plants.
Syzygium jambos (L.) Alston (Myrtaceae), is a medium-sized fruit tree commonly known as rose apple or “jambu mawar” in Malaysia and is found to be native to Southeast Asia and cultivated in other warm tropical regions, such as East India and South America (Baliga et al., 2017). S. jambos has a history of being extensively used as herb and for folk medicine. A decoction of the leaves could be used as diuretic for treating rheumatism, inflammatory pain, and diabetes. Moreover, ingestion of the juice of macerated leaves can help alleviate fever, sore throat, and sore eyes, and powdered leaves applied topically can act as a cooling agent for patients with smallpox (Avila-Pena et al., 2007; Bonfanti et al., 2013; Morton, 1987). Other plant parts, such as fruit, flower, seed, and bark, can also be used as various traditional remedies for diarrhea, dysentery, asthma, bronchitis, and dysphonia (Morton, 1987). S. jambos was reported to exhibit antinociceptive, analgesic (Avila-Pena et al., 2007), anti-microbial, anti-inflammatory (Sharma et al., 2013), anti-cancer (Yang et al., 2000), and hepato-protective (Islam et al., 2012) properties. Phytochemical studies on S. jambos leaves have identified several secondary metabolites, including flavonoids (catechin, rutin, quercetin, and ellagic acid) (Hossain et al., 2016), hydrolyzable tannins (pedunculagin, casuarinin, and castalagin) (Yang et al., 2000), and dihydrochalcones (myrigalone derivatives and phloretin) (Jayasinghe et al., 2007).
In view of the reported anti-hyperglycemic effect of S. jambos (Gavillán-suárez et al., 2015), an investigation into the possibility of additive effects among plant bioactive compounds and its pharmacological implications is warranted. Metabolomics provides a thorough phytochemical profile rather than an individual compound analysis, which reveals the variation of phytoconstituents in different environments and the corresponding interactions (Wolfender et al., 2013). The common techniques used in metabolomics studies are nuclear magnetic resonance (NMR) and mass spectrometry (MS). Proton NMR (1H NMR) is a robust instrument for the metabolites analysis qualitatively and quantitatively in a matrix, whereas two-dimensional NMR (2D-NMR) is usually employed to increase resolution, particularly in overlapping signals (Wolfender et al., 2013). NMR is widely used coupled with multivariate data analysis in phytochemical studies (Kim et al., 2011).
In the search for solvent efficiency in the extraction of bioactive metabolites from the S. jambos leaves, this study aimed to investigate the metabolite difference in S. jambos leaves extracted with four ethanol/water ratios using the NMR-based metabolomics technique. This allows the annotation of phytochemicals and the correlation between the antioxidant and anti-hyperglycemic properties of the plant. Moreover, the present study provides a thorough metabolite profile of the S. jambos leaf extract using ultrahigh-performance liquid chromatography electrospray ionization (UHPLC/ESI) Orbitrap MS. Hence, this study provides a comprehensive profile for the distribution of the bioactive compounds in S. jambos leaves extracts as a reliable source of functional foods which further proves its traditional usage in diabetes management.
2 Materials and methods
2.1 Chemicals and reagents
Deuterated methanol‑d4 (CD3OD), deuterium oxide (D2O), non-deuterated potassium dihydrogen phosphate (KH4PO4), sodium deuterium oxide (NaOD), Folin–Ciocalteu reagent, trimethylsilylpropionic acid-d4 sodium salt (TSP) and absolute ethanol were obtained from Merck (Darmstadt, Germany). 2,2-diphenyl-1-picrylhydrazyl (DPPH•), sodium carbonate, sodium nitroprusside (SNP), α-amylase enzyme, α-glucosidase enzyme, glycine, phosphate buffered saline, p-nitrophenyl-α-D-glucopyranose (PNPG), sodium chloride (NaCl), 3,5-Dinitrosalicylic acid (DNS), sodium potassium tartrate tetrahydrate, potato starch, quercetin, gallic acid and acarbose were supplied by Sigma-Aldrich Co. (St Louis, MO, USA). Dimethyl sulfoxide (DMSO), LCMS-grade methanol, LCMS-grade water and formic acid (FA) were obtained from Fisher Scientific (Geel, Belgium).
2.2 Plant collection and extraction
Syzygium jambos (voucher number MFI 0053/19) was collected from Universiti Putra Malaysia (UPM), Malaysia, identified and deposited by an in-house botanist from Biodiversity Unit, Dr. Mohd Firdaus Ismail at Institute of Bioscience, UPM. The leaves were collected from six trees to serve as biological replications. The cleaned leaves were air-dried at room temperature (23–26 °C) before being pulverized into powder using a laboratory pulverizer (Waring Commercial, Torrington, CT, USA) (Mediani et al., 2014). The extraction process was performed by adding 10 g of plant sample into 100 mL of four different ethanol/water ratios (0 %, 50 %, 70 %, and absolute ethanol) then subjected to ultra-sonication for 1 h (53 kHz) under temperature 26–40 °C using a Thermo-10D Ultrasonic machine (Fisher Scientific, Waltham, MA, USA). The crude extracts were then filtered, and vacuum concentrated at 40 °C. The extraction procedure was conducted twice using the same filtration residue. All the weighed crude extracts were subjected to freeze-dry using ScanVac CoolSafe Freeze dryer (Labogene, Lynge, Denmark), and kept at 4 °C for future analysis.
2.3 Total phenolic content (TPC) assay
TPC was determined by Folin–Ciocalteu (FC) procedure with slight amendments (Zhang et al., 2006). 100 µL of FC reagent was mixed with 20 µL of 100 µg/mL sample in a 96-well plate. The sample was prepared by dissolution of freeze-dried crude extract with 10 % DMSO for in-vitro biological assays. After 5 min incubation, 80 µL of 7.5 % sodium carbonate was added to the well, and the mixture was then incubated in the dark for 30 min. Then, the absorbance was detected at 750 nm using Tecan Infinite F200 micro-plate reader (Tecan Group Ltd, Männedorf, Switzerland). Gallic acid was used to obtain the calibration curve. The results obtained were then expressed in mg of gallic acid equivalent per crude extract of S. jambos (mg GAE/g).
2.4 DPPH• free radicals scavenging assay
The DPPH• assay was performed using the described method (Wan et al., 2012) with slight modifications. 100 µL of DPPH• reagent was added into 50 µL of the serial diluted sample. The absorbance was then detected at 515 nm after 30 min of incubation in the dark at room temperature. The scavenging capacity (SC) was calculated as SC%=[(A0 - As)/A0] x 100, where A0 is the absorbance of reagent blank, while As is the absorbance of test sample. The results were expressed in half maximal inhibitory concentration (IC50), and quercetin was served as a positive control.
2.5 Nitric oxide (NO•) radicals scavenging assay
NO• radical scavenging assay was performed by mixing 60 µL of 10 mM SNP into 60 µL of the serial diluted plant sample and incubated for 150 min at room temperature. 60 µL of Griess reagent (freshly prepared) was then added to the mixture before the absorbance was detected at 550 nm. The obtained results were expressed in IC50, gallic acid and quercetin served as a positive control (Tsai et al., 2007).
2.6 Anti-α-amylase (AA) assay
The anti-α-amylase assay was evaluated using the method with relevant adjustments (Telagari & Hullatti, 2015). The enzymatic reaction was achieved using 1 % potato starch solution as substrate and α-amylase enzyme, both prepared in 100 mM sodium phosphate buffer (pH 6.9). A mixture of 10 µL serial diluted test samples and 50 µL of 100 mM buffer was mixed with 10 µL of α-amylase enzyme (2U/mL) in 96-well plate. The blank solvent and blank sample were comprised of 60 µL of 100 mM sodium phosphate buffer and 10 µL of solvent, and 60 µL of 100 mM sodium phosphate buffer and 10 µL of test sample, respectively. Negative control was assembled by replacing the test sample with solvent, and the mixture was then incubated for 20 min at 37 °C. After that, 20 µL of 1 % starch solution was mixed into the wells of test sample, positive and negative controls, whereas the rest was added with 20 µL of 100 mM sodium phosphate buffer. A volume of 100 µL DNS reagent was then added into the mixture to quench the reaction after incubation at 37 °C for 30 min. Then, the mixture was brought to 90 °C for 10 min before the absorbance reading was measured at 550 nm. The percentage of inhibition was determined as %Inhibition = [(an – as)/an] x 100, where an is the absorbance difference between blank and negative control, and as is the absorbance difference between a sample and blank sample. Positive control used in this assay was acarbose and the results were expressed in IC50.
2.7 Anti-α-glucosidase (AG) assay
This experiment was performed by the method described (Lee et al., 2019). The enzymatic reaction was accomplished using PNPG (substrate) and α-glucosidase (enzyme). A mixture of 10 µL serial diluted test samples and 130 µL of 30 mM phosphate buffer was mixed with 10 µL of α-glucosidase enzyme in a 96-well plate. The blank sample, blank solvent and negative control were prepared accordingly, and the mixture was then incubated for 5 min at room temperature. After that, 50 µL of PNPG was mixed with test sample, positive and negative controls, while the rest was added with 50 µL of 30 mM sodium phosphate buffer. A volume of 50 µL 2 M glycine (pH 10) was added into the mixture to cease the reaction after incubation for 15 min. The absorbance reading was detected at 405 nm, and the percentage of inhibition was determined similarly to the anti-α-amylase assay. Positive control used in this assay was quercetin and acarbose, and the results obtained were expressed in IC50.
2.8 1H NMR analysis and spectra processing
The experiment of 1H NMR was conducted based on the method described (Abdul-Hamid et al., 2019). Generally, 10 mg of test sample was dissolved in 375 µL of CD3OD and 375 µL of KH2PO4 buffer in D2O containing 0.1 % TSP (pH 6.0). The mixture was sonicated for 15 min and centrifuged for 10 min at 13000 rpm. The supernatant was then subjected to 1H NMR analysis using a Varian INOVA 500 MHz NMR spectrometer (Varian Inc., Palo Alto, CA, USA). The PRESAT was employed to suppress the broad water signal. The acquisition time is 4.29 min with 64 scans, spectral width and relaxation delay were reported as 16 ppm, 2.045 s, 8.6 ms, and 2.0 s, respectively. 2D-NMR experiments were performed. J-res spectrum was measured through 8 K for the chemical shift axis and spectral width of 66 Hz with a relaxation delay of 1.5 s, 8 scans per 256 increments, and spectral width of 5000 Hz for the axis of the spin–spin coupling constant. TOCSY spectrum was performed with 16 scans per 256 increments, and 1.0 s relaxation delay in 2 h and 54 min total experiment run time, while HSQC and HMBC spectra were obtained by 32 scans per 256 increments, with relaxation delays of 1.0 s, in a total experiment run time of 5 h and 33 min, and 5 h and 43 min, respectively. All the processed spectra were binned using Chenomx software (Edmonton, AB, Canada). All spectra were binned and exported into Excel file for MVDA which were performed on the binned integrals of the 1H NMR data using SIMCA-P + software version 14.1 (Umetrics, Umeå, Sweden). Principal component analysis (PCA) and partial least squares (PLS) models were generated using the Pareto scaling method (Pramai et al., 2018).
2.9 UHPLC-MS/MS analysis
The measurement was conducted using method described (Wong et al., 2020). The UHPLC-MS/MS spectra in negative ion mode were acquired from 150 to 1500 m/z with collision-induced dissociation (CID) energy of 30 % using Thermo ScientificTM Q ExactiveTM Hybrid Quadrupole-Orbitrap mass spectrometer equipped with autosampler and surveyor UHPLC binary pump coupled with electrospray ionization (ESI) source (Thermo Fisher Scientific, Bremen, Germany). The mobile phase used was LCMS-grade water (solvent A) and acetonitrile (solvent B) with each consisting of 0.1 % FA. The programmed gradient was commenced with 95 % to 0 % solvent A from 0.5 to 30 min at a flow rate of 0.4 mL/min. The phytochemical separation was determined by an Acquity UPLC HSS T3 column (1.8 µm, 2.1 x 100 mm), with a 2 µL injection volume of 10 mg/mL sample. The obtained spectra were processed and analyzed using Thermo Xcalibur Qual Browser software 4.0 (Thermo Fisher Scientific Inc., Waltham, MA, USA). The peak identification was accomplished by comparison of deprotonated molecular ion and fragmentation patterns with mass tolerance at 0.01 Da (Olsen et al., 2005).
2.10 Statistical analysis
The results of biological activities were displayed as the mean ± standard deviation of six biological replicates. The significant difference of factors between groups at 95 % confidence level was determined using One-way ANOVA with Tukey’s post hoc test. Statistical analysis was accomplished by using IBM SPSS Statistics 20 (SPSS Inc., Chicago, IL, USA) and MS Excel 2013 (Microsoft, Redmond, WA, USA).
3 Results
3.1 Effects of extraction solvent ratio on biological activities
The impacts of solvent systems on the TPC and biological activities of S. jambos leaves are presented in Table 1. The use of 50 % and 70 % ethanol was able to extract higher yield from S. jambos leaves compared with the use of water and absolute ethanol extracts. The absolute ethanol extract retained the highest TPC without statistical difference with 70 % ethanolic extract followed by 50 % ethanolic and water extract. In antioxidant activities, the results (Table 1) indicated that the IC50 values of the DPPH• activity ranged from 2.49 ± 0.26 to 7.72 ± 0.42 µg/mL. The water extract (0 %) showed the highest DPPH• radical scavenging capacity without significant difference with 70 % ethanolic extract. Meanwhile, all the IC50 values of the extracts were higher than those of gallic acid and quercetin in the NO• assays (15.41 ± 0.63 and 15.85 ± 0.58 µg/mL, respectively). The water, absolute ethanol, and 70 % ethanolic extracts exhibited IC50 values close to 42 µg/mL without significant difference. The results are expressed as mean ± standard deviation of six replicates. Means with different superscript letters are significantly different (p < 0.05). “-” indicates the activities were not measured due to the irrelevance to the compounds.
Solvent system
Yield of extraction (%)
TPC (mg GAE / g crude extract)
DPPH• scavenging assay IC50 (µg/mL)
NO• scavenging assay IC50 (µg/mL)
Anti-α-amylase assay IC50 (µg/mL)
Anti-α-glucosidase assay IC50 (µg/mL)
0 %
21.26 ± 1.34 b
323.23 ± 20.46 b
2.49 ± 0.26 a
42.72 ± 3.51b
752.97 ± 103.66 c
30.41 ± 4.23 e
50 %
24.91 ± 1.35 a
324.78 ± 19.17 b
3.18 ± 0.25 b
53.62 ± 2.79 c
88.40 ± 5.36 b
5.62 ± 0.69 c
70 %
24.89 ± 0.98 a
386.37 ± 36.48 a
2.73 ± 0.07 a
42.08 ± 3.76 b
83.43 ± 3.40 b
0.52 ± 0.01 a
Abs
14.05 ± 1.44 c
425.04 ± 42.19 a
7.72 ± 0.42 c
42.34 ± 3.62 b
89.51 ± 6.75 b
0.90 ± 0.07 b
Quercetin
–
–
3.55 ± 0.28 b
15.85 ± 0.58 a
–
6.62 ± 0.03 c
Gallic acid
–
–
–
15.41 ± 0.63 a
–
–
Acarbose
–
–
–
–
0.68 ± 0.02 a
23.83 ± 0.55 d
The results (Table 1) indicated that the 70 % ethanolic extract demonstrated the most active in both anti-α-amylase and anti-α-glucosidase assays with IC50 values of 83.43 ± 3.40 µg/mL and 0.52 ± 0.01 µg/mL, respectively. In contrast, the water extract showed the lowest activity in both the α-amylase and α-glucosidase enzyme inhibition, with IC50 values of 752.97 ± 103.66 µg/mL and 30.41 ± 4.23 µg/mL, respectively.
3.2 Metabolite identification via NMR
The presence of metabolites in S. jambos leaves extracted at different ethanol/water ratios was investigated using a combination of the 1D 1H NMR and 2D-NMR (J-res, TOCSY, HSQC and HMBC) techniques. Fig. S1 presents the representative 1H NMR spectra of S. jambos leaves of different extracts. J-res analysis (Fig. S2) was conducted to elucidate the overlapping signals and to determine the coupling constants, TOCSY (Fig. S3) explained the correlations of protons within a spin system, whereas HSQC and HMBC (Fig. S4 and S5) provide proton-carbon single bond and multiple bonds correlations (Abdul-Hamid et al., 2019). The tentative metabolite identification was based on literature data comparison and quest for several available online databases (Chenomx database, KNApSAcK, NMRDB, and Human Metabolome Database). Previously, phenolics (gallic acid and bergenin), flavonoids (myricetin, kaempferol, and myrigalone G), triterpenoids (lupeol, β-amyrin, and friedelin), and ellagitannins (castalagin, casuarinin, vescalagin, and di-hexahydroxydiphenoyl (HHDP) glucose) were reported in the genus Syzygium (Sobeh et al., 2018). In the current study, 17 compounds comprising primary and secondary metabolites were further annotated via NMR (Table 2). “+” indicating presence and “-” indicating absence of the signals in the extracts.
No
Metabolites
1H NMR characteristic signals
TOCSY correlations δ1H
HSQC correlations δ13C
HMBC correlations δ13C
0 % EtOH
50 % EtOH
70 % EtOH
Abs EtOH
Primary metabolites
1
α-Glucose
5.18 (d, J = 3.5 Hz)
–
–
–
+
+
+
+
2
β-Glucose
4.57 (d, J = 8 Hz)
–
–
–
+
+
+
+
3
Fructose
4.10 (m)
–
–
–
+
+
+
–
4
Valine
3.61 (d, J = 5.0 Hz)
–
–
–
+
+
+
+
5
Alanine
1.47 (d, J = 7 Hz)
–
–
–
+
+
+
–
6
Formic acid
8.47 (s)
–
–
–
+
–
–
–
7
Choline
3.19 (s)
–
–
–
+
+
+
+
8
Gallic acid
7.15 (s)
–
–
–
+
+
+
+
Secondary metabolites
9
Stigmasterol
5.10 (m)
1.03 (s)–
–
–
+
+
+
+
10
Lupeol
4.54 (s)
1.07(s)
0.94 (s)–
–
–
–
+
+
+
11
Syringic acid
7.33 (s)
3.82 (s)–
–
–
+
+
+
+
12
β-amyrin
2.03 (s)
1.04 (s)
0.86 (s)–
–
–
+
+
+
+
13
Bergenin derivatives
4.06 (s)
4.02 (dd, J = 1.5, 12.0 Hz)
3.81 (m)
3.78 (m)–
74.53 (C8)
–
+
+
+
+
14
Myricetin derivatives
7.05 (s)
6.31 (d, J = 2.0 Hz)–
–
–
+
+
+
+
15
Kaempferol derivatives
8.04 (d, J = 8.7 Hz)
6.74 (d, J = 1.9 Hz)
6.42 (d, J = 1.9 Hz)
6.29 (d, J = 2.0 Hz)6.45 (H8)
97.23 (C8)102.02
(C6)–
+
+
+
+
16
Friedelin
1.57 (m)
1.31 (m)
1.02 (s)
0.92 (d, J = 6 Hz)
0.88 (s)
0.74 (s)1.57 (H18)
18.73 (C26/27)
–
–
+
+
+
17
Stigmasterol glucoside
4.51 (d, J = 8 Hz)
3.72 (m)
3.58 (m)
2.03 (m)
1.35 (d, J = 7.0 Hz)
1.06 (t, J = 7.2 Hz)
0.93 (d, J = 7.0 Hz)–
19.38 (C35)
–
–
+
+
+
18
Myrigalone G
7.18 (m)
3.80 (s)
3.34 (t, J = 8.0 Hz)
2.94 (t, J = 8.0 Hz)
1.90 (s)3.34 (Hα)
30.33 (Cβ)
–
–
+
+
+
19
Galloyl castalagin
7.14 (s)
6.78 (s)
6.58 (s)–
–
112.66 (C21), 141.77 (C23), 148.16 (C20)
–
+
+
+
20
Casuarinin
6.75 (s)
6.48 (s)
5.60 (d, J = 5.0 Hz)
5.30 (d, J = 7.0 Hz)–
–
–
+
+
+
+
21
Tellimagrandin II
7.16 (s)
7.11 (s)
7.01 (s)
3.80 (d, J = 14 Hz)–
112.79 (C3′)112.73
(C3)147.99 (C6),170.70
(C7)–
+
+
+
22
Coriariin B
7.35 (d, J = 2.0 Hz)
7.23 (d, J = 2.0 Hz)
7.00 (s)
6.98 (s)
6.65 (s)
6.49 (s)–
–
–
+
+
+
+
23
Coriariin A
6.70 (d, J = 2.0 Hz)
6.98 (s)
6.68 (s)
6.65 (s)–
–
–
+
+
+
+
24
Praecoxin A
7.13 (s)
6.58 (s)
6.49 (s)
3.71 (d, J = 13 Hz)–
110.90 (C3)110.03
(C6′’)–
+
+
+
+
25
Praecoxin B
7.11 (s)
6.39 (s)
5.18 (d, J = 3.5 Hz)–
–
–
+
+
+
+
26
Vescalagin
6.96 (s)
6.79 (s)
4.51 (d, J = 8 Hz)
4.18 (s)–
112.75 (C3′)112.70
(C3)
170.19 (C7),147.83 (C1)
,118.31
(C4)+
+
+
+
27
Vescalin
6.79 (s)
5.32 (s)
3.95 (t, J = 7 Hz)
3.88 (dd, J = 3, 12 Hz)–
–
–
+
+
+
+
28
Trigalloyl glucose
5.63 (d, J = 7.7 Hz)
5.26 (d, J = 7.7 Hz)
5.03 (m)
3.97 (d, J = 12.5)–
–
–
+
+
+
+
29
Di-HHDP glucose
7.49 (s)
6.65 (s)
5.34 (s)
5.18 (d, J = 3.5 Hz)
4.58 (d, J = 8 Hz)
3.70 (d, J = 12 Hz)
3.51
(d, J = 12 Hz)–
112.78 (C3′)
172.88 (C7)
+
+
+
+
30
Pomolic acid
3.46 (dd, J = 4.0, 11.0 Hz)
2.74 (s)
1.90 (s)
1.70 (m)
1.45 (m)
1.14 (d, J = 7.0 Hz)
1.07 (s)
0.92 (s)1.90 (H22)1.45
(H4)52.13 (C18‘)
–
–
+
+
+
3.3 Discrimination of S. jambos leaf extracts
The generated principal component analysis (PCA) score plot investigates the impact of extraction solvent polarity on clustering characteristics. Besides, a corresponding loading plot (Fig. 1b) was applied to determine the metabolite variation among S. jambos leaf extracts. The PCA score plot (Fig. 1a) showed the two major PCs contributed to a total variance of 80.3 %, with PC1 (67.1 %) and PC2 (13.2 %). Based on the plot, S. jambos leaves extracted with different solvent systems were divided into four distinct clusters without visible outliers. PC1 separated the absolute ethanol extract from hydroethanolic and water extracts, while PC2 discriminated the 50 % and 70 % ethanolic extracts from the absolute and water extracts. Meanwhile, the loading plot (Fig. 1b) demonstrated that the hydroethanolic extracts shared similarities in metabolite retention and hence were closely clustered.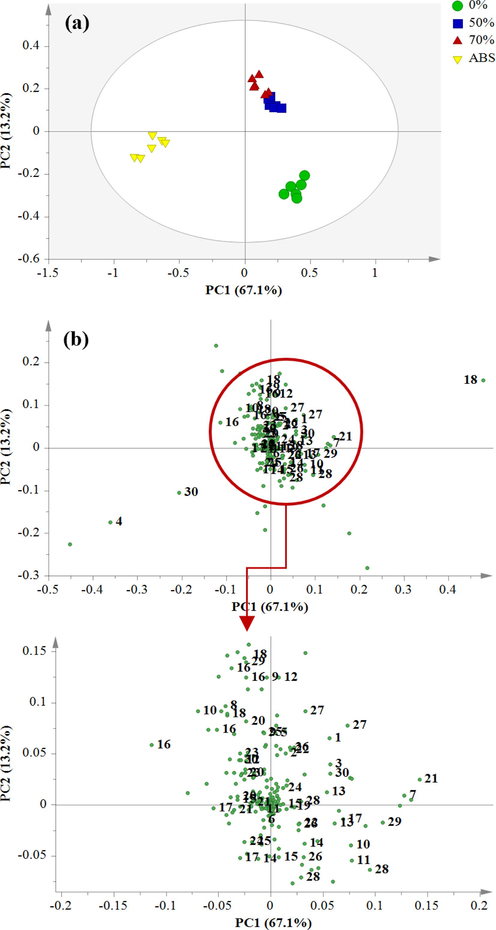
(a) Principal component analysis (PCA) score plot and (b) Loading plot of S. jambos leaves extracted with 0%, 50%, 70% and absolute ethanol.
3.4 Metabolite correlation with biological activities
Partial least square (PLS) analysis was employed to further evaluate the relationship between biological activities and the identified metabolites of the S. jambos leaf extracts. The PLS biplot (Fig. 2) is a composition of both score and loading plots in a single graphical presentation (Abdul-Hamid et al., 2015), To avoid bias analysis, two separate PLS biplots were generated to investigate the correlation between the X (binned 1H NMR spectral data) variables and Y (DPPH• and NO• radical scavenging, anti-α-amylase, and anti-α-glucosidase assays) variables for the inverse position of the Y variables in the plot.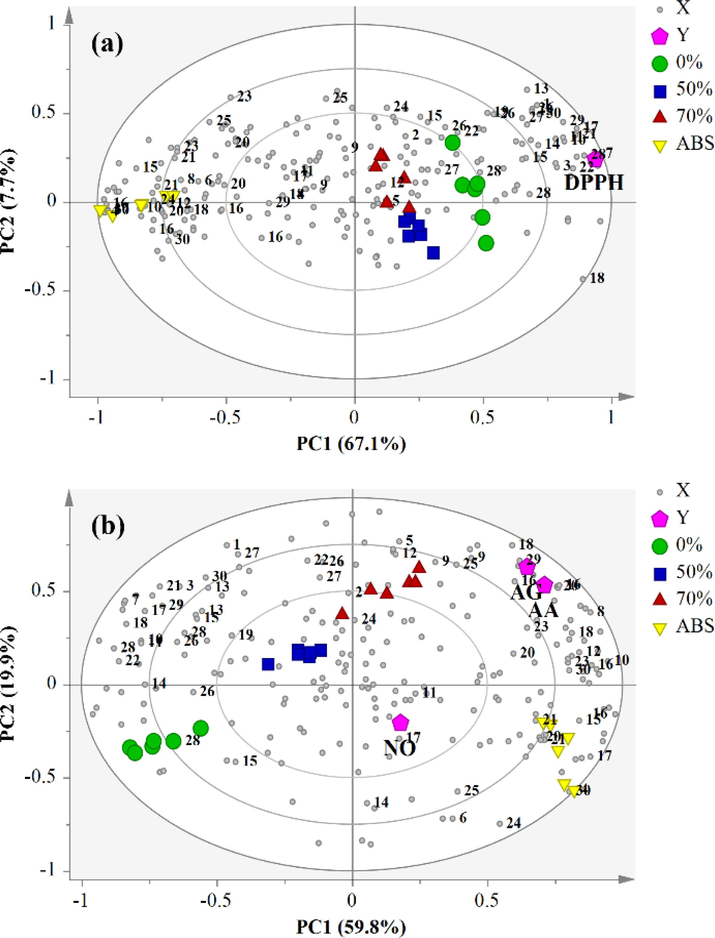
Partial least square (PLS) biplot illustrating the correlation of metabolites in S. jambos leaves with (a) DPPH• activity, (b) NO•, anti-α-amylase and anti-α-glucosidase activities; AA: anti-α-amylase, AG: anti-α-glucosidase.
3.5 Relative quantification of bioactive compounds
13 shared metabolites of both biplots that are strongly contributed to the bioactivities (VIP value > 1.0) were relatively quantified to the concentration of TSP according to the respective binned 1H NMR spectral data (Fig. 3 and Table S1).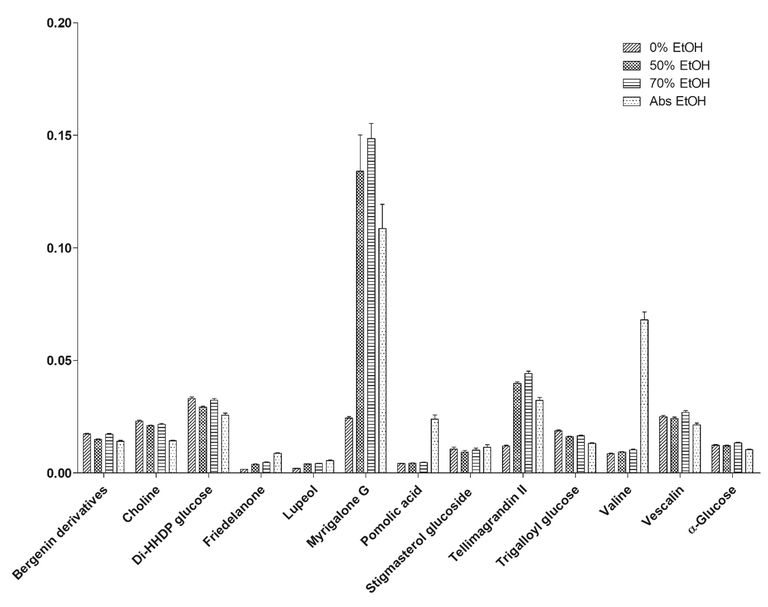
Relative quantification of the identified metabolites in S. jambos leaves extract.
3.6 Metabolite profiling via UHPLC-MS/MS
The most active extract, i.e., the 70 % ethanolic S. jambos leaf extract, was further characterized for its phytochemicals via ultrahigh-performance liquid chromatography–tandem mass spectrometry (UHPLC-MS/MS). It is worth mentioning that a number of 59 metabolites were conditionally identified in the S. jambos leaf extract (Table 3). The annotation was accomplished on literature data comparison and quest for various online databases (MassBank, PubChem, Metabolomics Workbench, KNApSAcK, and HMDB). The total ion chromatogram demonstrated the extensive compounds discovered between 0 and 30 min (Fig. S6). Tannins and flavonoids accounted for 34 % and 24 % of the 59 identified metabolites, respectively. Other classes of identified metabolites include terpenoids, phenols, and fatty acids.
Peak no.
Retention time, min
Exact mass
Deprotonated molecular ion [M−H]- (m/z)
Delta
MS/MS fragment ions
Tentative identification
Molecular formula
References
31
0.64
504.1618
503.1611
0.0007
221.0659, 179.0549, 161.0443, 113.0230, 101.0230
Raffinose
C18H32O16
(Kubica et al., 2012)
32
0.72
484.0780
483.0775
0.0005
331.0666, 313.0343, 300.9986, 169.0129
Digalloyl glucose
C20H20O14
(Slatnar et al., 2015)
33
0.99
192.0197
191.0187
0.0010
173.0079, 129.0180, 111.0074, 87.0073, 85.0281
Citric acid
C6H8O7
(Lantzouraki et al., 2015)
34
1.02
484.0780
483.0775
0.0005
331.0666, 313.0343, 300.9985, 169.0130
Digalloyl glucose isomer 1
C20H20O14
(Slatnar et al., 2015)
35
1.61
484.0780
483.0776
0.0004
331.0666, 313.0343, 300.9985, 169.0131
Digalloyl glucose isomer 2
C20H20O14
(Slatnar et al., 2015)
36
1.74
484.0780
483.0776
0.0004
331.0666, 313.0343, 300.9986, 169.0129
Digalloyl glucose isomer 3
C20H20O14
(Slatnar et al., 2015)
29
2.21
784.0681
783.0674
0.0007
481.0626, 300.9984, 275.0193, 257.0088, 229.0136
Di-HHDP glucose
C34H24O22
(Plaza et al., 2016)
26
2.29
934.0640
933.0623
0.0017
915.0509, 631.0547, 467.0239, 451.0322, 300.9985
Vescalagin
C41H26O26
(Sobeh et al., 2018)
37
2.55
784.0681
783.0673
0.0008
481.0626, 300.9984, 275.0193, 257.0088, 229.0136
Di-HHDP glucose isomer 1
C34H24O22
(Plaza et al., 2016)
38
2.86
934.0640
933.0631
0.0009
631.0565, 467.0230, 451.0292, 300.9985
Vescalagin isomer
C41H26O26
(Sobeh et al., 2018)
39
3.06
634.0734
633.0724
0.0010
463.0522, 300.9985, 275.0193
Strictinin
C27H22O18
(Taamalli et al., 2013)
40
3.11
634.0734
633.0724
0.0010
463.0508, 300.9985, 275.0193
Strictinin isomer 1
C27H22O18
(Taamalli et al., 2013)
41
3.34
784.0681
783.0673
0.0008
481.0597, 300.9986, 275.0194, 257.0089, 229.0135
Di-HHDP glucose isomer 2
C34H24O22
(Plaza et al., 2016)
42
3.39
784.0681
783.0670
0.0011
481.0597, 300.9986, 275.0194, 257.0083, 229.0135
Di-HHDP glucose isomer 3
C34H24O22
(Plaza et al., 2016)
43
3.78
484.0780
483.0773
0.0007
331.0666, 313.0561, 300.9980, 169.0131
Digalloyl glucose isomer 4
C20H20O14
(Slatnar et al., 2015)
44
3.88
1068.1213
1067.1205
0.0008
533.0539, 377.0291, 300.9987, 249.0396
Pterocarinin A
C46H36O30
(Díaz-de-Cerio et al., 2016)
45
3.93
484.0780
483.0772
0.0008
331.0666, 313.0561, 300.9988, 169.0131
Digalloyl glucose isomer 5
C20H20O14
(Slatnar et al., 2015)
46
4.12
634.0734
633.0725
0.0009
463.0503, 300.9986, 275.0194
Strictinin isomer 2
C27H22O18
(Taamalli et al., 2013)
20
4.29
936.0796
935.0798
0.0002
783.0668, 633.0737, 481.0637, 300.9991, 275.0195, 169.0131
Casuarinin
C41H28O26
(Díaz-de-Cerio et al., 2016)
47
4.56
786.0843
785.0833
0.0010
633.0725, 483.0795, 300.9985, 275.0193, 169.0132
HHDP-digalloylglucose (Tellimagrandin I)
C34H26O22
(Plaza et al., 2016)
48
4.70
306.0667
305.0696
0.0029
286.9944, 269.9857, 108.0201
(-)-Epigallocatechin
C15H14O7
(Savić et al., 2014)
49
5.00
478.0675
477.0663
0.0012
300.9986, 257.0090, 229.0137, 151.0027
Quercetin glucuronide
C21H18O13
(Lee et al., 2019)
50
5.10
464.0591
463.0581
0.0010
300.9982, 257.0091, 229.0127, 185.0236
Ellagic acid hexoside
C20H16O13
(Slatnar et al., 2015)
51
5.45
936.0796
935.0779
0.0017
633.0726, 481.0623, 300.9985, 275.0193, 169.0130
Casuarinin isomer (Casuarictin)
C41H28O26
(Díaz-de-Cerio et al., 2016)
52
5.70
480.0831
479.0823
0.0008
317.0269, 316.0220, 271.0242
Myricetin glucoside
C21H20O13
(Lee et al., 2019)
53
5.79
480.0831
479.0823
0.0008
317.0269, 316.0220, 271.0242
Myricetin glucoside isomer
C21H20O13
(Lee et al., 2019)
21
5.85
938.0953
937.0940
0.0013
785.0770, 635.0892, 483.0781, 300.9985
Eugeniin / Tellimagrandin II
C41H30O26
(Grace et al., 2014)
54
5.95
434.0776
433.0774
0.0002
300.9980, 283.9954, 271.9956
Quercetin xyloside
C20H18O11
(Lee et al., 2019)
55
6.10
616.0992
615.0989
0.0003
463.0880, 300.0273, 271.0237, 255.0295, 178.9976, 169.0132
Quercetin galloylglucoside
C28H24O16
(Lee et al., 2019)
56
6.19
626.1410
625.1406
0.0004
316.0220, 287.0195, 271.0244, 178.9977, 151.0023
Myricetin robinobioside
C27H30O17
(Lee et al., 2019)
57
6.42
301.9990
300.9981
0.0009
283.9954, 257.0089, 229.0135, 185.0235
Ellagic acid
C14H6O8
(Regueiro et al., 2014)
58
6.55
596.1305
595.1300
0.0005
316.0219, 287.0195, 271.0243, 178.9973, 151.0024
Myricetin xylosyl-rhamnoside
C26H28O16
(Lee et al., 2019)
59
6.64
464.0882
463.0875
0.0007
300.0272, 271.0244, 255.0299, 178.9974, 151.0025
Quercetin glucoside
C21H20O12
(Lee et al., 2019)
60
7.00
506.1352
505.1351
0.0001
359.1493, 344.1259, 329.1021, 300.0955
Jaceidin rhamnoside
C24H26O12
(Taamalli et al., 2013)
61
7.08
434.0776
433.0774
0.0002
300.0271, 271.0245, 255.0296, 178.9992, 151.0028
Quercetin arabinoside
C20H18O11
(Lee et al., 2019)
62
7.21
448.0933
447.0928
0.0005
301.9967, 300.9941, 270.9876, 255.5956
Quercetin rhamnoside
C21H20O11
(Lee et al., 2019)
63
7.41
580.1356
579.1348
0.0008
429.0834, 300.0272, 271.0245, 255.0295
Quercetin xylosyl-rhamnoside
C26H28O15
(Lee et al., 2019)
64
8.15
564.1406
563.1401
0.0005
413.0867, 285.0372, 255.0294, 227.0342, 178.9976, 151.0024
Kaempferol rhamnoside-xyloside
C26H28O14
(Lee et al., 2019)
65
8.87
478.1002
477.1031
0.0029
331.0663, 313.0565, 169.0131
Gallic acid coumaroyl-hexose
C22H22O12
(Sobeh et al., 2018)
66
11.18
328.2177
327.2170
0.0007
229.1440, 211.1331, 171.1015
Trihydroxy octadecadienoic acid
C18H32O5
(Taamalli et al., 2013)
67
11.96
330.2334
329.2327
0.0007
312.9938, 293.2107, 242.3922, 229.1438, 211.1331, 183.1379
Trihydroxy octadecenoic acid
C18H34O5
(Taamalli et al., 2013)
68
12.47
504.3378
503.3371
0.0007
485.3265, 453.3004, 427.3229, 407.2961, 370.8087, 174.9547
Madecassic acid
C30H48O6
(Xia et al., 2015)
69
13.20
504.3378
503.3373
0.0005
485.3270, 453.3011, 407.2961, 369.7016
Madecassic acid isomer
C30H48O6
(Xia et al., 2015)
70
15.29
488.3429
487.3415
0.0014
419.2439, 409.3099, 377.0679, 373.9642, 151.4273
Asiatic acid
C30H48O5
(Xia et al., 2015)
71
18.11
ND
723.3798
ND
695.0396, 415.1442, 279.2323
Fatty acid derivative
ND
(Minkler & Hoppel, 2010)
72
18.26
ND
476.2773
ND
416.2551, 279.2325, 214.0470, 196.0373
Fatty acid derivative
ND
(Hsu et al., 2007)
73
18.39
ND
564.3293
ND
505.3116, 279.2323, 481.8533
Fatty acid derivative
ND
(Hsu et al., 2007)
74
18.62
ND
559.3112
ND
496.2796, 412.5738, 277.2167
Fatty acid derivative
ND
(Hsu et al., 2007)
75
18.88
ND
699.3796
ND
437.1535, 415.1462, 255.2323, 172.3546
Fatty acid derivative
ND
(Hsu et al., 2007)
76
19.00
ND
452.2773
ND
255.2325, 214.0476, 196.0372
Fatty acid derivative
ND
(Hsu et al., 2007)
77
19.16
ND
540.3295
ND
436.6639, 417.5592, 255.2324
Fatty acid derivative
ND
(Hsu et al., 2007)
78
19.46
ND
431.2196
ND
294.7976, 277.2172, 171.0053
Fatty acid derivative
ND
(Hsu et al., 2007)
30
19.92
472.3480
471.3469
0.0011
454.1732, 427.1954, 393.3149, 246.9636, 201.9723
Pomolic acid
C30H48O4
(Guo et al., 2011)
79
20.15
472.3480
471.3470
0.0010
454.1732, 427.1954, 264.9446, 246.9636, 146.4775
Maslinic acid
C30H48O4
(Cheng & Cao, 1992)
80
23.19
456.3531
455.3525
0.0006
406.6367, 378.5817, 327.2702, 231.1748
Oleanolic acid
C30H48O3
(Song et al., 2006)
81
26.15
592.2613
591.2598
0.0015
559.2346, 515.2444, 500.2210, 487.2533, 460.2217
Pheophorbide A
C35H36N4O5
(Park et al., 2003)
82
26.39
356.3218
355.3209
0.0009
338.3106, 309.3160, 295.2102, 280.1995, 254.8060, 238.9417, 102.1235
Hydroxy-docosanoic acid
C22H44O3
(Muchembled et al., 2005)
83
26.85
370.3447
369.3410
0.0037
353.1471, 325.2532, 309.2224
Methyl hydroxydocosanoate
C23H46O3
(Gorst-Allman & Spiteller, 1988)
84
27.01
370.3447
369.3410
0.0037
353.1471, 325.2532, 309.2224
Methyl hydroxydocosanoate isomer
C23H46O3
(Gorst-Allman & Spiteller, 1988)
4 Discussion
The present study evaluated the bioactive phytoconstituents in S. jambos leaves exhibiting antidiabetic properties through in-vitro anti-hyperglycemic activities by providing a comprehensive profile using 1H NMR metabolomics and UHPLC-MS/MS techniques. Four different solvent extraction systems were investigated on the retention of potential bioactive metabolites from the plant matrix. Ultra-sonication is a non-thermal extraction technique used in this study as it facilitates effective extraction of phytoconstituents from plant matrix in a shorter duration, requires less solvent, produces higher yield and better retention of the bioactive compounds (Kumar et al., 2021). The higher yield of hydroethanolic S. jambos extracts may be attributed to the combination of solvents used in the extraction, which facilitated the solubility of both polar and less polar compounds from the plant matrices (Abdul-Hamid et al., 2015). The nonalignment of the TPC with the extraction yield might be attributed to the higher amount of non-phenol compounds such as terpenes and the carbohydrates being extracted in more polar solvents. In addition, complex formation of phenolic compounds can be possibly found in the extract, and these compounds may have more phenol groups than the more polar extracts (Do et al., 2014). Oxidative stress has been postulated in chronic diseases including diabetes (Lobo et al., 2010), thus, two in-vitro antioxidant activities, specifically 2,2-diphenyl-1-picrylhydrazyl (DPPH•) and nitric oxide (NO•), radical scavenging activities were conducted to evaluate the antioxidant potential of S. jambos leaf extracts. Interestingly, a different trend was observed in both antioxidant assays. This may be due to the different antioxidant mechanisms (Sumanont et al., 2004). In the DPPH• assay, antioxidants normally act by donating electron or hydrogen atoms to scavenge DPPH• free radicals, whereas the NO• radicals could be scavenged by receiving hydrogen atom from antioxidant or the formation of antioxidant cation with successive loss of hydrogen atom (Sumanont et al., 2004). Thus, this study suggested that the antioxidant potential of S. jambos leaf extracts is higher in DPPH• free radicals than in NO• free radicals.
Furthermore, the inhibition of two carbohydrate hydrolyzing enzyme activities, particularly anti-α-amylase and anti-α-glucosidase assays, was evaluated on the anti-hyperglycemic properties of S. jambos leaf extracts. A slightly different trend was discovered in the anti-α-amylase and anti-α-glucosidase activities, even though the most active extract is 70 % ethanolic extract and the water extract is the least active in both cases. This may be attributed to the distinct principle of the enzymatic reaction. During normal carbohydrate ingestion, the complex starch structure is first hydrolyzed by α-amylase enzyme, which is further hydrolyzed by membrane-bound intestinal α-glucosidase enzyme to glucose and other monosaccharides for absorption (Alqahtani et al., 2020). In the anti-α-amylase assay, the DNS reagent detected and estimated the amount of reducing sugar as the product of enzymatic reaction, whereas in the anti-α-glucosidase assay, the potential to prevent enzyme from hydrolyzing the substrate for glucose production was evaluated (Miller, 1959). Thus, the inhibition of both α-amylase and α-glucosidase enzyme assays could suppress carbohydrate digestion, which could delay glucose uptake and reduce the blood glucose level. The positive results from in-vitro antioxidant and anti-hyperglycemic activities might justify the ethnobotanical use of S. jambos leaves in indigenous communities to treat diabetes and inflammatory-associated diseases. Furthermore, the in-vitro outcomes supported research on in-vivo models demonstrated antidiabetic properties of S. jambos bark and leaves in Type I diabetic model by protecting pancreatic β-cells against oxidative stress and enhancing insulin signaling pathway (Mahmoud et al., 2021). In view of this bioactive medicinal plant, S. jambos is considered harmless for consumption as its safety dose is up to 5 g/kg body weight in acute toxicity rat model evaluation (Dhanabalan et al., 2014). Different extracts and fractions of the leaves exhibited various toxicity levels on brine shrimp Artemia salina assay (Ochieng et al., 2022). However, there is limited information on the toxicological data of this plant to date which is yet to be fully explored.
The highly overlapping signals in the sugar region (δ 3.10–5.50) of the NMR spectra complicate the metabolite identification. The water extract exhibited a higher peak intensity in this region compared with the ethanolic extract, with signals at δH 5.18 (d, J = 3.5 Hz) and δH 4.57 (d, J = 8 Hz) could be annotated as α-glucose (1) and β-glucose (2), respectively. In the aliphatic region (δ 0.50–3.00), the water extract generated lesser peaks with lower concentration, interpreting the absence of lupeol (10), friedelin (16), stigmasterol glucoside (17), myrigalone G (18) and pomolic acid (30). In addition, the water extract did not increase galloyl castalagin (19) and tellimagrandin II (21) in the aromatic region (δH 5.00–8.50). Meanwhile, the primary metabolites were absent in the absolute ethanol extract including fructose (3) and alanine (5) (Khoo et al., 2015).
Based on the TOCSY spectrum, identification of kaempferol derivatives (15) was further supported by the correlation via spin–spin coupling at resonating signals at δH 6.29 and 6.42 in position H6 and H8 respectively in the aromatic A-ring of the structure. HSQC experiment also showed correlations of H6 to C6 at δc 102.02, and H8 to C8 at δc 97.23, indicating direct attachment of hydrogen atoms to carbon atoms in the compound. Besides, bergenin derivatives (13) could be characterized by a direct correlation of H8 (δH 4.02) to C8 at δc 74.53 by HSQC analysis. Besides, myrigalone G (18) could be further identified with TOCSY correlations of Hβ and Hα resonating at δH 2.94 and 3.34 respectively, where Hβ (δH 2.94) showed a direct correlation of HSQC experiment at δC 30.33 (Hβ-Cβ) (Jayasinghe et al., 2007).
Several ellagitannins could be further characterized using 2D-NMR experiments. HMBC analysis supported the structure of galloyl castalagin (19) by 3 J correlation between H21 (δH 7.14) of the galloyl moiety with C21 (δC 112.66), C23 (δC 141.77) and C20 (δC 148.16) of the aromatic ring in HHDP group (Yang et al., 2000). Galloyl moiety was anticipated attached to castalagin relatively higher compared to its isomer vescalagin, as galloyl castalagin was previously identified in S. jambos leaves (Ochieng et al., 2022). As for tellimagrandin II (21), proton at δH 7.11 (H3) and δH 7.18 (H3′) showed a direct correlation in the HSQC experiment at δC 112.73 (H3-C3) and δC 112.79 (H3′-C3′) of the HHDP moiety, while HMBC showed H3 correlate to C6 (δC 147.99) and C7 (δC 170.70). Furthermore, HSQC showed the direct correlation of proton H6′’ at δH 6.49 to C6′’ (δC 110.03) of the valoneoyl group, and H3 (δH 6.58) to C3 (δC 110.90) of HHDP moiety in praecoxin A (24). Vescalagin (26) showed HSQC correlation between H3 (δH 6.79) and C3 at δC 112.70, while the correlation of H3 to C1 (δC 147.83), C4 (δC 118.31) and C7 (δC 170.19) were demonstrated in HMBC of D-ring and its carbonyl group of the structure. HSQC correlation demonstrates a direct correlation of H3′ (δH 6.96) to C3′ (δC 112.75) in the E-ring of the structure (Puech et al., 1999). The HHDP hydrogen in metabolite 29 showed a direct HSQC correlation of H3′ at δH 7.49 to C3′ at δC 112.78 of the aromatic ring, while a 3 J correlation of H3 (δH 6.65) to carbonyl carbon C7 at δC 172.88 could be assigned with HMBC experiment.
Identification of triterpenoids such as pomolic acid (30) was further strengthened by the TOCSY correlation of protons resonating at H16 (δH 1.70) and H22 (δH 1.90), H2 (δH 0.92), and H24 (δH 1.45), while a direct correlation of H18 (δH 2.74) to C18 at δC 52.13 could be assigned with HSQC experiment. Besides, identification of friedelin (16) was identified based on the hydrogen–hydrogen correlation in a spin system at δH 1.31 (H14) and δH 1.57 (H18), while a direct HSQC correlation could be assigned on H26/27 at δH 1.02 to C26/27 at δC 18.73 (Manguro et al., 2018).
Based on the identified metabolites, the discrimination of S. jambos leaves extracted by different solvent systems was manifested by the PCA score and loading plots, where less polar compounds, such as valine (4), friedelin (16), and pomolic acid (30), were more predominant in the absolute ethanol extract, and these metabolites positioned on the negative side of PC1. The water extract retained a higher amount of syringic acid (11), trigalloyl glucose (28), and di-HHDP glucose (29). Furthermore, the loading plot demonstrated that the hydroethanolic extracts had higher extraction of most metabolites, including myrigalone G (18), galloyl castalagin (19), casuarinin (20), tellimagrandin II (21), praecoxin A (24), praecoxin B (25), vescalagin (26), vescalin (27), and β-amyrin (12). This may be attributed to the use of a solvent mixture that is more effective in the dissolution of metabolites from plant matrices and possibly contribute to biological activities (Abdul-Hamid et al., 2015). The plot was validated with five evaluated principal components (PCs), with R2X (cum) = 0.949 and Q2 (cum) = 0.863. The difference between Q2 and R2X is less than 0.3 (0.086), indicating that the generated model is reliable and gained a strong predictive power (Khoo et al., 2015).
Biplot (a) (Fig. 2) shows the correlation of the metabolites with DPPH• radical scavenging activity, affording PC1 (67.1 %) and PC2 (7.7 %) of the total variance. The biplot showed that PC1 clearly separates the absolute ethanol extract from the hydroethanolic and water extracts, while the water extract is closely associated to the DPPH• activity. The plot was aligned with the bioactivities result in the current study, in which the water extract exhibited the highest DPPH• radical scavenging activity, and the lowest was absolute ethanol extract in the particular case. Meanwhile, the metabolites contributing to the DPPH• activity with variable importance in the projection (VIP) values higher than 1.0 including mostly flavonoids and some tannins such as syringic acid (11), bergenin (13), myricetin (14), and kaempferol (15) derivatives, myrigalone G (18), tellimagrandin II (21), coriariin B (22), vescalagin (26), vescalin (27), trigalloyl glucose (28), di-HHDP glucose (29), and several primary metabolites, including α-glucose (1), fructose (3), valine (4), and choline (7). The presence of free and glycosylated flavonoids in the extracts could strengthen antioxidant activity (Pereira et al., 2018). In contrast, the contradiction of high phenolic content with low DPPH• antioxidant power of the absolute ethanol extract in the present study might be explained by the chemical structure of phenolic compounds, including the number and position of hydroxyl groups that possibly influence the antioxidant capacity of the leaf extracts (Rice-Evans et al., 1996).
Another PLS biplot (b) (Fig. 2) was generated to examine the correlation of the metabolites with NO• radical scavenging, anti-α-amylase, and anti-α-glucosidase activities, affording PC1 (59.8 %) and PC2 (19.9 %) of the total variance. The biplot showed that PC1 separated the 70 % ethanolic and absolute ethanol extracts from the 50 % ethanolic and water extracts, while PC2 discriminated the hydroethanolic extracts from the water and absolute ethanol extracts. The 70 % ethanolic extract was found to be closely correlated with the biological activities, which is aligned with the bioactivities result in the current study. Furthermore, metabolite signals with VIP values > 1.0 indicate potential X variables granted to this model. For instance, most of the tannins, including myrigalone G (18), tellimagrandin II (21), coriariin A (23), praecoxin A (24), praecoxin B (25), vescalin (27), trigalloyl glucose (28), and di-HHDP glucose (29), contribute to the biological activities. Tannins isolated in the genus Syzygium have been previously reported to exhibit antioxidant and anti-hyperglycemic properties that strengthen the current PLS biplot (Gavillán-suárez et al., 2015; Sobeh et al., 2018). Several phenolic compounds, including bergenin derivatives (13) and gallic acid (8), also contributed to the activities.
Interestingly, a total of 13 metabolites shared between both biplots contributed significantly (VIP value > 1.0) to the antioxidant and anti-hyperglycemic activities of the S. jambos leaf extracts. Among these bioactive compounds, five tannins (di-HHDP glucose (29), myrigalone G (18), tellimagrandin II (21), trigalloyl glucose (28), and vescalin (27)) and three triterpenoids (pomolic acid (30), friedelin (16), and lupeol (10)) were found to be highly associated with the bioactivities. Moreover, bergenin derivatives (13), stigmasterol glucoside (17), and several primary metabolites, including choline (7), valine (4), and α-glucose (1), also strongly correlated with the bioactivities. Valine is known to be involved in the shikimate pathway for the biosynthesis of phenolic acids, which explains the significant contribution to the activity as observed in the model (Pramai et al., 2018). These significantly contributing metabolites in S. jambos leaves to antioxidant and anti-hyperglycemic activities revealed by PLS analysis substantially justified the traditional use of S. jambos leaves for treating diabetes and various inflammatory-associated illnesses.
Both PLS models were verified using the internal cross-validation technique. Two PCs with R2Y (cum) = 0.935 and Q2 = 0.901 and five PCs with R2Y (cum) = 0.98 and Q2 = 0.913 were evaluated in biplots (a) and (b), respectively. This indicated that the models were statistically valid. Furthermore, the permutation test (Fig. S7) was employed to further validate the models in order to explain the overfitting degree and predictive ability. Both models obtained Y-intercepts of R2 (<0.4) and Q2 (<0.05), indicating the robustness and reliability of the PLS models (Eriksson et al., 2006). In addition, the models were also validated by regression analysis, with the r values of DPPH•, NO•, anti-α-amylase, and anti-α-glucosidase assays being 0.9351, 0.6886, 0.9305, and 0.9817, respectively (Fig. S8). Overall, the PLS models were statistically validated and are thus reliable.
The bioactive metabolites were relatively quantified where the water extract retained a higher amount of bergenin derivatives, choline, and several tannins, including di-HHDP glucose and vescalin, without significant difference with 70 % ethanolic extract. Meanwhile, the 70 % ethanolic extract contained a significantly higher concentration of myrigalone G, tellimagrandin II, and α-glucose compared with the other extracts. Less polar compounds, such as friedelin, lupeol, pomolic acid, and valine, are significantly predominant in the absolute ethanol extract. It is important to note that both hydroethanolic extracts contained most of the metabolites, such as choline, lupeol, myrigalone G, trigalloyl glucose, and valine without significant difference. These findings agree with those of the PCA and PLS analysis in the present study, in which the solubilization of bioactive compounds from S. jambos leaves was influenced by the polarity of the solvent used in the extraction.
In view of the contribution of S. jambos leaves’ bioactive phytochemicals to the antioxidants and anti-hyperglycemic activities, a higher sensitivity of the MS technique was applied to detect low-abundance metabolites that might appear to be invisible and ambiguous in NMR due to its shortcoming. Hydrolyzable tannins including ellagitannins and gallotannins are major polyphenolic compounds identified in the S. jambos leaf extract. Gallotannins comprised of polyesters of sugar moiety and gallic acid; where ellagitannins contained hexahydroxydiphenic acid (HHDP) ester which metabolically derived from gallotannins (Plaza et al., 2016). The radicals scavenging capacity of hydrolyzable tannins depends on molecular size, as a higher number of galloyl and HHDP groups in the molecule enhanced radicals scavenging activities by the formation of stable quinone (Valverde Malaver et al., 2019). Several hydrolyzable ellagitannins identified including strictinin and eugeniin were previously identified in the genus Syzygium, which could be annotated in this study. Strictinin (39) was identified based on fragment ions at m/z 463 and 300, which correspond to the cleavage of gallic acid (169 u) and glucosyl moiety (162 u) from the deprotonated molecular ion at m/z 633.0724 (Taamalli et al., 2013). Eugeniin or tellimagrandin II (21) was identified based on the deprotonated molecular ion at m/z 937.0940 with fragment ions at m/z 785, 635, and 483 due to the subsequent loss of three galloyl moieties (152 u) and 300, which is consistent with a previously reported MS/MS data (Grace et al., 2014). Furthermore, both tellimagrandin I (47) and casuarinin (20) could be observed with the loss of subsequent galloyl moieties (152 u) to yield fragment ions at m/z 783, 633, and 483 from deprotonated molecular ions at m/z 785.0833 and 935.0798, respectively (Díaz-de-Cerio et al., 2016). Gallotannins, such as digalloyl glucose isomers (32, 34, 35, 36, 43 and 45), were tentatively identified based on deprotonated molecular ion at m/z 483.0775 and fragment ion at m/z 331, which correspond to a galloyl glucose molecule, 313 and 169, based on the subsequent loss of hydroxyl and glucose moieties, respectively (Slatnar et al., 2015). Moreover, ellagitannins, such as di-HHDP glucose and its isomers (29, 37, 41, 42), could be identified based on deprotonated molecular ion at m/z 783.0673 and fragmentation patterns at m/z 481 (loss of HHDP), 300 (elimination of HHDP-glucose), and the corresponding fragments to the HHDP unit at m/z 275, 257, and 229, which the experimental fragment pattern is in agreement with previous study (Plaza et al., 2016). Other polyphenols, such as citric acid (33), ellagic acid (57), and its derivative (50), could be detected in the S. jambos leaf extract (Lantzouraki et al., 2015; Regueiro et al., 2014).
Apart from hydrolyzable tannins, flavonoids were identified as another major class of compounds in S. jambos leaf extract. Hydroxyl configuration of the B ring of flavonoids is the most significant factor in radicals scavenging capacity as it provides a greater stability for flavonoid radical formation. Furthermore, antioxidant activities were influenced by the occurrence and number of glycoside moieties in flavonoids (Kumar & Pandey, 2013). A total of seven quercetin derivatives were relatively annotated in the S. jambos leaf extract, based on the characteristic aglycone fragment ion at m/z 301. Quercetin glucuronide (49) could be identified based on deprotonated molecular ion at m/z 477.0663 and fragment ions at m/z 300 due to the loss of glucuronide moiety (177 u). Peaks 59, 61, 62, and 63 were assigned as quercetin glucoside, quercetin arabinoside, quercetin rhamnoside, and quercetin xylosyl-rhamnoside with the deprotonated molecular ions at m/z 463.0875, 433.0774, 447.0928, and 579.1348, respectively (Lee et al., 2019). These metabolites were identified according to their corresponding loss of arabinosyl (132 u), rhamnosyl (146 u), xylosyl (150 u), and glucose (162 u) residues to yield quercetin aglycone. Furthermore, myricetin derivatives could be identified in the extract as myricetin glucoside (52), myricetin robinobioside (56), and myricetin xylosyl-rhamnoside (58) based on the presence of characteristic myricetin aglycone fragment ion at m/z 316 (Lee et al., 2019). Other flavonoids, such as jaceidin rhamnoside (60), could be identified based on deprotonated molecular ion at m/z 505.1351 and fragment ions at m/z 359 (loss of rhamnosyl moiety), 344 (loss of methyl group), and 329 after subsequent deduction of two methyl groups (Taamalli et al., 2013).
Triterpenoids were annotated in the S. jambos leaf extract, including pomolic acid (30) and maslinic acid (79). Both metabolites shared virtually similar deprotonated molecular weight at m/z 471.3469 and fragmentation patterns at m/z 454 (loss of hydroxyl group), 427 (loss of carboxyl group, 44 u), and 393 (loss of hydroxyl, carboxyl, and methyl moieties). Nevertheless, metabolite 30 was identified as pomolic acid and the later eluted metabolite as maslinic acid (79) based on the elution order (Cheng & Cao, 1992; Guo et al., 2011). In addition, madecassic acid (68) was identified based on deprotonated molecular ion at m/z 503.3371 and fragment ions at 485 (loss of H2O), 453 (subsequent loss of HCH2OH), 407 (loss of HCOOH, HCH2OH, and H2O), 370, and 174, which is consistent with previous data (Xia et al., 2015). Asiatic acid (70) was identified with deprotonated molecular ion at m/z 487.3415 and fragment ions at m/z 419, 409 (loss of HCOOH and HCH2OH), 377 (subsequent loss of HCH2OH), 373, and 151 (Xia et al., 2015). Application of UHPLC-MS/MS further strengthened the metabolites identification providing a comprehensive phytochemical characterization of S. jambos leaf extract.
5 Conclusion
In conclusion, the present study provides a comprehensive insight into the Syzygium jambos leaf extracts with different solvent polarities in the retention of bioactive compounds. The application of 1H NMR metabolomics and ultrahigh-performance liquid chromatography-tandem mass spectrometry (UHPLC-MS/MS) allowed the detailed identification of metabolites in S. jambos leaves and the correlation with biological activities, demonstrating distinct clusters among the extracts with various ethanol concentrations. A total of 59 and 30 metabolites were identified via UHPLC-MS/MS and NMR, respectively. The PLS model demonstrated 13 bioactive compounds, including tannins, triterpenoids, and flavonoids, which may significantly contribute to the antioxidant and anti-hyperglycemic activities. This study provides the first comprehensive insights into S. jambos metabolome and serves as a reference for the use of an ingredient in functional food development. Future research into the absolute quantification of bioactive compounds in active extracts and the underlying mechanisms involved will be valuable. However, extensive dose- and time-repetitive toxicity studies were recommended to further validate the safety of this plant.
CRediT authorship contribution statement
Pei Lou WONG: Data curation, Formal analysis, Investigation, Methodology, Software, Writing – original draft. Nurul Shazini RAMLI: Conceptualization, Project administration, Supervision. Chin Ping TAN: Methodology, Software, Supervision, Validation. Azrina AZLAN: Conceptualization, Project administration, Supervision. Faridah ABAS: Conceptualization, Funding acquisition, Project administration, Resources, Supervision, Validation, Visualization, Writing – review & editing.
Acknowledgements
The authors wish to thank Universiti Putra Malaysia for the grant (GP-IPS/2021/9699000) provided under the Putra Graduates Initiative Grant Scheme. The first author also gratefully acknowledges the support from Universiti Putra Malaysia for funding her study under the Graduate Research Fellowship scheme.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Metabolites and biological activities of Phoenix dactylifera L. pulp and seeds: A comparative MS and NMR based metabolomics approach. Phytochem. Lett.. 2019;31:20-32.
- [Google Scholar]
- Abdul-Hamid, N.A., Abas, F., Ismail, I.S., Shaari, K., Lajis, N.H., 2015. Influence of different drying treatments and extraction solvents on the metabolite profile and nitric oxide inhibitory activity of Ajwa dates. J. Food Sci. 80(11), H2603–H2611.
- Alpha-amylase and alpha-glucosidase enzyme inhibition and antioxidant potential of 3-oxolupenal and katononic acid isolated from Nuxia oppositifolia. Biomolecules. 2020;10(61):1-19.
- [Google Scholar]
- Antinociceptive activity of Syzygium jambos leaves extract on rats. J. Ethnopharmacol.. 2007;112:380-385.
- [Google Scholar]
- Baliga, M.S., Ranganath Pai, K.S., Saldanha, E., Ratnu, V.S., Priya, R., Adnan, M., Naik, T.S., 2017. Rose Apple (Syzygium jambos (L.) Alston), in: Fruit and Vegetable Phytochemicals, pp. 1235–1242.
- Syzygium jambos and Solanum guaraniticum show similar antioxidant properties but induce different enzymatic activities in the brain of rats. Molecules. 2013;18(8):9179-9194.
- [Google Scholar]
- Pomolic acid derivatives from the root of Sanguisorba officinalis. Phytochemistry. 1992;31(4):1317-1320.
- [Google Scholar]
- In vivo antiplasmodial activity of four folklore medicinal plants used among tribal communities of Western Ghats, Coimbatore, Tamil Nadu. J. Pharm. Res.. 2014;8(6):751-759.
- [Google Scholar]
- Determination of guava (Psidium guajava L.) leaf phenolic compounds using HPLC-DAD-QTOF-MS. J. Funct. Foods. 2016;22:376-388.
- [Google Scholar]
- Effect of extraction solvent on total phenol content, total flavonoid content, and antioxidant activity of Limnophila aromatica. J. Food Drug Anal.. 2014;22(3):296-302.
- [Google Scholar]
- Eriksson, L., Byrne, T., Johansson, E., Trygg, J., Vikstrom, C., 2006. Process Analytical Technology (PAT) and Quality by Design (QbD), in: Multi- and Megavariate Data Analysis: Basic Principles and Applications, pp. 323–355.
- Chemical profile and in vivo hypoglycemic effects of Syzygium jambos, Costus speciosus and Tapeinochilos ananassae plant extracts used as diabetes adjuvants in Puerto Rico. BMC Complem. Altern. m.. 2015;15(244):1-15.
- [Google Scholar]
- Investigation of lipoxygenase-like activity in strawberry homogenates. Z. Lebensm. Unters. Forsch.. 1988;187(4):330-333.
- [Google Scholar]
- Efficient preparative isolation and identification of walnut bioactive components using high-speed counter-current chromatography and LC-ESI-IT-TOF-MS. Food Chem.. 2014;158:229-238.
- [Google Scholar]
- Guo, S., Duan, J.ao, Tang, Y., Qian, Y., Zhao, J., Qian, D., Su, S., Shang, E., 2011. Simultaneous qualitative and quantitative analysis of triterpenic acids, saponins and flavonoids in the leaves of two Ziziphus species by HPLC-PDA-MS/ELSD. J. Pharmaceut. Biomed. 56(2), 264–270.
- HPLC profiling, antioxidant and in vivo anti-inflammatory activity of the ethanol extract of Syzygium jambos available in Bangladesh. BMC Res. Notes. 2016;9(191):1-8.
- [Google Scholar]
- Electrospray ionization multiple stage quadrupole ion-trap and tandem quadrupole mass spectrometric studies on phosphatidylglycerol from Arabidopsis leaves. J. Am. Soc. Mass Spectr.. 2007;18(4):783-790.
- [Google Scholar]
- Antioxidant and hepatoprotective activity of an ethanol extract of Syzygium jambos (L.) leaves. Drug Discov. Ther.. 2012;6(4):205-211.
- [Google Scholar]
- Dihydrochalcones with radical scavenging properties from the leaves of Syzygium jambos. Nat. Prod. Res.. 2007;21(6):551-554.
- [Google Scholar]
- Phytochemical diversity of Clinacanthus nutans extracts and their bioactivity correlations elucidated by NMR based metabolomics. Phytochem. Lett.. 2015;14:123-133.
- [Google Scholar]
- NMR-based plant metabolomics: Where do we stand, where do we go? Trends Biotechnol.. 2011;29(6):267-275.
- [Google Scholar]
- Modern approach for determination of lactulose, mannitol and sucrose in human urine using HPLC-MS/MS for the studies of intestinal and upper digestive tract permeability. J. Chromatogr. B. 2012;907:34-40.
- [Google Scholar]
- Chemistry and biological activities of flavonoids: An overview. Sci. World J. 2013:1-16.
- [Google Scholar]
- Ultrasound assisted extraction (UAE) of bioactive compounds from fruit and vegetable processing by-products: A review. Ultrason. Sonochem.. 2021;70:105325
- [Google Scholar]
- Total phenolic content, antioxidant capacity and phytochemical profiling of grape and pomegranate wines. RSC Adv.. 2015;5:101683-101692.
- [Google Scholar]
- Lee, S.Y., Mediani, A., Ismail, I.S., Maulidiani, Abas, F., 2019. Antioxidants and α-glucosidase inhibitors from Neptunia oleracea fractions using 1H NMR-based metabolomics approach and UHPLC-MS/MS analysis. BMC Complem. Altern. M. 19(1), 1–15.
- Free radicals, antioxidants and functional foods: Impact on human health. Phcog. Rev.. 2010;4(8):118-126.
- [Google Scholar]
- Syzygium jambos extract mitigates pancreatic oxidative stress, inflammation and apoptosis and modulates hepatic IRS-2/AKT/GLUT4 signaling pathway in streptozotocin-induced diabetic rats. Biomed. Pharmacother.. 2021;142:112085
- [Google Scholar]
- Isolation, characterization and biological activities of phytoconstituents from Lonchocarpus eriocalyx Harms leaves. Trends Phytochem. Res.. 2018;2(3):135-146.
- [Google Scholar]
- Effects of different drying methods and storage time on free radical scavenging activity and total phenolic content of Cosmos caudatus. Antioxidants. 2014;3(2):358-370.
- [Google Scholar]
- Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem.. 1959;31(3):426-428.
- [Google Scholar]
- Separation and characterization of cardiolipin molecular species by reverse-phase ion pair high-performance liquid chromatography-mass spectrometry. J. Lipid Res.. 2010;51(4):856-865.
- [Google Scholar]
- Morton, J., 1987. Rose Apple, in: Fruits of warm climates, pp. 383–386.
- Methoxylated fatty acids in Blumeria graminis conidia. Phytochem.. 2005;66(7):793-796.
- [Google Scholar]
- Syzygium jambos L. Alston: An insight into its phytochemistry, traditional uses, and pharmacological properties. Front. Pharmacol.. 2022;13:1-16.
- [Google Scholar]
- Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics. 2005;4(12):2010-2021.
- [Google Scholar]
- Separation and characterization of chlorophyll degradation products in silkworm using HPLC-UV-APCI-MS. J. Liq. Chromatogr. R. T.. 2003;26(19):3183-3197.
- [Google Scholar]
- Carbohydrates, volatile and phenolic compounds composition, and antioxidant activity of calabura (Muntingia calabura L.) fruit. Food Res. Int.. 2018;108:264-273.
- [Google Scholar]
- Characterization of antioxidant polyphenols from Myrciaria jaboticaba peel and their effects on glucose metabolism and antioxidant status: A pilot clinical study. Food Chem.. 2016;211:185-197.
- [Google Scholar]
- Metabolite profiling, antioxidant, and α-glucosidase inhibitory activities of germinated rice: nuclear-magnetic-resonance-based metabolomics study. J. Food Drug Anal.. 2018;26(1):47-57.
- [Google Scholar]
- Puech, J.L., Mertz, C., Michon, V., Le Guernevé, C., Doco, T., Hervé Du Penhoat, C., 1999. Evolution of castalagin and vescalagin in ethanol solutions. Identification of new derivatives. J. Agr. Food Chem. 47(5), 2060–2066.
- Comprehensive identification of walnut polyphenols by liquid chromatography coupled to linear ion trap-orbitrap mass spectrometry. Food Chem.. 2014;152:340-348.
- [Google Scholar]
- Structure-antioxidant activity relationships of flavonoids and phenolic acids. Free Radic. Bio. Med.. 1996;20(7):933-956.
- [Google Scholar]
- The qualitative analysis of the green tea extract using ESI-MS method. Adv. Technol.. 2014;3(1):30-37.
- [Google Scholar]
- Antibacterial and anti-inflammatory effects of Syzygium jambos L. (Alston) and isolated compounds on acne vulgaris. BMC Complem. Altern. M.. 2013;13(292):1-10.
- [Google Scholar]
- Identification and quantification of phenolic compounds in kernels, oil and bagasse pellets of common walnut (Juglans regia L.) Food Res. Int.. 2015;67:255-263.
- [Google Scholar]
- Phenolic compounds from Syzygium jambos (Myrtaceae) exhibit distinct antioxidant and hepatoprotective activities in vivo. J. Funct. Foods. 2018;41:223-231.
- [Google Scholar]
- Determination of oleanolic acid in human plasma and study of its pharmacokinetics in Chinese healthy male volunteers by HPLC tandem mass spectrometry. J. Pharmaceut. Biomed.. 2006;40(1):190-196.
- [Google Scholar]
- Evaluation of the nitric oxide radical scavenging activity of manganese complexes of curcumin and its derivative. Biol. Pharm. Bull.. 2004;27(2):170-173.
- [Google Scholar]
- IDF Diabetes Atlas: Global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Res. Clin. Pr.. 2022;183:109119
- [Google Scholar]
- UPLC-QTOF/MS for a rapid characterisation of phenolic compounds from leaves of Myrtus communis L. Phytochem. Anal.. 2013;25(1):89-96.
- [Google Scholar]
- In-vitro α-amylase and α-glucosidase inhibitory activity of Adiantum caudatum Linn. and Celosia argentea Linn. extracts and fractions. Indian J. Pharmacol.. 2015;47(4):425-429.
- [Google Scholar]
- Comparison of NO-scavenging and NO-suppressing activities of different herbal teas with those of green tea. Food Chem.. 2007;103:181-187.
- [Google Scholar]
- Hydrolysable tannins and biological activities of Meriania hernandoi and Meriania nobilis (Melastomataceae) Molecules. 2019;24(4):746-761.
- [Google Scholar]
- Antioxidant and α-glucosidase inhibitory phenolics isolated from highbush blueberry flowers. Food Chem.. 2012;135(3):1929-1937.
- [Google Scholar]
- Plant metabolomics: From holistic data to relevant biomarkers. Curr. Med. Chem.. 2013;20:1056-1090.
- [Google Scholar]
- Biological activities of selected plants and detection of bioactive compounds from Ardisia elliptica using UHPLC-Q-Exactive Orbitrap mass spectrometry. Molecules. 2020;25(13):3067-3082.
- [Google Scholar]
- World Health Organization (WHO), 2016. Global report on diabetes, pp 1-83.
- Structural analysis of metabolites of asiatic acid and its analogue madecassic acid in zebrafish using LC/IT-MS. Molecules. 2015;20(2):3001-3019.
- [Google Scholar]
- Induction of apoptosis by hydrolyzable tannins from Eugenia jambos L. on human leukemia cells. Cancer Lett.. 2000;157:65-75.
- [Google Scholar]
- A simple 96-well microplate method for estimation of total polyphenol content in seaweeds. J. Appl. Phycol.. 2006;18(3–5):445-450.
- [Google Scholar]
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105546.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







