Translate this page into:
Optimization of Portulaca oleracea L. extract using response surface methodology and artificial neural network and characterization of bioactive compound by high-resolution mass spectroscopy
⁎Corresponding author at: Department of Food Science and Biotechnology, Kyungpook National University, Daegu 41566, Republic of Korea. sang@knu.ac.kr (Sang-Han Lee)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
The well-known medicinal plant Portulaca oleracea L. (PO) is used as a traditional medicine and culinary herb to treat various diseases. Fatty acids, essential oils, and flavonoids were extracted from PO seeds and leaves using ultrasonic, microwave, and supercritical fluid extraction with RSM techniques. However, investigations on the secondary metabolites and antioxidant capabilities of the aerial part of PO (APO) are scarce. In order to extract polyphenols and antioxidants from APO as effectively as possible, this study used heat reflux extraction (HRE), response surface methodology (RSM), and artificial neural network (ANN) modeling. It also used high-resolution mass spectrometry to identify the APO secondary metabolite. A central-composite design (CCD) was used to establish the ideal ethanol content, extraction time, and extraction temperature to extract the highest polyphenolic compounds and antioxidant activity from APO. According to RSM, the highest amount of TPC (8.23 ± 1.06 mgGAE/g), TFC (43.12 ± 1.15 mgCAE/g), DPPH-scavenging activity (43.01 ± 1.25 % of inhibition) and FRAP (35.98 ± 0.19 µM ascorbic acid equivalent) were obtained at 60.0 % ethanol, 90.2 % time, and 50 °C. Statistical metrics such as the coefficient of determination (R2), root-mean-square error (RMSE), absolute average deviation (AAD), and standard error of prediction (SEP) revealed the ANN's superiority. Ninety-one (91) secondary metabolites, including phenolic, flavonoids, alkaloids, fatty acids, and terpenoids, were discovered using high-resolution mass spectrometry. In addition, 21 new phytoconstituents were identified for the first time in this plant. The results revealed a significant concentration of phytoconstituents, making it an excellent contender for the pharmaceutical and food industries.
Keywords
Antioxidant
Artificial neural network
Portulaca oleracea
Response surface methodology
Secondary metabolites
1 Introduction
Portulaca oleracea L. (PO) is a well-known medicinal plant used both as a traditional medicine and as an edible herb to treat various ailments. This herb is widely used in European folk medicine. Additionally, PO is mentioned in some pharmacopeias, such as the Ayurvedic Pharmacopoeia of India and the Pharmacopoeia of PR China (Iranshahy et al., 2017). Pharmacological investigations have demonstrated that PO has a wide range of biological effects, including anti-inflammatory, a bronchodilator, anti-microbial, antioxidant, and neuroprotective characteristics (Malek et al., 2004, Wang et al., 2007, Hozayen et al., 2011, Karimi et al., 2011, Du et al., 2017). Animal studies have demonstrated its hepatoprotective, antiulcerogenic, and antifertility benefits (Kumar et al., 2010, Nayaka et al., 2014, Eidi et al., 2015). Additionally, investigations on phytochemistry have revealed that this plant includes minerals, vitamins, fatty acids, flavonoids, alkaloids, and terpenoids (Sakai et al., 1996, Xiang et al., 2005, Yan et al., 2012, Petropoulos et al., 2016).
Numerous studies have demonstrated that the solvent concentration, incubation time, and temperature affect the effectiveness of polyphenol extraction, while some thermolabile bioactive substances may degrade during extraction (Saha et al., 2011). There are several extraction strategies for bioactive molecules, including reflux, soxhlet, microwave-aided, ultrasonicator-assisted, and supercritical fluid extraction (Pandey and Banik 2012). Heat reflux extraction has several benefits over more traditional extraction methods, (1) the solvent is replenished in the extraction; (2) the mass transfer driving force is more substantial, (3) the extraction takes less time, (4) the solvent is used less because it has already been used, and (5) the extraction yield is increased. This technology is a promising substitute for extracting bioactive natural compounds due to its benefits over traditional extraction methods (Gong et al., 2014, Ma et al., 2022).
Extraction is the initial and most crucial step in collecting and purifying bioactive chemicals from plant sources; yet, lengthy extraction times and low extraction efficiency limit these approaches (Samuel and Emovon 2018, Sedraoui et al., 2020). Analytical techniques were optimized using multivariate statistical methodologies to address this problem. Response surface methodology (RSM) combines mathematical and statistical methods that have proven effective in developing, improving, and optimizing complex processes (David Samuel et al., 2021). RSM provides a wealth of information and is more cost-effective because it reduces the required experiments. In addition, RSM assesses the simultaneous influence of several factors and anticipates the system's response to each new condition to find the optimal circumstances for the predicted response (Kusuma et al., 2021, Kusuma et al., 2022). Nevertheless, nonlinearity and inaccurate data are not handled precisely by RSM approaches. It has also been demonstrated that artificial neural networks (ANNs) are effective data-driven computational tools with the flexibility to capture complex and non-linear data (Okwu et al., 2020, Okwu et al., 2021). The operation of ANN as a prediction tool is similar to that of the human brain. The brain's neurons, basic processing units connected by networks and used to transmit messages between the neurons, served as the model's primary source of inspiration. The sigmoid function controls the network (Samuel and Okwu 2019, Okwu et al., 2021, Zadhossein et al., 2021). However, the black box learning technique associated with the ANN cannot be utilized to correlate input factors and output variables (Gupta and Sharma 2014). This problem is circumvented by incorporating an additional method, such as RSM, to analyze the interaction between the input and response variables. Hence, Combining RSM with ANN resulted in a more precise forecast (Samuel and Okwu 2019).
Mounting studies over the past few decades have shown how to extract fatty acids, essential oils, and flavonoids from PO seeds and leaves using ultrasonic, microwave, and supercritical fluid extraction with RSM methods (Stroescu et al., 2013, Wang et al., 2014, Sodeifian et al., 2018). Most of the research disclosed just process optimization. However, the authors did not compare the efficacy of predictive modeling with better methodologies, such as ANN, and there was a dearth of secondary metabolite profiling of the improved extracts. To the best of our knowledge, however, heat reflux extraction (HRE) using RSM and ANN was used for the first time in this study to increase the polyphenol content and antioxidant activity of the aerial portions of the Portulaca oleracea (APO). This study aimed to examine and improve extraction parameters, including extraction temperature and duration, as well as ethanol concentration, using the RSM central composite design (CCD) tool to obtain the highest polyphenolic content and antioxidant potential from APO. Additionally, for the first time, we have profiled the secondary metabolites of APO using high-resolution mass spectrometry analysis.
2 Materials and methods
2.1 Sample collection and preparation
Wild Portulaca oleracea L. was collected in September 2021 in Daegu, Korea. The Department of Food Science and Technology, Kyungpook National University, Daegu, Korea (voucher specimen # FT1005), identified the sample. Heat reflux extraction (HRE) was done in an oven with a condenser (Soxhlet water bath C-WBS-D6, Changshin Science, Seoul, Korea). Dry powder samples (10.0 g) were extracted using 100 ml of solvent following the instructions in supplemental Table 1. The extracted materials were filtered on Whatman No. 1 filter paper (Schleicher & Schuell, Keene, New Hampshire) and then dried in a freeze drier (Il-shin Biobase, Goyang, Korea). The APO extract was kept at −20 °C for the ensuing investigations. X1. Ethanol concentration (%); X2. time (min); X3. temperature (°C); TPC. total phenolic content (mg gallic acid equivalent/g dry weight extract); TFC. total flavonoid content (mg catechin equivalent/g dry weight extract); DPPH. DPPH radical scavenging activity (% inhibition); FRAP. ferric reducing antioxidant power (µM ascorbic acid equivalent).
Run
Independent variables
Responses
(X1)
(X2)
(X3)
TPC (Y1)
TFC (Y2)
DPPH (Y3)
FRAP (Y4)
1
50
140
50
5.09 ± 0.68
21.20 ± 0.73
20.87 ± 0.72
19.03 ± 0.05
2
50
40
50
3.89 ± 0.83
19.42 ± 0.52
15.36 ± 0.39
16.36 ± 0.02
3
50
90
50
8.12 ± 0.34
41.89 ± 0.25
41.75 ± 0.37
35.19 ± 0.16
4
50
90
30
3.15 ± 0.26
29.07 ± 0.65
9.18 ± 0.24
12.04 ± 0.10
5
0
90
50
1.25 ± 0.41
19.76 ± 0.32
0.66 ± 1.00
3.04 ± 0.07
6
75
60
60
6.58 ± 0.82
29.09 ± 0.10
27.21 ± 0.21
26.02 ± 0.08
7
50
90
50
7.61 ± 1.02
41.75 ± 0.56
41.93 ± 0.25
35.19 ± 0.06
8
75
120
40
5.06 ± 0.49
31.02 ± 0.26
25.07 ± 0.45
23.12 ± 0.02
9
75
60
40
5.03 ± 0.42
32.02 ± 0.95
22.70 ± 0.54
22.26 ± 0.04
10
25
120
60
5.06 ± 0.23
25.75 ± 0.35
20.08 ± 0.98
17.01 ± 0.08
11
50
90
50
7.59 ± 0.62
43.20 ± 0.26
40.56 ± 0.10
35.19 ± 0.04
12
100
90
50
5.75 ± 0.06
31.50 ± 0.33
30.25 ± 1.02
25.93 ± 0.02
13
50
90
50
7.49 ± 0.04
43.02 ± 0.53
40.05 ± 0.56
35.17 ± 0.06
14
75
120
60
6.84 ± 0.08
29.02 ± 0.35
29.88 ± 0.46
27.96 ± 0.16
15
25
60
60
3.17 ± 0.06
24.52 ± 0.15
14.02 ± 0.29
15.33 ± 0.13
16
50
90
70
5.53 ± 0.24
31.01 ± 0.60
22.53 ± 0.37
21.33 ± 0.19
17
25
60
40
2.81 ± 0.68
23.13 ± 0.72
7.01 ± 0.19
10.46 ± 0.15
18
50
90
50
7.98 ± 0.68
43.11 ± 0.72
41.89 ± 0.73
35.09 ± 0.07
19
50
90
50
8.12 ± 0.83
43.23 ± 0.39
41.05 ± 0.52
36.09 ± 0.05
20
25
120
40
2.82 ± 0.34
21.03 ± 0.37
8.51 ± 0.25
9.19 ± 0.16
2.2 Total phenolic content (TPC), total flavonoid content (TFC) and antioxidant activities
The TPC and TFC of APO extracts were assessed using the Folin-Ciocalteu assay and the aluminum chloride colorimetric method, respectively (Alam et al., 2017). The corresponding regression equations for the calibration curves were used to determine the TPC (y = 0.0582x + 0.0038; r2 = 0.9955) and TFC (y = 0.059x + 0.0081; r2 = 0.9879). The gallic acid equivalent (mg)/dry weight sample (g) and catechin equivalent (mg)/dry weight sample (g) were used as the units of measurement for the TPC and TFC, respectively. DPPH-radical scavenging test and ferric reducing antioxidant power (FRAP) assay were used to assess the antioxidant properties of APO extracts (Alam et al., 2021).
2.3 RSM design and extraction process
The RSM model was designed to extract phenolic chemicals from APO using ethanol concentration (X1), extraction duration (X2), and temperature (X3) as independent process factors. Respondent factors included TPC, TFC, DPPH-scavenging activity and FRAP (Y1-Y4, respectively). A three-component, five-layer CCD was employed for the extractions (supplementary data Table S1). The CCD is widely utilized due to its adaptability. The early experimental results from a factorial design can be used in a CCD that only makes the axial points, eliminating resource waste. Nonetheless, the most distinctive parameters are the axial points (−α, α). These locations are outside the minimum and maximum limits of the factors, guaranteeing the response surface's curvature and allowing the construction of ideal conditions. In contrast to the BBD, a CCD can distinguish between axial point estimates based on orthogonal and rotational characteristics. An orthogonal design allows for an independent examination of the primary, interaction, and quadratic effects, simplifying the interpretation of the results. The second-order polynomial model equation (Eq.1) describes the link between independent factors and replies.
The model's adequacy was assessed using the determination coefficient (R2), the adjusted determination coefficient (Adj.R2), and the lack of fit test. The F-value (p < 0.05) was significant. Three-dimensional (3D) surface plots showed each factor's effect on response value. The RSM analysis and multiple linear regression were carried out using Design Expert 11 (Stat-Ease, Minneapolis, Minnesota, USA).
2.4 Artificial neural networks (ANN) modeling
The ANN modelling was systematically conducted by using the dataset presented in Table 1. MATLAB R2020a software (The Neural Network Toolbox, Inc., USA) was employed to create the ANN model. The ANN design consists of data collection; model development using different functions and algorithms; configuration of the model; weights and biases initialization; training, testing, and validation of the model. The MLP (multilayer perceptron network) topology consists of input, hidden, and output layers. Independent variables (X1, X2, and X3) were used as input vectors, and four responses (Y1, Y2, Y3, and Y4) were applied as target vectors (Fig. 1A). The data were divided into three subsets, where 70, 15, and 15 % of the whole data points were utilized for training, validation, and testing, respectively. In the training step, the feed-forward network and cascade feed-forward network with the Broyden-Fletcher-Goldfarb-Shanno algorithm (BFGS) and Levenberg-Marquardt back-propagation algorithm were used to lessen the mean square error (MSE). The MSE was calculated using Eq. (2)
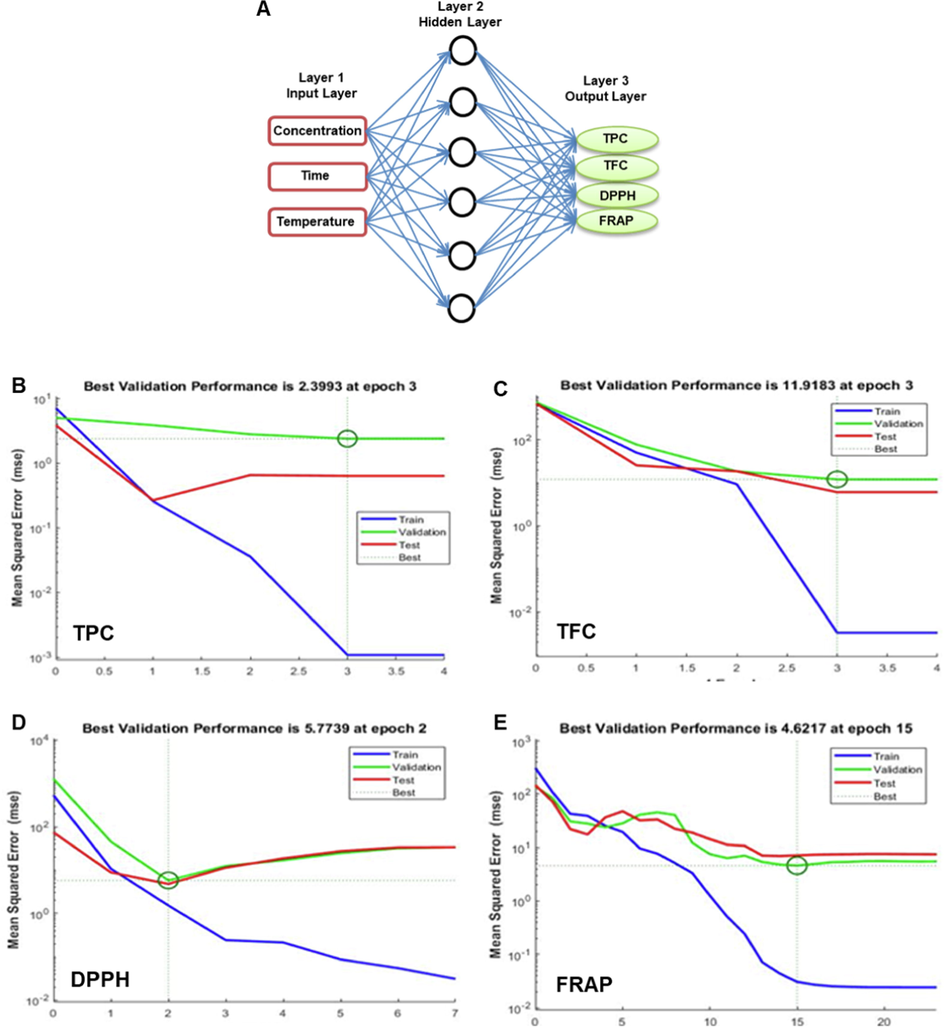
ANN model topology. The best ANN model in terms of architecture (A), network training curves for trained subsets with epoch numbers for TPC (B), TFC (C), DPPH (D) and FRAP (E) using the MATLAB software.
2.5 Comparison of the RSM and ANN models' predictive abilities
To evaluate the estimation skills of RSM and ANN, several statistical metrics such as R2, RMSE, AAD, and SEP were calculated using the following equations.
2.6 Model validation
The optimal extraction parameters were determined using response surface and desirability function analysis. A series of three experiments was conducted under ideal conditions to ensure the model's accuracy, with the average experimental results compared to predictions. In addition, the electrospray ionization mass spectrometry (ESI-MS)/MS profiles of bioactive compounds were identified under optimum conditions.
2.7 Identification of bioactive compounds by ESI-MS/MS analysis
The negative (-) mode ESI-MS was performed on a Q-Exactive Orbitrap mass spectrometer (Thermo Fisher Scientific Inc., San Jose, CA, USA). Immersing the sample in the ESI source required a 500 L graded syringe (Hamilton Company Inc., Reno, NV, USA) and a 15 L/min syringe pump (Model 11, Harvard, Holliston, MA, USA). The normal negative mode ESI-MS conditions were as follows: mass resolution of 140,000 (full width at half maximum, FWHM), sheath gas flow rate of 5, seep gas flow rate of 0, auxiliary gas flow rate of 0, spray voltage of 4.20 kV, capillary temperature of 320 °C, S-lens Rf level, and automatic gain control of 5 E 6. The MS/MS investigations used the same apparatus with three stepwise normalized collision energies (10, 20, and 30) (Alam et al., 2021). Mass spectrum data was processed using the Xcalibur 3.1 with Foundation 3.1 (Thermo Fisher Scientific Inc., Rockford, IL, USA). The compounds were probably found by comparing the calculated (exact) masses of deprotonated (M−H) adducts with the m/z values and ESI-MS/MS fragmentation patterns from the in-house MS/MS database and online databases like FooDB (Naveja et al., 2018), METLIN (Guijas et al., 2018), CFM-ID 4.0 (Wang et al., 2021). The ChemDraw Professional 15.0 (PerkinElmer, Waltham, MA, USA) was used to draw the chemical structure.
2.8 Statistical analysis
All data were reported as the mean ± standard deviation of at least three independent experiments (n = 3), each with three sample replicates. Differences were considered significant at p < 0.001, p < 0.01, and p < 0.05.
3 Results and discussion
3.1 Fitting of the RSM and ANN models
For each extraction circumstance, Table 1 describes the experimental settings and conclusions. All response variables were transformed into second-order quadratic polynomial equations to account for changes in answers as a function of extraction factors. ANOVA was used to determine whether the fitted second-order quadratic model equations were statistically significant. The regression coefficient (β), adjusted correlation factor (R2), coefficient of variation (CV), and adequate precision were used to describe how well the model fit (Table 2). The nonsignificant terms (p > 0.05) were taken out of the models to improve the fit and predictions. We used the p-values to figure out how vital each coefficient was. When the p-values were<0.05, 0.01, and 0.001, the model terms were significant, very significant, and strikingly significant, respectively. RC. Regression coefficient; SS. sum of squares; MS. mean square.
ANOVA for quadratic model for TPC
Source
RC
SS
DF
MS
F-value
p-value
Model
81.54
9
9.06
102.01
< 0.0001
Significant
Intercept
7.82
Linear terms
X1
1.17
21.74
1
21.74
244.77
< 0.0001
Significant
X2
0.3091
1.30
1
1.30
14.58
0.0034
Significant
X3
0.6681
7.14
1
7.14
80.42
< 0.0001
Significant
Interaction terms
X1X2
−0.2012
0.3240
1
0.3240
3.65
0.0852
X1X3
0.0912
0.0666
1
0.0666
0.7500
0.4068
X2X3
0.2637
0.5565
1
0.5565
6.27
0.0313
Significant
Quadratic terms
X12
−1.08
29.78
1
29.78
335.27
< 0.0001
Significant
X22
−1.20
19.41
1
19.41
218.56
< 0.0001
Significant
X32
−0.8696
19.32
1
19.32
217.52
< 0.0001
Significant
Lack of Fit
0.4767
5
0.0953
1.16
0.4379
not significant
Pure error
0.4115
5
0.0823
R2
0.9892
Adjusted R2
0.9795
Adeq Precision
31.5543
C.V. %
5.47
ANOVA for quadratic model for TFC
Model
1414.37
9
157.15
249.82
< 0.0001
Significant
Intercept
42.67
Linear terms
X1
3.14
157.50
1
157.50
250.38
< 0.0001
Significant
X2
0.0757
0.0778
1
0.0778
0.1236
0.7324
X3
0.2375
0.9025
1
0.9025
1.43
0.2586
Interaction terms
X1X2
−0.0250
0.0050
1
0.0050
0.0079
0.9307
X1X3
−1.38
15.24
1
15.24
24.22
0.0006
Significant
X2X3
0.5325
2.27
1
2.27
3.61
0.0868
Quadratic terms
X12
−4.30
472.67
1
472.67
751.40
< 0.0001
Significant
X22
−8.14
894.88
1
894.88
1422.58
< 0.0001
Significant
X32
−3.12
248.70
1
248.70
395.35
< 0.0001
Significant
Lack of Fit
3.93
5
0.7861
1.67
0.2946
Non-Significant
Pure error
2.36
5
0.4720
R2
0.9956
Adjusted R2
0.9916
Adeq Precision
41.8660
C.V. %
2.54
ANOVA for quadratic model for DPPH
Model
3330.37
9
370.04
412.69
< 0.0001
Significant
Intercept
41.13
Linear terms
X1
7.15
818.25
1
818.25
912.55
< 0.0001
Significant
X2
1.61
35.01
1
35.01
39.04
< 0.0001
Significant
X3
3.41
186.32
1
186.32
207.80
< 0.0001
Significant
Interaction terms
X1X2
−0.3150
0.7938
1
0.7938
0.8853
0.3689
X1X3
−1.16
10.72
1
10.72
11.95
0.0061
Significant
X2X3
0.6075
2.95
1
2.95
3.29
0.0997
Quadratic terms
X12
−6.52
1085.41
1
1085.41
1210.51
< 0.0001
Significant
X22
−8.49
974.81
1
974.81
1087.16
< 0.0001
Significant
X32
−6.42
1052.36
1
1052.36
1173.65
< 0.0001
Significant
Lack of Fit
5.90
5
1.18
1.92
0.2449
Non-significant
Pure error
3.07
5
0.6132
R2
0.9973
Adjusted R2
0.9949
Adeq Precision
60.2976
C.V. %
3.78
ANOVA for quadratic model for FRAP
Model
1975.64
9
219.52
900.96
< 0.0001
Significant
Intercept
35.30
Linear terms
X1
5.82
542.31
1
542.31
2225.80
< 0.0001
Significant
X2
0.5651
4.33
1
4.33
17.77
0.0018
Significant
X3
2.49
99.35
1
99.35
407.77
< 0.0001
Significant
Interaction terms
X1X2
0.2988
0.7140
1
0.7140
2.93
0.1177
X1X3
−0.5112
2.09
1
2.09
8.58
0.0151
Significant
X2X3
0.5038
2.03
1
2.03
8.33
0.0162
Significant
Quadratic terms
X12
−5.23
698.20
1
698.20
2865.65
< 0.0001
Significant
X22
−6.39
551.56
1
551.56
2263.76
< 0.0001
Significant
X32
−4.68
559.01
1
559.01
2294.37
< 0.0001
Significant
Lack of Fit
1.72
5
0.3435
2.39
0.1806
Non-significant
Pure error
0.7190
5
0.1438
R2
0.9988
Adjusted R2
0.9977
Adeq Precision
93.2690
C.V. %
2.14
Table 2 shows that model terms are significant (p < 0.0001). The R2 values (0.9892–0.9988) of the built regression models indicate a high level of statistical significance. The appropriate precision indicates a signal-to-noise ratio, and > 4 is ideal (Alam et al., 2022). The ratio ranged between 31.55 and 93.26, showing a significant signal and suitability for this method. The coefficient of variation (CV) measures the repeatability of a model, and the range of 2.14 to 5.47 indicates that the model is reproducible. Multiple linear regression equations were used to create 3D surfaces and contour plots to show independent variable interactions (Fig. 2A-D).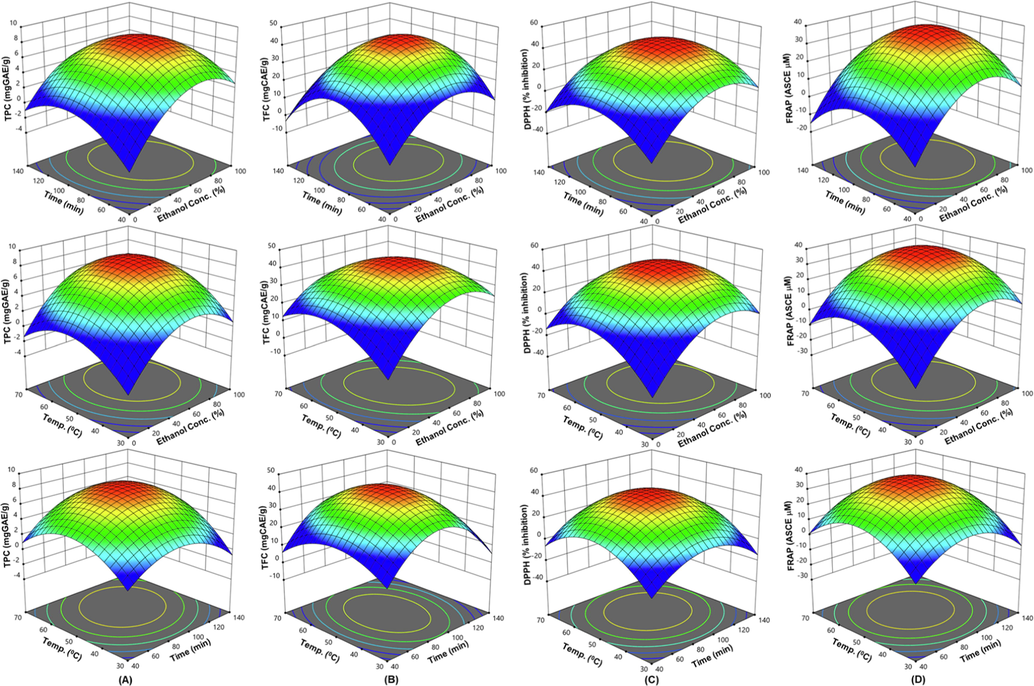
The three-dimensional (3D) response surface plots of APO extraction for TPC (A), TFC (B), DPPH-radical scavenging activity (C), and FRAP (D) for ethanol concentration, time, and temperature as a function of key interaction factors for RSM.
Mounting evidence revealed that ANN modeling is superior and more sophisticated than RSM, and ANNs are a feasible alternative to RSM for complicated nonlinear multivariate modeling. ANNs are more exact than RSM at fitting experimental responses, predicting, and modeling biological processes (Huang et al., 2017). ANN modeling was used to verify the experimental values. The trained ANN model's predicted values are in Table S2. The ANN predicts nonlinear relationships between extraction parameters (X1, X2, and X3) and response variables (Y1, Y2, Y3, and Y4). The ANN model predicted values that were pretty close to the actual values, proving its accuracy. By comparing network training and testing errors, the hit-and-try strategy modified the number of hidden layer neurons. The experiment investigated the lowest practicable error between training and testing and the minimal number of epochs to prevent model overfitting; the results were consistent with earlier efforts (Choi et al., 2022). The Levenberg-Marquardt approach produced the best validation result for all dependent variables Y1, Y2, Y3, and Y4 (Fig. 1B-E.
3.2 Comparison of the RSM and ANN models' predictive abilities
Both the RSM and ANN models' prediction and estimation skills were examined. Comparative similarity plots were utilized to examine the ANN model's four target response predictions (Y1, Y2, Y3, and Y4). In terms of fitting experimental data to all target responses, the ANN model was more accurate, precise, and assessable than the RSM model (supplementary data Table S2). The RSM model had a larger discrepancy between projected and actual data, whereas the ANN model's residuals remained steady.
To compare RSM with ANN, R2, RMSE, AAD, and SEP were calculated (Table 3). A better model has lower RMSE, AAD, and SEP while higher R2. R2 values of the trained ANN model were greater than those of the RSM model, suggesting the ANN model's superiority in predicting all four dependent variables. The AAD gauges the deviation between projected and actual data, while RMSE shows model fit. The ANN outperformed RSM by having lower AAD and RMSE values. The ANN model also showed low SEP values, which ranged from 0.0813 to 0.3126. The ANN model is more predictive than the RSM model because it can approximate nonlinear systems, while the RSM model requires second-order polynomial regression. The ANN model is also unaffected by experimental design and calculates several replies in a single run, while the RSM model takes multiple runs for multi-response optimization (Dadgar et al., 2015). R2. correlation coefficient; RMSE. root-mean-square error; AAD. absolute average deviation; SEP. standard error of prediction.
Parameters
TPC
TFC
DPPH
FRAP
RSM
ANN
RSM
ANN
RSM
ANN
RSM
ANN
R2
0.9892
0.9976
0.9956
0.9997
0.9973
0.9985
0.9988
0.9995
RMSE
0.1299
0.0753
2.3413
1.7888
1.6655
1.0867
1.3608
1.1492
AAD (%)
2.2909
1.1408
6.6496
3.0372
28.8295
14.4468
4.9237
3.1221
SEP (%)
0.1401
0.0813
0.4091
0.3126
0.4675
0.3050
0.3567
0.3012
3.3 Influence of HRE parameters on TPC and TFC
In APO extracts, TPC and TFC contents ranged from 1.25 ± 0.41 to 8.12 ± 0.34 mgGAE/g and 19.76 ± 0.32 to 43.23 ± 0.91 mgCAE/g, respectively (Table 1). Both the TPC and TFC exhibited a substantial linear influence of X1 and the quadratic component of (X12), (X22), and (X32) (supplementary data Figure S1). The second-order polynomial equations in eqs. (8) and (9) illustrate the relationships between TPC, TFC, and their variables.
The TPC and TFC showed nonsignificant lack of fit values (F = 1.16 and 1.67, respectively) showing the model accurately predicted R2 = 0. 9892 (TPC) and 0.9956 (TFC) and Adj.R2 = 0. 9795 (TPC) and 0.9949 (TFC) (Table 2). The RSM model accurately predicted the parameter impacts on TPC and TFC of the APO extract. As depicted in Fig. 2(A, B), at 50 °C, 50 % ethanol produced the most TPC and TFC in 90 min. Previous studies revealed that medium-concentration ethanol may make the solvent more polar, dissolving more polar and moderately polar phenolic compounds (Sedraoui et al., 2020). Moderate ethanol in water can affect the architecture and structure of membrane phospholipids. This affects plant cell penetrability, allowing higher polyphenol extraction and diffusion (Gurtovenko and Anwar 2009). Experiments in a prior comparison investigation found that extraction of phenolic compounds from green tea leaves under high hydrostatic pressure increased with ethanol in the solvent; peaked at 50 % ethanol and fell after that (Xi and Wang 2013).
3.4 Effect of HRE parameters on the in vitro antioxidant capacity (AC)
A linear significant influence of ethanol content (X1), a quadratic effect of concentration (X1), time (X2) and temperature (X3) as well as interaction between concentration and temperature (X1X3) (supplementary data Figure S1) on antioxidant activity were found using DPPH radical scavenging activity and FRAP analyses. Eqs. (10) and (11) display the fitted second-order polynomial equations for DPPH (% inhibition) and FRAP (ascorbic acid equivalent μM):
The AC values ranged from 0.66 ± 1.00 % to 41.89 ± 0.22 % inhibition of DPPH and from 3.04 ± 0.07 to 36.09 ± 0.34 μM ascorbic acid equivalent (Table 1). The ANOVA results show that the data fitted the model results for DPPH (R2 = 0.9973 and Adj. R2 = 0.9949) and FRAP response (R2 = 0.9988 and Adj. R2 = 0.9977), and the lack of fit was nonsignificant (F = 1.92 for DPPH and 2.39 for FRAP) (Table 2). As depicted in Fig. 2(C, D), at 50 °C, 50 % ethanol produced the highest DPPH inhibition and FRAP value in 90 min. This indicates that the capacity for electron and proton donation improves with increasing amounts of the organic solvent. This outcome is in line with the earlier discovery for TFC that maximum extraction calls for 75 percent ethanol (Do et al., 2014). The extraction of considerable polyphenolics from APO, both in terms of quality and quantity, is made possible by raising the ethanol concentration. There is growing evidence that ethanol concentrations affect antioxidant activity and polyphenolic compound quality and amount (Zhu et al., 2011, Do et al., 2014).
3.5 Model validation
The desirability function optimizes TPC, TFC, DPPH, and FRAP simultaneously. Derringer's desirability function was used to anticipate the parameters, allowing a multivariate analysis to find the best level for all replies in a single extraction. In this study, the following conditions (X1, 60 %), (X2: 90.5 min), and (X3, 50 °C), was used to achieve the maximal overall desirability D = 0.999 (on a scale of 0 to 1). Under these optimal conditions, the predicted values for TPC, TFC, percentage inhibition of DPPH, and µM ascorbic acid equivalent FRAP are 8.12 mgGAE/g, 43.23 mgCAE/g, and 42.98 %, and 36.81, respectively. To verify the sufficiency of the model equations, a duplicate experiment was conducted in the optimal conditions predicted by Derringer's desire model. The following results were obtained: TPC = 8.23 ± 1.06 mgGAE/g, TFC = 43.12 ± 1.15 mgCAE/g, % inhibition of DPPH = 43.01 ± 1.25 %, and µM ascorbic acid equivalent FRAP = 35.98 ± 0.19. The model efficiently optimized the common extraction parameters for all responses, as evidenced by the good agreement between experimental and expected values (supplementary data Table S3).
Additionally, comparison research between this study and earlier studies was carried out to confirm the high extractability of hydro-alcoholic solvents for polyphenols and the antioxidant properties of APO. The hydro-alcoholic solvent had more TPC and DPPH scavenging action than other solvents, as indicated in Table 4. When compared to alternative solvents, which were typically utilized in earlier studies, these comparisons showed that the HRE technique using hydro-alcoholic solvent was a high-efficiency technique.
3.6 Identification of secondary metabolites in APO by high-resolution mass spectroscopy
The ESI-MS/MS in negative ionization techniques detected secondary metabolites in APO extracts. Table 4 shows that 93 compounds were identified in negative mode utilizing MSn data from the precursor ion mass, fragments, recognized fragmentation patterns for the provided classes of compounds, and neutral mass loss, as well as literature and online database searches. The confidence level determined the significance of these results. Level 2 shows the likely structure of the detected substance, whereas Level 3 denotes a speculative candidate. (Schymanski et al., 2014). HM. 60% methanol; M. 100% methanol; E. 100% ethanol and W. 100% aqueous solvent.
Methods
Solvent
TPC (mgGAE/g)
DPPH IC50 (mg/ml)
Ref.
HRE
HM
8.23
1.01
Present study
HRE
M
4.78
1.78
HRE
M
6.98
2.52
HRE
E
3.60
3.56
HRE
W
4.41
2.35
3.6.1 Phenolic acids
A phenolic acid can lose methyl (15 Da), hydroxyl (18 Da), or carboxyl (44 Da) to form a fragment ion. Fragmentation of a phenolic acid glycoside begins with cleavage of the glycosidic bond to give phenolic acid m/z and sugar loss (neutral mass loss of 162 Da) (Choi et al., 2022). In addition, hydroxycinnamic acid conjugates yield quinate (m/z 191) by loss of the acyl group and dehydrated quinate (m/z 173), coumarate (m/z 163), caffeate (m/z 179), ferulate (m/z 193) and sinapate (m/z 223) through β-elimination of a carboxylic acid (Jaiswal et al., 2010, Parveen et al., 2011). Thus, compounds 1–3, 5, 7, 8, 10–16, and 20 were tentatively identified as hydroxy benzoic acid, coumaric acid, ferulic acid, caffeic acid phenethyl ester, ellagic acid, protocatechuic acid glucoside, coumaroylquinic acid, caffeic acid hexoside, ferulylshikimic acid, ferulic acid hexoside, syringoylquinic acid, caffeic acid derivatives, feruloyl galactaric acid and hexosyl caffeoyl hexose (Islam et al., 2020, Alam et al., 2021, Fernández-Poyatos et al., 2021, Choi et al., 2022). In addition, by comparing the fragmentation patterns to those previously published in the literature, compound 4, 6, 17, 18 and 21 was identified as maclurin (C13H10O6), uralenneoside (C12H14O8), picraquassioside A (C18H22O10), methylpicraquassioside A (C19H24O10) and arillatose B (C22H29O14), respectively which yielded a precursor ion [M−H]¯ at m/z 261.0401, 285.0612, 397.1142, 411.1302, and 517.1563, respectively (Berardini et al., 2004, Abdelrahman et al., 2017, Llorach et al., 2019, Tang et al., 2020). All of these substances were intriguingly discovered for the first time in APO. Furthermore, compound 9 generated a monoisotopic ion [M−H]¯ at m/z 333.0618 and produced fragment ions at m/z 289.07 by lose of carboxyl (-44 Da) group and at m/z 271.06 through successive loss of H2O. It also yielded a characteristic ion at m/z 167.03 by cleavage of ether bond between C7-C9 and tentatively confirmed as berceloneic acid B, which has been identified as first time in APO (Fig. 3A). In addition, compound 19 was tentatively identified as paederol B with molecular formula (C20H28O12), generated a deprotonated ion at m/z 459.1511 and yielded the following fragment ions: m/z at 399.12 ([M−H−61 Da]), 341.08 ([M−H−C4H10O3−CH3]), 281.08 ([M−H−178 Da]) and 193.05 (ferulate ion) through β-elimination of a carboxylic acid (Fig. 3B). This compound has also been first time identified in APO.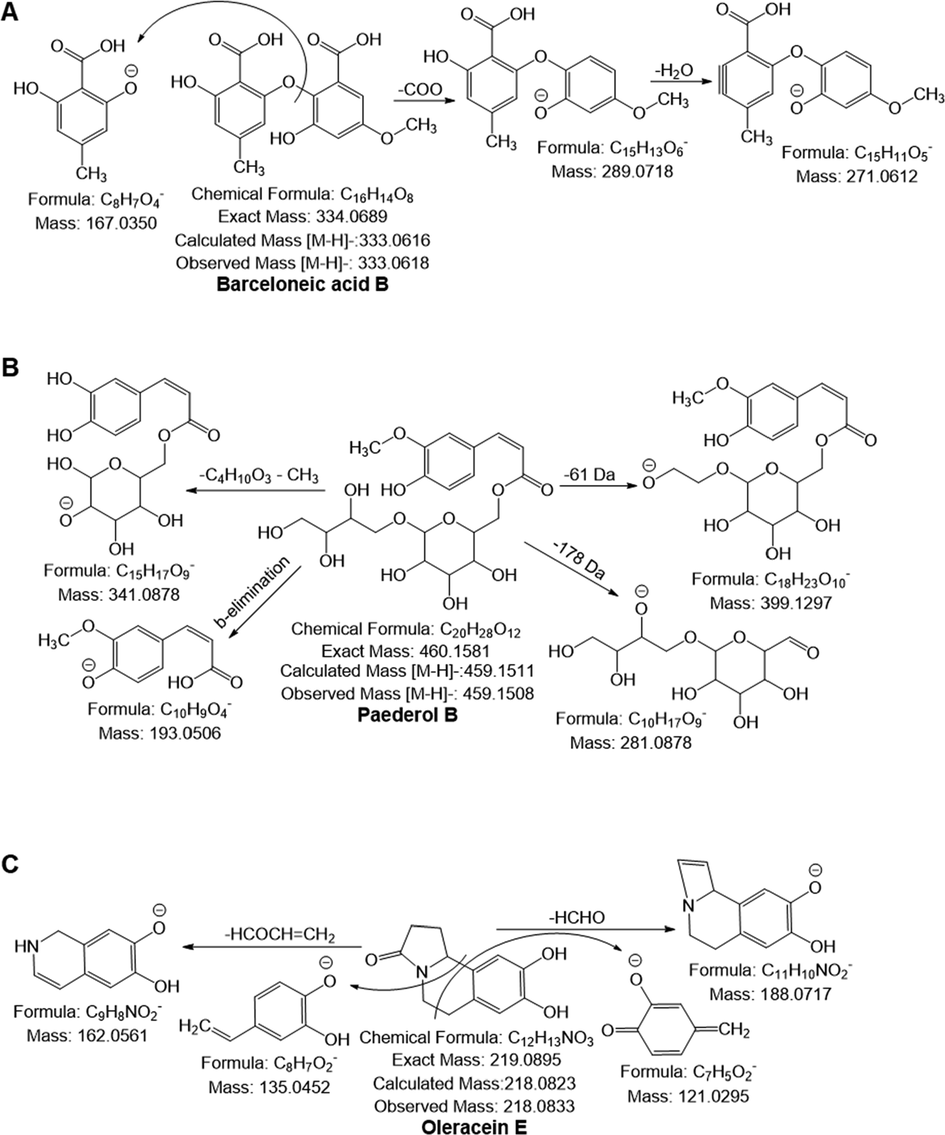
Possible mass fragmentation pattern of (A) barceloneic acid B, (B) paederol B and (C) Oleracein E.
3.6.2 Flavonoids
According to a prior study, each subgroup of flavonoids exhibits a distinct fragmentation pattern during mass analysis. The most common fragmentation of flavonoids is the cleavage of the C-ring bonds (retro-Diels-Alder, i.e., RDA mechanism), which forms ions with the A- or B-ring and a portion of the C-ring. There may also be significant losses of tiny neutral molecules, such as CO (28 Da), C2H2O (42 Da), COO (44 Da), 2CO (56 Da), CO + COO (72 Da), and 3CO (84 Da). A unique ion [M−H−CH3], distinguished by the loss of 15 Da, is also present in methylated flavonoids (Alam et al., 2021, Choi et al., 2022). Flavonoids typically undergo glycosylation. O-glycosides, C-glycosides, and O—C—glycosides are formed when the glycoside residues are connected to the O and C atoms of the flavonoids. Hexoses (162 Da), deoxyhexoses (146 Da), pentoses (132 Da), and an aglycone ion are the neutral species that result from the usual fragmentation of O-glycosides. As opposed to this, C-glucosides result in a series of fragments due to the cleavage of the C—C bonds with the sugar moiety. Some examples of these fragments include [M−H−60]−, [M−H−90]−, and [M−H−120]−, which are used as the distinctive diagnostic ions of glycone (Vukics and Guttman 2010, Kachlicki et al., 2016). Compounds 22–29 were identified as eriodictyol, catechin, dactylorhin C, taxifolin-7-sulfate, diadzin, cajanone, phenethylrutinoside, and kaempferol glucoside respectively, based on the similarities noticed in their fragmentation behaviors and the behaviors mentioned in the literature (Alam et al., 2021, Islam et al., 2021, Choi et al., 2022).
3.6.3 Alkaloids
APO contains oleraceins, a type of indoline amide glycosides. Many of these compounds are glucosylated and have 5,6-dihydroxyindoline-2-carboxylic acid N-acylated with cinnamic acid derivatives such as hydroxybenzoyl, coumaroyl, caffeoyl, feruloyl, and sinapoyl. The following fragment ions indicate the types of hydroxy cinnamic acid N-linked to the indoline core, at m/z 340.08, 356.07, and 370.09 for coumaroyl, caffeoyl and feruloyl, respectively. Furthermore, oleraceins also yielded characteristics ions by neutral loss of CO (28 Da), COO (44 Da), hydroxybenzoyl (120 Da), coumaroyl (146 Da), caffeoyl (162 Da), feruloyl (176 Da), sinapoyl (206 Da), glucosyl (162 Da), double glucosyl (324 Da) and triple glucosyl (486 Da). The first oleracein compound found in this study was oleracein E (compound 30), which has the chemical formula C12H13NO3 and produces a deprotonated ion at m/z 218.0833. The loss of an HCHO molecule and an HCOCH = CH2 molecule, respectively, resulted in the production of the fragment ions with m/z 188.07 and 162.05 respectively. It also undergoes i-cleavage of the middle ring's phenyl and C—C connections, producing fragment ions with m/z values of 135.04 and 121.02, respectively (Fig. 3C). Moreover, compounds 36–44 were identified as oleracein U, A, B, C, I, P, N/S, L/J, and O, respectively, based on commonalities seen between their fragmentation behaviors and those reported in the literature (Voynikov et al., 2021).
Furthermore, hydroxycinnamic acid amide yielded the base ion at m/z 147.04 (coumaroyl), m/z 163.04 (caffeoyl), m/z 177.05 (feruloyl) and m/z 207.06 (sinapoyl) by elimination of tyramine (137 Da) moiety. Further fragmentation was generated by the loss of a molecule of CO from the base peak. In addition, the tyramine moiety was further loss of NH3 to yield ion at m/z 121 (Liu et al., 2021). On the basis of the above fragmentation behavior, compound 32–36 was identified as feruloylglycine, coumaroyltyramine, caffeoyltyramine, feruloyltyramine and feruloyloctopamine, respectively (Zhou et al., 2015).
3.6.4 Carboxylic acids, fatty acids and amino acids
From comparisons of the mass and the fragmentation behaviors of the precursor ion based on mass spectroscopic analysis reported in literature and various online databases, compounds 45–49, and 52–64 were identified as carboxylic acids, and fatty acids, respectively (Table 5). In addition, compound 67–74 were characterized as amino acids (Guijas et al., 2018, Naveja et al., 2018, Nematallah et al., 2018, Ruan et al., 2019, Islam et al., 2020, Alam et al., 2021, Najm et al., 2021, Wang et al., 2021). Furthermore, molecule 50, 51, 65, and 66 were recognized as jasmonic acid and its derivatives (tuberonic acid, tuberonic acid glucoside, and methyl tuberonic acid glucoside) based on the mass fragmentation behavior described by Quirantes-Pine et al (Quirantes-Piné et al., 2010) (Fig. 4A). The jasmonic acid and its all derivaties are discovered for the first time in APO. EF. elemental formula; OM. observed mass; CM. calculated mass; CL. confidence level; (-). Negative mode. # First time identification in Portulaca oleracea.
No.
Compound name
EF
OM (m/z)-
CM (m/z)-
MS/MS (negative mode)
CL
Phenolic acid
1
4-Hydroxy benzoic acid
C7H6O3
137.0253
137.0244
119.03, 93.01
2
2
Coumaric acid
C9H8O3
163.0402
163.0401
119.04,
2
3
Ferulic acid
C10H10O4
193.0521
193.0506
179.03, 149.06, 135.04
2
4
Maclurin#
C13H10O6
261.0401
261.0405
151.00, 107.01
3
5
Caffeic acid phenethyl ester
C17H16O4
283.0967
283.097
265.08, 239.07, 179.03, 163.04, 135.04
3
6
Uralenneoside#
C12H14O8
285.0612
285.0616
153.01, 109.02
3
7
Ellagic acid
C14H6O8
300.9893
300.9984
283.99, 245.00, 229.01, 200.01, 185.02
2
8
Protocatechuic acid glucoside
C13H16O9
315.072
315.0716
162.02, 153.01
2
9
Barceloneic acid B#
C16H14O8
333.0618
333.0618
289.07, 271.06, 167.03
3
10
Coumaroylquinic acid
C16H18O8
337.0924
337.0626
191.05, 163.03
2
11
Caffeic acid hexoside
C15H18O9
341.1084
341.0872
215.03, 179.06, 161.04, 135.04
2
12
Ferulylshikimic acid
C17H18O8
349.0927
349.0923
193.05, 177.01, 173.04, 155.03, 129.02
2
13
Ferulic acid hexoside
C15H16O10
355.0666
355.0665
193.05, 179.02, 149.05, 134.02
2
14
Syringoylquinic acid
C16H20O10
371.0981
371.0978
353.08, 191.05, 173.04, 135.04
2
15
caffeic acid derivatives
C18H18O9
377.0885
377.0878
341.11, 215.03, 179.05, 161.03
2
16
Feruloyl-galactaric acid
C16H18O11
385.0828
385.0776
341.08, 209.03, 191.03, 147.02
2
17
Picraquassioside A#
C18H22O10
397.1142
397.1135
235.06, 217.05, 191.07, 187.04, 177.05
3
18
Methylpicraquassioside A#
C19H24O10
411.1302
411.1297
397.11, 249.07, 231.06, 219.06, 201.05
3
19
Paederol B#
C20H28O12
459.1511
459.1508
399.12, 341.12, 281.10, 193.05
3
20
Hexosyl caffeoyl hexose
C21H28O14
503.1393
503.1401
341.0.08, 179.03, 161.02
2
21
Arillatose B#
C22H29O14
517.1563
517.1557
313.05, 193.05
3
Flavonoids
22
Eriodictyol
C15H12O6
287.0565
287.0555
179.01, 151.00, 135.04, 125.01, 107.03
2
23
Catechin
C15H14O7
289.0721
289.0712
245.08, 205.05, 179.03, 135.04
2
24
Dactylorhin C
C14H24O10
351.1293
351.1291
189.07, 179.05, 171.06, 163.06, 127.07
2
25
Taxifolin-7-sulfate
C15H12O10S
383.0105
383.0079
303.05, 285.04, 275.05, 151.00, 125.03
2
26
Daidzin
C21H20O9
415.1029
415.1035
253.05, 235.04, 225.02, 135.00, 119.05
2
27
Cajanone
C25H26O6
421.1662
421.1651
383.12, 217.05, 197.09, 165.07, 151.00
2
28
Phenethylrutinoside
C20H30O10
429.1767
429.1761
249.09, 205.01, 161.04, 145.05, 119.05,
2
29
Kaempferol-3-O-glucoside
C21H20O11
447.0932
447.0928
285.04, 271.06, 256.02, 240.04, 151.00
2
Alkaloids
30
Oleracein E
C12H13NO3
218.0833
218.0823
200.07, 190.08, 160.04, 121.02
2
31
Feruloylglycine
C12H13NO5
250.0724
250.0721
206.08, 191.07, 177.05, 149.06
2
32
Coumaroyltyramine
C17H17NO3
282.1128
282.1123
279.01, 162.03, 145.03, 134.02, 119.02
2
33
Caffeoyltyramine
C17H17NO4
298.1085
298.1079
280.09, 178.05, 160.04, 136.07, 121.06
2
34
Feruloyltyramine
C18H19NO4
312.1235
312.1241
177.05, 149.06, 136.07, 121.06, 119.05
2
35
Feruloyloctopamine
C18H19NO5
328.1181
328.1184
310.02, 161.05, 133.02
2
36
Oleracein U
C18H15NO6
340.0831
340.0827
322.07, 296.09, 194.04, 145.02, 132.04
2
37
Oleracein A
C24H25NO11
502.1351
502.1349
340.08, 296.09, 194.05, 145.02
2
38
Oleracein B
C25H27NO12
532.1463
532.1455
370.09, 326.10, 194.05, 175.04, 161.02
2
39
Oleracein C
C30H35NO16
664.1883
664.1877
502.13, 340.08, 296.09, 194.04, 145.02
2
40
Oleracein I
C31H37NO17
694.1989
694.1983
518.15, 370.09, 326.10, 194.04, 175.04
2
41
Oleracein P
C36H45NO21
826.2395
826.2405
664.18, 502.13, 340.08, 194.04, 145.02
2
42
Oleracein N/S
C40H43NO19
840.2336
840.2351
694.19, 664.18, 340.08, 194.04, 145.02
2
43
Oleracein L/J
C40H43NO20
856.2276
856.2300
694.19, 518.15, 326.10, 194.04, 161.02
2
44
Oleracein O
C41H45NO20
870.2443
870.2456
694.19, 518.15, 194.04, 175.04, 161.02
2
Fatty acids
45
Citramalic acid
C5H8O5
147.0319
147.0299
129.01, 115.00, 103.04
2
46
2-Isopropylmalic acid
C7H12O5
175.0625
175.0612
157.05, 115.04, 113.06
2
47
Citric acid
C6H8O7
191.0217
191.0197
145.01, 129.01, 111.00
2
48
Oxaloglutaric acid
C7H8O7
203.0189
203.0197
141.01, 97.02, 69.03
2
49
Homocitric acid
C7H10O7
205.0351
205.0354
161.04, 143.04, 117.05
2
50
Jasmonic acid#
C12H18O3
209.1176
209.1183
165.09, 133.01, 109.03
3
51
Tuberonic acid#
C12H18O4
225.1125
225.1127
207.10, 181.12, 163.11, 135.08
3
52
Palmitic acid
C16H32O2
255.2314
255.2324
237.22, 211.24, 195.21, 59.01
2
53
2-Hydroxypalmitic acid
C16H32O3
271.2274
271.2273
253.21, 227.12,
2
54
Linolenic acid
C18H30O2
277.2165
277.2169
259.20, 233.22, 205.21, 179.25, 165.23
2
55
alpha-Linoleic acid
C18H32O2
279.2331
279.233
261.22
2
56
Oleic acid
C18H34O2
281.2487
281.2486
263.25, 181.21, 127.25
2
57
Stearic acid
C18H36O2
283.2643
283.2637
265.24, 239.25, 209.22, 183.19, 171.12
2
58
Hydroxy octadecatrienoic acid
C18H30O3
293.2112
293.2116
275.20, 223.03, 195.13, 183.13, 171.10
2
59
Hydroxy octadecadienoic acid
C18H32O3
295.2312
295.2276
277.20, 253.02, 223.03, 167.05
2
60
Hydroxy octadecenoic acid
C18H34O3
297.2433
297.2429
279.23, 255.12, 225.05, 127.05
2
61
Arachidonic acid
C20H32O2
303.2326
303.2324
285.22, 269.19, 259.24, 205.12
2
62
Dihydroxy octadecatrienoic acid
C18H30O4
309.2075
309.2069
291.19, 199.85, 179.14, 110.03
2
63
Trihydroxy-octadecadienoic acid
C18H32O5
327.217
327.2171
299.12, 285.21, 229.14, 211.13, 171.10
2
64
Pinellic acid#
C18H34O5
329.2329
329.2328
229.14, 211.13, 171.10
3
65
Tuberonic acid glucoside#
C18H27O9
387.1656
387.1655
207.10, 163.11, 101.02
3
66
Methyl tuberonic acid glucoside#
C19H30O9
401.1823
401.1817
239.12, 221.11, 207.10, 163.06
3
Amino acids
67
Phenylalanine
C9H11NO2
164.0732
164.0717
147.04, 120.08
2
68
Tyrosine
C9H11NO3
180.0674
180.0666
163.04, 134.06
2
69
3,4-Dihydroxyphenylalanine
C9H11NO4
196.0571
196.0615
181.05, 152.07
2
70
N-acetyl phenylalanine
C11H13NO3
206.0816
206.0823
164.07, 147.04
2
71
N-Acetyl tyrosine
C11H13NO4
222.0766
222.0772
180.06, 178.08, 163.04
2
72
N-benzoylaspartic acid
C11H11NO5
236.0558
236.0564
218.05, 192.06, 174.05, 120.04, 115.00
2
73
N-glucosyl ethanolamine
C8H17NO7
238.0927
238.0932
220.08, 202.07, 139.00
2
74
N-Acetyl tryptophan
C13H14N2O3
245.0925
245.0932
203.08, 185.07, 170.06, 116.05,
2
Terpenoids
75
Triptophenolide A1#
C20H24O3
311.1682
311.1653
295.13, 283.16, 267.17, 251.14, 237.12,
3
76
Menthane-1,2,8,9-tetrol 2-glucoside#
C16H30O9
365.1807
365.1812
204.13, 186.12, 168.11
3
77
α,g-Onoceradienedione
C30H46O2
437.3426
437.342
219.17, 205.15
2
78
4,5-dioxo 10-epi-4,5-seco-γ-eudesmol 2′-O-acetyl-fucopyranoside#
C23H38O8
441.2508
441.2488
399.23, 253.18, 221.15, 191.14
3
79
Oleanolic acid
C30H48O3
455.353
455.3525
407.33, 391.30, 377.28, 363.26
2
Others
80
Glucose
C6H12O6
179.0572
179.0561
163.06, 147.06, 115.04
2
81
Psoralen
C11H6O3
185.025
185.0244
157.02, 141.03, 129.03, 115.01
2
82
Gluconic acid
C6H12O7
195.0522
195.051
177.01, 151.06, 129.02, 121.04
2
83
Ethyl glucoside
C8H16O6
207.0854
207.0847
179.05, 163.06, 147.06, 115.04
2
84
Bargapten
C12H8O4
215.0348
215.0344
185.02, 157.02, 141.03, 129.03, 115.01
2
85
Glucosylglycolate
C8H14O8
237.0619
237.0616
220.05, 207.05, 193.07,163.06, 147.02
2
86
Oxyresveratrol
C14H12O4
243.0656
243.0663
225.05, 199.05, 161.06, 135.04
2
87
2-deoxy-2,3-dehydro-N-acetylneuraminic acid#
C11H17NO8
290.0876
290.0876
230.06, 200.05, 169.01, 128.07
3
88
Diphyllin
C21H16O7
379.0823
379.0817
363.05, 347.01, 333.04, 319.06, 305.04
2
89
Piceatannol glucoside
C20H22O9
405.1172
405.1178
243.06, 201.05, 159.04
2
90
Benzyl alcohol dihexoside
C19H28O11
431.1564
431.1553
341.10, 269.10, 251.09, 179.05, 163.06
2
91
Daphylloside
C19H26O12
445.135
445.1346
409.11, 387.12, 267.08, 179.05
2
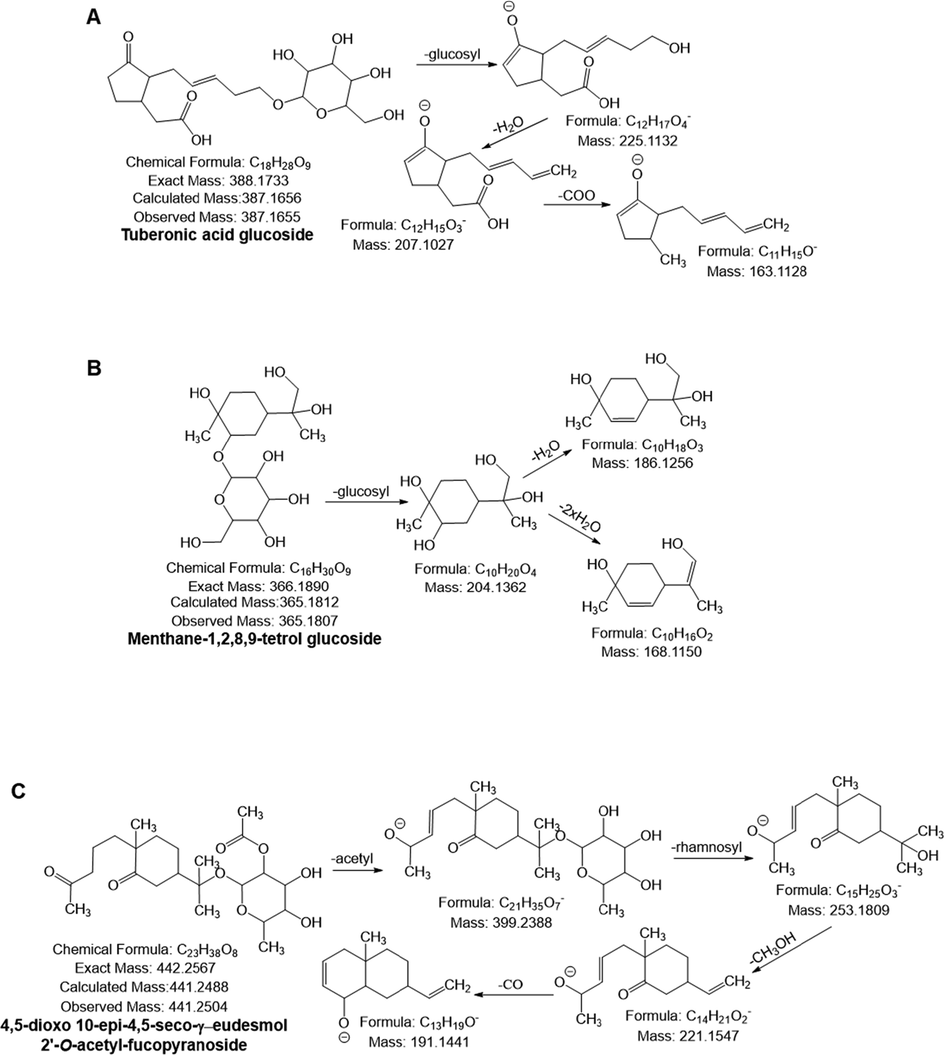
Possible mass fragmentation pattern of (A) tuberonic acid glucoside, (B) menthane-1,2,8,9-tetrol glucoside and (C) 4–5-dioxo 10-epi-4,5-seco-γ-eudesmol 2′-O-acetyl-fucopyranoside.
3.6.5 Terpenoids
For the first time in APO, compounds 75–79 was tentatively identified as terpenoids (Table 5). Compounds 75 was identified as triptophenolide A1 (m/z 311.1682) with molecular formula C20H24O3, based on the mass fragmentation behaviour described by Li et al., (Li et al., 2008). Compounds 76 produced the deprotonated ion [M−H]- at m/z 365.1812, yielded a fragment ion at m/z 204.13 by losing the glucosyl (162 Da) moiety, which was followed by the loss of one and two molecules of H2O to form the fragment ions at m/z 186.12 and 168.11, respectively. Accordingly, the compound was tentatively identified as menthane-1,2,8,9-tetrol glucoside (Fig. 4B) (Matsumura et al., 2002). Moreover, compound 78 generated a monoisotopic mass [M−H]- at m/z 441.2504, yielded fragment ions at m/z 399.23 ([M−H−acetyl]-), 253.18 ([M−H−acetyl−rhamnosyl]-), 221.15 ([M−H−253.18-CH3OH]-) and 191.14 ([M−H−221.15-CO]-) and was tentatively identified as 4,5-dioxo 10-epi-4,5-seco-γ-eudesmol 2′-O-acetyl-fucopyranoside with molecular formula C23H38O8 (Fig. 4C).
3.6.6 Others
compounds 80–91 were identified as glucose (m/z 179.0572), psoralen (m/z 185.0250), gluconic acid (m/z 195.0522), ethyl glucoside (m/z 207.0854), bergapten (m/z 215.0384), glucosylglycolate (m/z 237.0619), oxyresveratrol (m/z 243.0656), 2-deoxy-2,3-dehydro-N-acetylneuraminic acid (m/z 290.0876), diphyllin (m/z 379.0823), piceatannol glucoside (m/z 405.1178), benzyl alcohol glucoside (m/z 431.1564) and daphylloside (m/z 445.1346), based on the similarities noticed in their fragmentation behaviors and the behaviors mentioned in the literature.
4 Conclusions
This work, which was the first investigation into optimizing the HRE conditions on APO using two modeling approaches (RSM and ANN), revealed the presence of phenolic acids, flavonoids, alkaloids, fatty acid and terpenoids, through high-resolution mass spectroscopy examination. The ANN model is more accurate and sophisticated than the RSM model, as evidenced by the fact that it had a higher R2 and lower RMSE, AAD, and SEP values than the latter. The ideal parameters were identified as 60 % ethanol, extraction time of 90.5 min of extraction time, and 50 °C of extraction temperature. The highest values of TPC, TFC, DPPH radical scavenging effect, and ascorbic acid equivalent FRAP were found as 8.23 ± 1.06 mgGAE/g, 43.12 ± 1.15 mgCAE/g, 43.01 ± 1.25 %, and 35.98 ± 0.19, respectively, under these circumstances. These results lead us to the conclusion that APO, a viable candidate for an antioxidant functional food, can be widely used commercially in the nutraceutical food and pharmaceutical industries.
CRediT authorship contribution statement
Fanar Alshammari: Methodology, Formal analysis, Investigation, Writing – original draft. Md Badrul Alam: Conceptualization, Investigation, Formal analysis, Project administration, Writing – review & editing. Marufa Naznin: Methodology, Investigation, Formal analysis. Sunghwan Kim: Conceptualization, Supervision, Writing – review & editing. Sang-Han Lee: Conceptualization, Methodology, Supervision, Funding acquisition, Project administration, Writing – review & editing.
Acknowledgments
This study was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (2020R1A2C2011495 and 2021R1IA1A01058062). Fanar Alshammaria received the financial support for his Ph.D. studying project from Education Ministry of Kingdom of Saudi Arabia (EMSA).
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Isolation and characterization of Cepa2, a natural alliospiroside A, from shallot (Allium cepa L. Aggregatum group) with anticancer activity. Plant Physiol. Biochem. : PPB.. 2017;116:167-173.
- [CrossRef] [Google Scholar]
- DNA protecting activities of Nymphaea nouchali (Burm. f) flower extract attenuate t-BHP-induced oxidative stress cell death through Nrf2-mediated induction of Heme Oxygenase-1 expression by activating MAP-kinases. Int. J. Mol. Sci.. 2017;18
- [CrossRef] [Google Scholar]
- High resolution mass spectroscopy-based secondary metabolite profiling of Nymphaea nouchali (Burm. f) stem attenuates oxidative stress via regulation of MAPK/Nrf2/HO-1/ROS pathway. Antioxidants. 2021;10:719.
- [Google Scholar]
- High resolution mass spectroscopy-based secondary metabolite profiling of Nymphaea nouchali (Burm. f) stem attenuates oxidative stress via regulation of MAPK/Nrf2/HO-1/ROS pathway. Antioxidants (Basel Switzerland). 2021;10
- [CrossRef] [Google Scholar]
- Response surface methodology (RSM)-based optimization of ultrasound-assisted extraction of Sennoside A, Sennoside B, Aloe-Emodin, Emodin, and Chrysophanol from Senna alexandrina (Aerial Parts): HPLC-UV and antioxidant analysis. Molecules (Basel, Switzerland).. 2022;27
- [CrossRef] [Google Scholar]
- Characterization of gallotannins and benzophenone derivatives from mango (Mangifera indica L. cv. 'Tommy Atkins') peels, pulp and kernels by high-performance liquid chromatography/electrospray ionization mass spectrometry. Rapid Commun. Mass Spectromet. : RCM.. 2004;18:2208-2216.
- [CrossRef] [Google Scholar]
- Optimization of the extraction conditions of Nypa fruticans Wurmb. using response surface methodology and artificial neural network. Food Chem.. 2022;381:132086
- [CrossRef] [Google Scholar]
- Comparison between artificial neural network and response surface methodology in the prediction of the parameters of heat set polypropylene yarns. J. Text. Inst.. 2015;106:417-430.
- [CrossRef] [Google Scholar]
- Prandtl number of optimum biodiesel from food industrial waste oil and diesel fuel blend for diesel engine. Fuel. 2021;285:119049
- [CrossRef] [Google Scholar]
- Effect of extraction solvent on total phenol content, total flavonoid content, and antioxidant activity of Limnophila aromatica. J. Food Drug Anal.. 2014;22:296-302.
- [CrossRef] [Google Scholar]
- Flavonoids extract from Portulaca oleracea L. induce Staphylococcus aureus death by apoptosis-like pathway. Int. J. Food Prop.. 2017;20:S534-S542.
- [Google Scholar]
- Hepatoprotective effects of Portulaca oleracea extract against CCl4-induced damage in rats. Pharm. Biol.. 2015;53:1042-1051.
- [CrossRef] [Google Scholar]
- Phytochemical composition and antioxidant activity of Portulaca oleracea: influence of the steaming cooking process. Foods (Basel, Switzerland). 2021;10
- [CrossRef] [Google Scholar]
- Optimization of the ethanol recycling reflux extraction process for saponins using a design space approach. PLoS One. 2014;9:e114300.
- [Google Scholar]
- METLIN: a technology platform for identifying knowns and unknowns. Anal. Chem.. 2018;90:3156-3164.
- [CrossRef] [Google Scholar]
- A survey on stock market prediction using various algorithms. Int. J. Comput. Technol. Appl.. 2014;5:530-533.
- [Google Scholar]
- Interaction of ethanol with biological membranes: the formation of non-bilayer structures within the membrane interior and their significance. J. Phys. Chem. B. 2009;113:1983-1992.
- [CrossRef] [Google Scholar]
- Effects of aqueous Purslane (Portulaca oleracea) extract and fish oil on gentamicin nephrotoxicity in albino rats. J. Nat. Sci.. 2011;9:47-62.
- [Google Scholar]
- RSM and ANN modeling-based optimization approach for the development of ultrasound-assisted liposome encapsulation of piceid. Ultrason. Sonochem.. 2017;36:112-122.
- [Google Scholar]
- A review of traditional uses, phytochemistry and pharmacology of Portulaca oleracea L. J. Ethnopharmacol.. 2017;205:158-172.
- [CrossRef] [Google Scholar]
- Identification of secondary metabolites in Averrhoa carambola L. bark by high-resolution mass spectrometry and evaluation for α-glucosidase, tyrosinase, elastase, and antioxidant potential. Food Chem.. 2020;332:127377
- [CrossRef] [Google Scholar]
- Metabolite profiling of Manilkara zapota L. leaves by high-resolution mass spectrometry coupled with ESI and APCI and in vitro antioxidant activity, α-Glucosidase, and Elastase inhibition assays. Int. J. Mol. Sci.. 2021;22:132.
- [Google Scholar]
- Profile and characterization of the chlorogenic acids in green Robusta coffee beans by LC-MS n: identification of seven new classes of compounds. J. Agric. Food Chem.. 2010;58:8722-8737.
- [Google Scholar]
- Structural characterization of flavonoid Glycoconjugates and their derivatives with mass spectrometric techniques. Molecules. 2016;21
- [CrossRef] [Google Scholar]
- Protective effects of aqueous and ethanolic extracts of Nigella sativa L. and Portulaca oleracea L. on free radical induced hemolysis of RBCs. Daru. 2011;19:295-300.
- [Google Scholar]
- Antiulcerogenic effect of ethanolic extract of Portulaca oleracea experimental study. Pharmacologyonline. 2010;1:417-432.
- [Google Scholar]
- Evaluation of extract of Ipomoea batatas leaves as a green coagulant–flocculant for turbid water treatment: Parametric modelling and optimization using response surface methodology and artificial neural networks. Environ. Technol. Innov.. 2021;24:102005
- [CrossRef] [Google Scholar]
- A comparative evaluation of statistical empirical and neural intelligence modeling of Manihot esculenta-derived leaves extract for optimized bio-coagulation-flocculation of turbid water. Ind. Crop. Prod.. 2022;186:115194
- [CrossRef] [Google Scholar]
- Analysis of triptophenolide and its related compounds from Tripterygium wilfordii Hook.f by electrospray ionization tandem mass spectrometry. Int. J. Mass Spectrom.. 2008;278:38-49.
- [CrossRef] [Google Scholar]
- Identification of α-glucosidase inhibitors from Cortex Lycii based on a bioactivity-labeling high-resolution mass spectrometry–metabolomics investigation. J. Chromatogr. A. 2021;1642:462041
- [Google Scholar]
- Comparative metabolite fingerprinting of legumes using LC-MS-based untargeted metabolomics. Food Res. Int. (Ottawa, Ont.). 2019;126:108666
- [CrossRef] [Google Scholar]
- Reflux extraction optimization and antioxidant activity of phenolic compounds from Pleioblastus amarus (Keng) Shell. Molecules (Basel, Switzerland). 2022;27
- [CrossRef] [Google Scholar]
- Bronchodilatory effect of Portulaca oleracea in airways of asthmatic patients. J. Ethnopharmacol.. 2004;93:57-62.
- [Google Scholar]
- Water-soluble constituents of caraway: carvone derivatives and their glucosides. Chem. Pharm. Bull.. 2002;50:66-72.
- [CrossRef] [Google Scholar]
- Identification of phytochemicals of Phoenix dactylifera L. Cv Ajwa with UHPLC-ESI-QTOF-MS/MS. Int. J. Fruit Sci.. 2021;21:848-867.
- [CrossRef] [Google Scholar]
- Naveja, J. J., M. P. Rico-Hidalgo and J. L. Medina-Franco, 2018. Analysis of a large food chemical database: chemical space, diversity, and complexity. F1000Research. 7, Chem Inf Sci-993. https://doi.org/10.12688/f1000research.15440.2.
- Evaluation of potential antifertility activity of total flavonoids, isolated from Portulaca oleracea L on female albino rats. Int. J. PharmTech. Res.. 2014;6:783-793.
- [Google Scholar]
- Polyphenols LC-MS2 profile of Ajwa date fruit (Phoenix dactylifera L.) and their microemulsion: potential impact on hepatic fibrosis. J. Funct. Foods. 2018;49:401-411.
- [CrossRef] [Google Scholar]
- Development of ternary models for prediction of biogas yield in a novel modular biodigester: a case of fuzzy Mamdani model (FMM), artificial neural network (ANN), and response surface methodology (RSM) Biomass Convers. Biorefin. 2020
- [CrossRef] [Google Scholar]
- Estimation of biogas yields produced from combination of waste by implementing response surface methodology (RSM) and adaptive neuro-fuzzy inference system (ANFIS) Int. J. Energy Environ. Eng.. 2021;12:353-363.
- [CrossRef] [Google Scholar]
- Predictive ability of Response Surface Methodology (RSM) and artificial Neural Network (aNN) to approximate Biogas Yield in a Modular Biodigester. Springer International Publishing; 2021.
- Optimization of extraction conditions for colchicine from Gloriosa superba tubers using response surface methodology. J. Agric. Technol.. 2012;8:1301-1315.
- [Google Scholar]
- Isolation, identification and quantitation of hydroxycinnamic acid conjugates, potential platform chemicals, in the leaves and stems of Miscanthus× giganteus using LC–ESI-MSn. Phytochemistry. 2011;72:2376-2384.
- [Google Scholar]
- Phytochemical composition and bioactive compounds of common purslane (Portulaca oleracea L.) as affected by crop management practices. Trends Food Sci. Technol.. 2016;55:1-10.
- [Google Scholar]
- Characterization of phenolic and other polar compounds in a lemon verbena extract by capillary electrophoresis-electrospray ionization-mass spectrometry. J. Sep. Sci.. 2010;33:2818-2827.
- [CrossRef] [Google Scholar]
- Comprehensive chemical profiling in the ethanol extract of Pluchea indica aerial parts by liquid chromatography/mass spectrometry analysis of its silica gel column chromatography fractions. Molecules (Basel, Switzerland). 2019;24
- [CrossRef] [Google Scholar]
- Response surface optimisation of antioxidant extraction from kinema, a Bacillus-fermented soybean food. Food Chem.. 2011;129:507-513.
- [CrossRef] [Google Scholar]
- Portuloside A, a monoterpene glucoside, from Portulaca oleracea. Phytochemistry. 1996;42:1625-1628.
- [Google Scholar]
- The development of expeller for kernel based seed and its oil characterization. Int. J. Integr. Eng.. 2018;10
- [Google Scholar]
- Comparison of Response Surface Methodology (RSM) and Artificial Neural Network (ANN) in modelling of waste coconut oil ethyl esters production. Energy Sources Part A. 2019;41:1049-1061.
- [CrossRef] [Google Scholar]
- Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ. Sci. Tech.. 2014;48:2097-2098.
- [CrossRef] [Google Scholar]
- Optimization of the ultrahigh-pressure–assisted extraction of phenolic compounds and antioxidant activity from palm dates (Phoenix dactylifera L.) Food Anal. Methods. 2020;13:1556-1569.
- [CrossRef] [Google Scholar]
- Properties of Portulaca oleracea seed oil via supercritical fluid extraction: experimental and optimization. J. Supercrit. Fluids. 2018;135:34-44.
- [Google Scholar]
- Optimization of fatty acids extraction from Portulaca oleracea seed using response surface methodology. Ind. Crop. Prod.. 2013;43:405-411.
- [Google Scholar]
- Analysis of chemical constituents in Ficus Hirta Vahl. by LCMS-IT-TOF and GC-MS. IOP Conf. Ser.: Mater. Sci. Eng.. 2020;730:012027
- [CrossRef] [Google Scholar]
- UHPLC-Orbitrap-MS tentative identification of 51 Oleraceins (Cyclo-Dopa Amides) in Portulaca oleracea L. cluster analysis and MS(2) filtering by mass difference. Plants (Basel Switzerland). 2021;10
- [CrossRef] [Google Scholar]
- Structural characterization of flavonoid glycosides by multi-stage mass spectrometry. Mass Spectrom. Rev.. 2010;29:1-16.
- [CrossRef] [Google Scholar]
- Protective effect of Portulaca oleracea extracts on hypoxic nerve tissue and its mechanism. Asia Pac. J. Clin. Nutr.. 2007;16:227.
- [Google Scholar]
- Optimization of ultrasonic-assisted extraction of flavonoid from Portulaca oleracea L. by response surface methodology and chemical composition analysis. J. Korean Soc. Appl. Biol. Chem.. 2014;57:647-653.
- [Google Scholar]
- CFM-ID 4.0: more accurate ESI-MS/MS spectral prediction and compound identification. Anal. Chem.. 2021;93:11692-11700.
- [CrossRef] [Google Scholar]
- Optimization of ultrahigh-pressure extraction of polyphenolic antioxidants from green tea by response surface methodology. Food Bioproc. Tech.. 2013;6:2538-2546.
- [CrossRef] [Google Scholar]
- Homoisoflavonoids from the medicinal plant Portulaca oleracea. Phytochemistry. 2012;80:37-41.
- [Google Scholar]
- Exergy and energy analyses of microwave Dryer for Cantaloupe slice and prediction of thermodynamic parameters using ANN and ANFIS algorithms. Energies. 2021;14:4838.
- [Google Scholar]
- Zhou, Y. X., H. L. Xin, K. Rahman, et al., 2015. Portulaca oleracea L.: a review of phytochemistry and pharmacological effects. BioMed research international. 2015, 925631. https://doi.org/10.1155/2015/925631.
- Antioxidant activities and total phenolic contents of various extracts from defatted wheat germ. Food Chem.. 2011;126:1122-1126.
- [CrossRef] [Google Scholar]
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2022.104425.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







