Translate this page into:
Phytochemical inspection and anti-inflammatory potential of Euphorbia milii Des Moul. integrated with network pharmacology approach
⁎Corresponding authors. walaa.negm@pharm.tanta.edu.eg (Walaa A. Negm), rabinsuwaidan@pnu.edu.sa (Reem Binsuwaidan),
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
The inflammatory reaction is induced by several factors like stimulation of different enzymes, oxidative stress, and production of various proinflammatory mediators. Here, we elucidated the phytochemical profile of Euphorbia milii ethanol extract (EMEE) then, we studied its anti-inflammatory potential using a carrageenan-induced paw edema model in rats. It was found that the average paw edoema weight substantially declined (p < 0.05) by treatment with EMEE (100 mg/kg and 200 mg/kg). Also, it was noticed the immunostaining with cyclooxygenase-2 and tumor necrosis factor-alpha antibodies decreased to a mild positive (score 1) after treatment with EMEE (200 mg/kg). This is along with improvement in the histological findings after examination of the paw tissues with hematoxylin and eosin as it revealed a decrease in the inflammation. Also, EMEE (200 mg/kg) resulted in a significant decline (p < 0.05) in the levels of the proinflammatory mediators (granulocyte–macrophage colony-stimulating factor, monocyte chemoattractant protein-1, inducible nitric oxide synthase, and interleukin-5). Interestingly, it resulted in a substantial increase (p < 0.05) in the levels of the anti-inflammatory interleukins (IL-10 and IL-12). Using the Swiss Target Prediction database, 127 genes were related to the identified metabolites in EMEE, of which 55 target genes were associated with 11 inflammatory disorders, as demonstrated by the DisGeNET database. The PI3K-Akt signaling route, the MAPK signaling pathway, and the Ras signaling pathway were identified as the dominant signaling pathways by the study of annotated genes. Therefore, EMEE requires further clinical investigations as an anti-inflammatory agent.
Keywords
Cyclooxygenase-2
Tumor necrosis factor-alpha
qRT-PCR
LC/MS
Network pharmacology
Signaling pathway
1 Introduction
The inflammatory process is carried out by the human cells as a defensive response against different inducers such as injury of tissues, specific chemical compounds, and infections. During this protective mechanism, particular reactions are provoked via mediators of the inflammatory response (Sychrová et al., 2020). Interleukin-5 (IL-5), interleukin-6 (IL-6), and tumor necrosis factor-alpha (TNF-α) are examples of these proinflammatory mediators (Kaneko et al., 2019) and they can launch and boost the inflammatory process. Moreover, certain enzymes are involved in this inflammatory process, such as cyclooxygenase-2 (COX-2) which can manage the prostaglandin (PGs) production (Rea et al., 2018). Also, reactive oxygen species (ROS) are considered important supporters of the inflammatory process. Such molecules are released by the inflammatory cells and can exaggerate the process of oxidative stress (Lin et al., 2021, Elekhnawy et al., 2021).
Unfortunately, finding safe and efficient anti-inflammatory compounds is challenging. Anti-inflammatory medications currently in use, such as non-steroidal anti-inflammatory agents, have numerous side effects (Bindu et al., 2020). These drugs, when used for a long term, can cause renal, gastrointestinal, and cardiac adverse reactions. Therefore, there is an urgent necessity to identify novel, effective anti-inflammatory products with no or relatively low side effects. Currently, natural agents, like plants, are regarded as a rich source for many bioactive compounds, and consequently, their therapeutic efficacy is being widely inspected. This is attributed to their numerous advantages like efficacy, safety, and biocompatibility (Attallah et al., 2022b).
With about 2000 species, the genus Euphorbia is a well-known angiosperm that may be found all over the world, especially in Asia, Africa, Central and South America. This genus's members range in size from tiny annual or perennial herbs to woody shrubs, vines, trees, and even succulent plants. They also produce an irritant milky latex. Generally, the entire plants, stems, leaves, latex, roots, and seeds of the Euphorbia species have been the subject of chemical and pharmacological investigation (Zhao et al., 2022, Xu et al., 2021).
Euphorbia roots or tubers are well-known for their therapeutic properties, particularly in China. Many of them are utilized as Chinese herbal medicines (Zhao et al., 2022, Salehi et al., 2019). Additionally, a recent study showed distinctions between the geographic distribution of Euphorbia plants used in traditional medicines around the world (Ernst et al., 2015). Three uses in particular were found to be the most frequently used in this regard: treating digestive system disorders (very common worldwide with Australasia being an exception); treating infections and infestations (primarily in Southern Africa and America, the Pacific, Asia-tropical, and Asia-temperate); and treating skin/subcutaneous cellular tissue disorders (especially in Australasia, Europe, Asia, and Northern America) (Salehi et al., 2019, Pascal et al., 2017).
One of the most significant medicinal plants in this genus is Euphorbia milii, sometimes known as “Christ thrown” or “Christ plant,” which has been used traditionally to cure hepatitis, cancer, and warts (Saleem et al., 2019, Yu et al., 2022). Additionally, this plant's antifungal, antinociceptive, antibacterial, and molluscicidal capabilities have been documented (Saleh et al., 2023b, Saleh et al., 2023a).
Network pharmacology is a new approach to virtual biological–chemical interactions that helps in identifying the highly represented genes related to identified or isolated compounds from a specific species in relation to a specific disease under study. It is used in different disorders like inflammatory disorders (Alnusaire et al., 2023), cancer (El-Hawary et al., 2023, Mohammed et al., 2023, Ahmed et al., 2022), wounds (Elmaidomy et al., 2023), neurodegenerative disorders (Bakhsh et al., 2023) and microbial infections (Alyami et al., 2023). Since this virtual technique can help to save money and effort for identifying target genes, biological pathways, cellular components, biological processes, and molecular functions of natural origin.
The objective of the current study was to look into the anti-inflammatory activity of EMEE in carrageenan-induced paw edoema after elucidating its secondary metabolites by LC/MS/MS. Also, network pharmacology studies would be employed to reveal the potential anti-inflammatory action of the phytochemicals of EMEE.
2 Materials and methods
Chemicals and plant materials
The utilized chemicals and plant were displayed in supplementary data.
LC-ESI-MS/MS profile analysis
Extract of E. milii aerial parts was analysed by adopting procedures previously reported by Attallah et al. (Attallah et al., 2022b).
Network pharmacology study
3.1 Compound targets for Euphorbia milli (supplementary data)
3.2 Metabolites-Targets network
The NCBI Gene databas (https://www.ncbi.nlm.nih.gov/gene/) (Boutet et al., 2016), also the Online SwissTargetPrediction database (Gfeller et al., 2014) were used to identify the related targets.
3.3 Targets-inflammation network
The target genes were specifically identified for the inflammatory disorders related to osteoarthritis, joints, and bone inflammations through the DisGeNET database (https://www.disgenet.org/search) (Piñero et al., 2020). The built in filter option in the database was used. The disease names were specified as ‘’Rheumatoid Arthritis’’, ‘’Inflammatory polyarthritis’’, ‘’ Bone Diseases’’, ‘’Inflammation’’, ‘’ Inflammatory pain’’, ‘’ inflammatory joint disease’’, ‘’ Arthritis’’, ‘’ Degenerative polyarthritis’’, ‘’ Arthritis, Adjuvant-Induced’’, ‘’ Osteoarthritis, Knee’’, ‘’ Arthritis, Reactive’’, ‘’Inflammation’’.
3.4 Protein–protein interaction (PPI) data (supplementary data)
3.5 Network construction
Using the network analysis programme Cytoscape, the resultant networks were visualised and reorganised (Kohl et al., 2011), version 3.9.0. The difference was considered statistically significant at p0.05.
3.6 Gene ontology and pathway analysis
ShinyGo database (Ge et al., 2020) (last accessed on November 25, 2022) and FunRich software (Pathan et al., 2017) were used to research the involved gene pathways, cellular components, molecular activities, and biological processes.
Anti-inflammatory activity of EMEE (supplementary data)
Statistics
The experimental studies were performed in triplicates and the obtained results are revealed as the mean ± standard deviation (SD). The utilized software was Graph-Pad prism version 8 and ANOVA test followed by post-hoc test was utilized. The significance level was recognized at p < 0.05.
3 Results
3.1 LC-ESI-MS/MS analysis
A total of 16 compounds were tentatively identified in E. milii using LC-ESI-MS/MS in a negative mode. The main substances were several types of flavonoids and their glycosidic derivatives, phenolics, and carboxylic acids. The comprehensive profile is listed in Table 1. While Fig. S1 shows the total ion chromatogram of detected metabolites in negative ion mode.
Metabolite Name
RT. (min.)
Molecular Formula
Precursor m/z
(Telesford et al., 2015)-
Error
MS/MS
Class
Homogenentisic acid
1.385
C8H8O4
167.03
−12.5
99.01,124.01, 125.02,152.01, 167.03
2(hydroxyphenyl) acetic acids
Quercitrin
7.274
C21H20O11
447.09
4.3
163.02, 256.93, 285.02, 296.91, 301.02, 334.90, 402.89, 447.08
Flavonoid-3-O-glycosides
Kaempferol-3-O-alpha-L-rhamnoside
7.964
C21H20O10
431.09
1.5
112.98, 227.03, 284.05, 431.09
Flavonoid-3-O-glycosides
Astragalin
8.877
C21H20O11
447.094
−0.5
447.09, 285.18, 284.03
Flavonoid-3-O-glycosides
l-Methyl-6-hydroxy1,2,3,4-Tetra hydro isoquinoline-3-carboxylic acid
10.370
C11H13NO3
206.082
−0.5
206.08
Isoquinoline-3-carboxylic acid
Baicalein-7-O-glucuronide
10.468
C21H18O11
445.1552
−7.7
261.08, 263.10, 445.12
Flavonoid-7-O-glucuronides
6,7-Dihydroxycoumarin
10.481
C9H6O4
177.05
−5.5
117.03, 119.00, 145.02, 177.04
6,7-dihydroxycoumarins
Quercetin-3-O-(6″-
galloyl)-β-galactopyranoide)10.770
C28H24O16
615.099
1.5151.01, 178,02, 301.04, 331.06, 615.09
Flavonoid-3-O-glycosides
Sinapyl aldehyde
10.852
C11H12O4
207.0665
−1.0
133.02, 149.02, 161.02, 177.01, 179.03, 188.93, 192.04, 207.06
Methoxyphenols
Luteolin
10.865
C15H10O6
285.18
5.7
285.04
Flavones
3′-methoxy-4′,5,7-trihydroxyflavonol
11.530
C16H12O7
315.04
3.6
243.01, 246.89, 269.25, 297.08, 300.02, 314.89
Flavonols
Gamma-Linolenic acid
11.875
C18H30O2
277.1451
−0.6
59.01, 102.95, 120.02, 121.03, 205.15, 208.92, 233.14, 277.17
Lineolic acids and derivatives
(+/-)-Taxifolin
13.559
C15H12O7
303.1588
1.8
190.95, 258.93, 285.14, 303.15
Flavanonols
3, 5, 7-Trihydroxy-4′-methoxyflavone
14.564
C16H12O6
299.09
42.3
177.02, 178.02, 281.21, 299.09
Flavones
3,4,5′-Trihydroxy-3′-glucopyranosylstilbene
19.296
C20H22O9
405.2002
0.5
405.16
Stilbene glycosides
7-Hydroxy-4-methylcoumarin
27.004
C10H8O3
174.9572
−5.5
86.98, 130.96, 146.96, 174.95
7-hydroxycoumarins
3.2 Network analysis
3.2.1 Plant –metabolite network analysis
Using LC/MS/MS analysis technique (negative mode), the identification of the active metabolites (16 metabolite) from Euphorbia milli was established. A simple illustrating network was constructed and visualized by Cytoscape software (Fig. 1).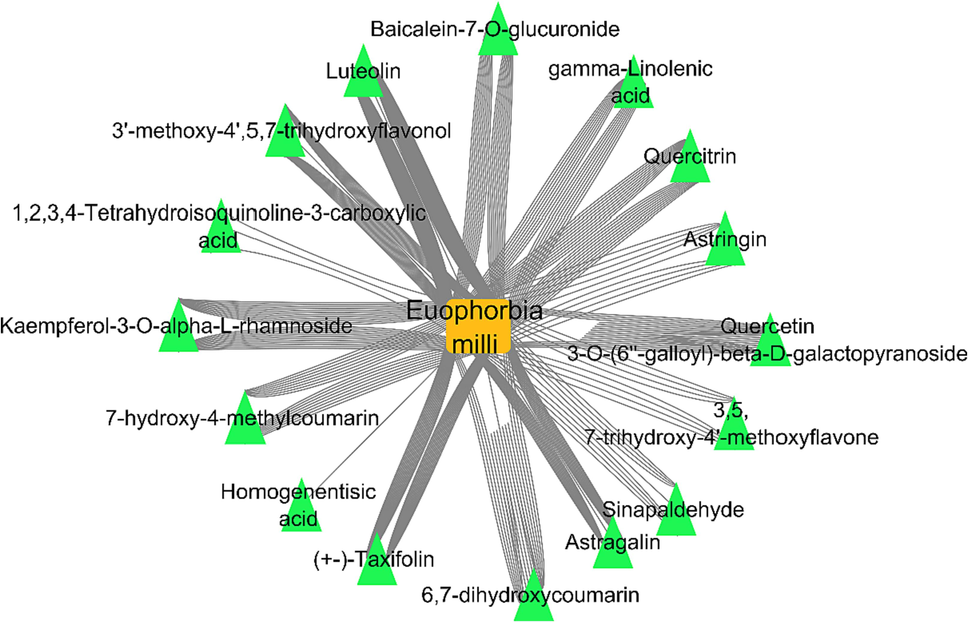
Plant-metabolite network; a network illustrates the identified compounds from Euphorbia mill plant, the orange square is the plant name, the green triangles represent the identified metabolites. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
3.2.2 Metabolite-target network analysis
The identified compounds of Euphorbia milli by LC/MS/MS analysis were introduced in PubChem database and Swiss Target Prediction in order to extract the target hits. Interestingly, a total of 127 target hits were obtained. A network of metabolites-target was established. The nodes of the network were 142 node for the metabolites and 127 node for the targets, with network heterogenicity of 1.682 (Fig. 2).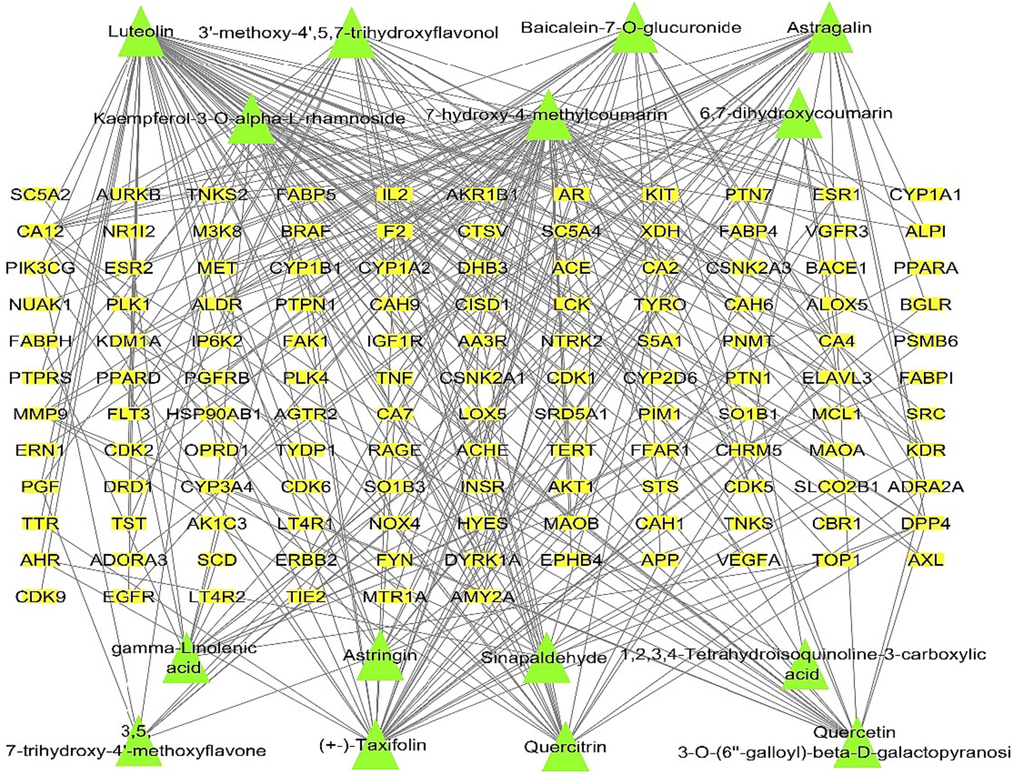
Metabolites-targets network; a network illustrating the targets of the identified compounds. Green triangles represent the identified compounds; yellow rectangles represent the targets. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
3.2.3 Protein–protein interaction (PPI) data
The PPI network was constructed with 76 nodes, 501 edges, and an average node degree of 9.19. The interactions between the target proteins were performed and visualized using the online STRING database (Fig. 3). With clustering coefficient of 2.095 and network centralization of 0.384 (Alqahtani et al., 2022).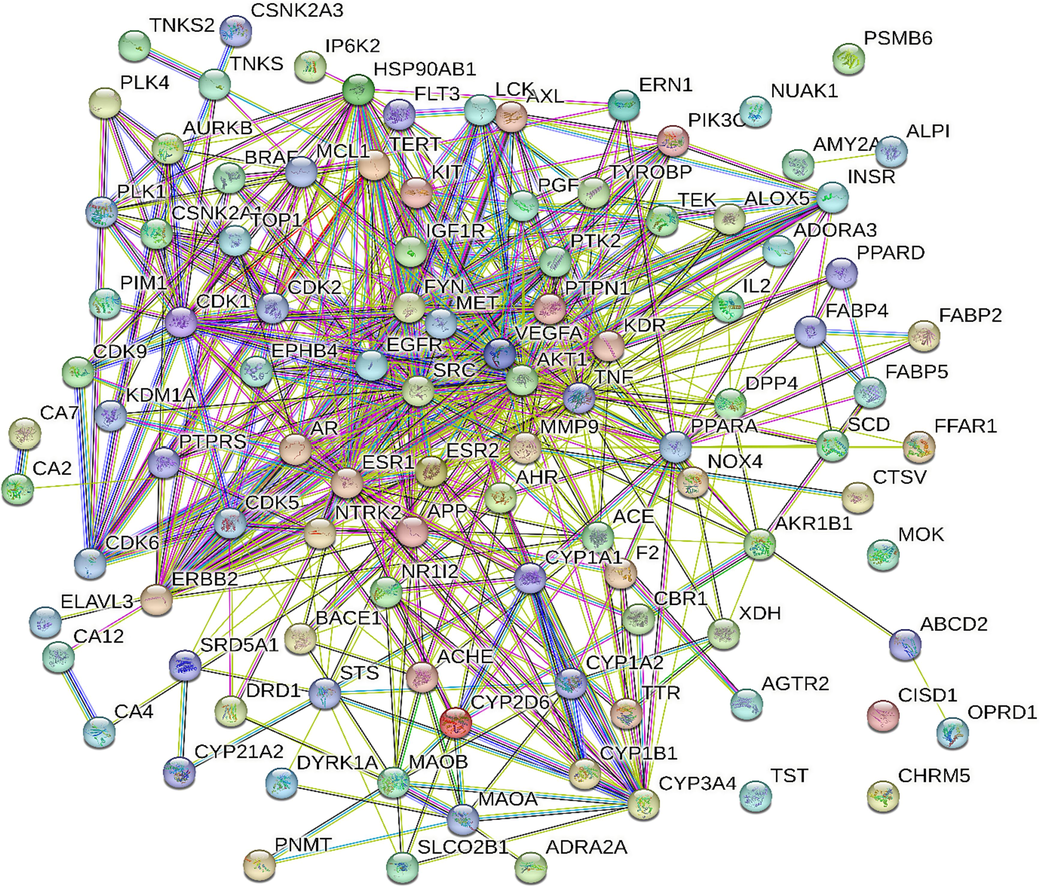
PPI network, illustrating the interaction between proteins of identified targets.
3.2.4 Common targets-inflammation network analysis
A network assessment was performed to establish the relations between the identified targets and the different kinds of inflammations. A sum of 55 genes was connected to 11 types of inflammation (Fig. 4) and the top revealed genes were TNF, VEGFA, PIK3CG, EGFR, and MMP9 with number of edges of 16, 8, 8,7, and 7, in the same order.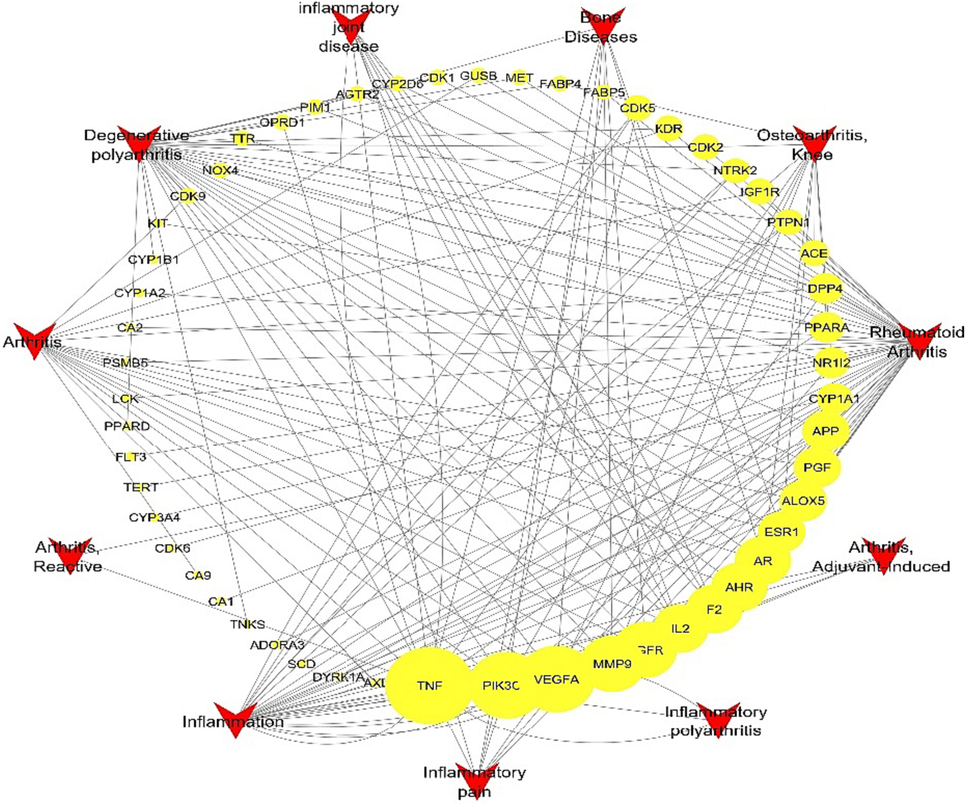
Targets-inflammation network, a network illustrating the relations between identified targets and inflammatory disorders, yellow circles represent the targets (higher size indicate higher representation), the red triangles represent the types of inflammatory disorders. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
3.2.5 Complete pharmacology network (Metabolites- targets-inflammation)
The whole pharmacology network was created by combining and integrating the networks (plant-metabolites, metabolite-targets, targets-inflammation) to analyse the interactive connections between the identified compounds from Euphorbia milli to the target proteins and to the inflammatory disorders (Fig. 5). This network showed the genes related to inflammation in this gene set and other genes not involved in inflammatory disorders.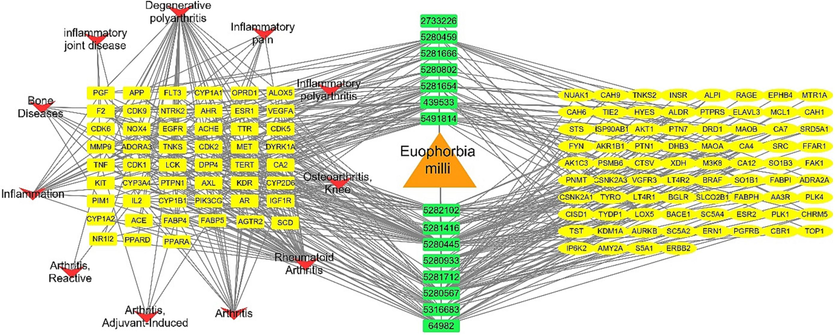
Complete pharmacology network analysis (Metabolites-targets-inflammation), orange triangle is the plant names, green rectangles represent compounds PCIDs, yellow oval shapes represent targets not involved in inflammation, yellow rectangles represent targets involved in inflammations, red arrow heads represent types of inflammation. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
3.2.6 Target genes-pathways analysis
The top cellular components identified with this gene through the FunRich software were: cytoplasm followed by plasma membrane, the top molecular functions were the catalytic activity followed by transmembrane receptor protein tyrosine kinase activity, while the top biological processes were signal transduction and cell communication (Fig. 6).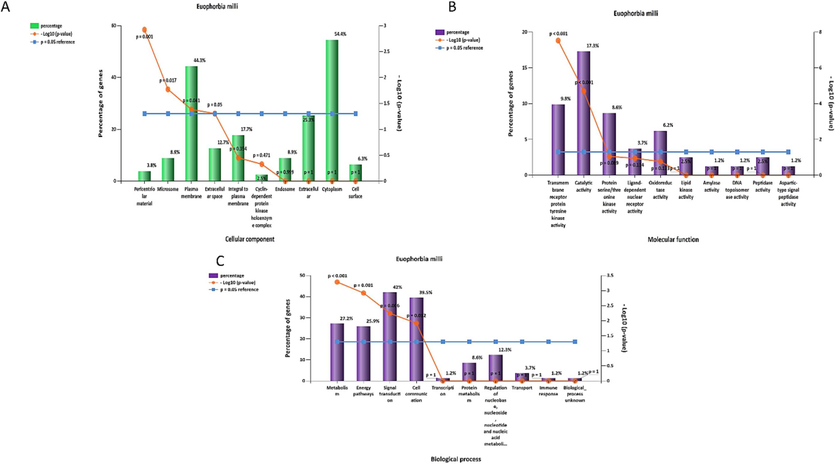
Gene enrichment analysis of the targets of Euphorbia milli; (A) cellular components, (B) molecular functions, (C) biological process.
We used the KEGG and ShinyGO databases to create the network of the hit genes targeted by the detected compounds interacting with the defined types of inflammatory illnesses in order to develop target genes-pathways networking. The PI3K-Akt signalling route, MAPK signalling pathway, and the Ras signalling pathways were the top signalling pathways identified by the databases (Fig. 7 & Table S2).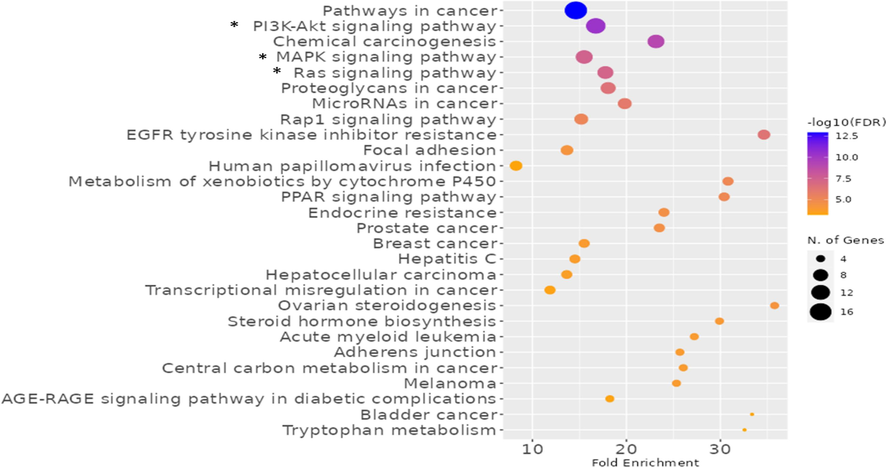
Biological pathways of the genes of Euphorbia milli related to inflammation disorders, the top signalling pathways marked with (*), the more intense colour the more no. of genes represented in the pathway.
3.3 In vivo study
The in vivo study was employed to reveal the potential anti-inflammatory action of EMEE in an animal model.
3.3.1 Effect of EMEE on the paw edema weight
The influence of the EMEE on the paw edoema average weight was inspected as revealed in Fig. 8. Compared to groups II and IV, group V's average paw edoema weight significantly decreased (p < 0.05). Regarding group II (carrageenan group), there was a significant increase in the edema weight by 925.64 % in comparison with group I. EMEE, with concentrations of 100 and 200 mg/kg, resulted in a marked decrease in the paw edema weight by 65 % and 80.75 %, respectively, in comparison with group II.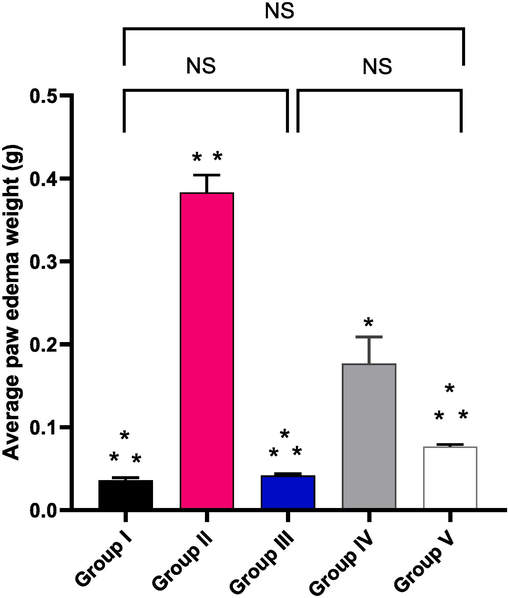
Average weight of the paw edema of the studied groups. The symbol (*) denotes a significant difference (p < 0.05) as regards to group II. The symbol (**) denotes a significant difference (p < 0.05) as regards to group IV. The abbreviation (NS) denotes non-significant difference among the groups (p > 0.05).
3.3.2 Histological studies
Tissue sections of the paws from the different studied groups were stained with H&E as exhibited in Fig. 9.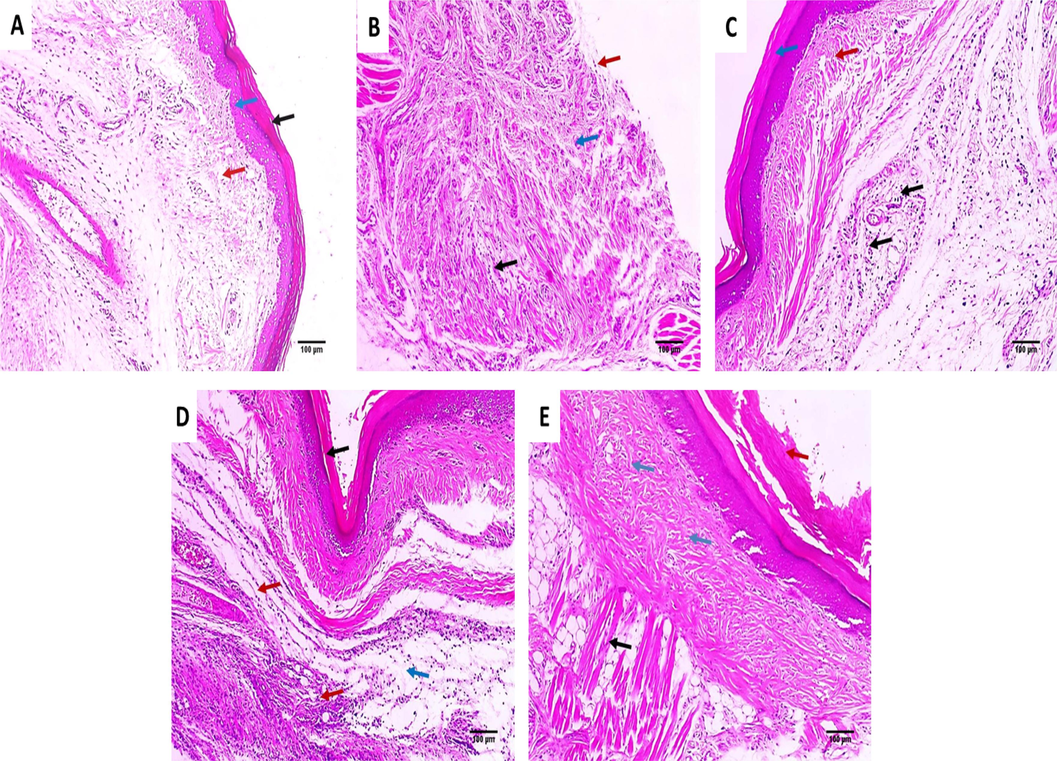
H&E stained paw tissue sections of: A) Group I presenting normal skin consisting of epidermis of average thickness (blue arrow) lined with thick keratin (black arrow) and underlying normal dermis (red arrow) (×100). B) Group II presenting surface ulceration (red arrow) with underlying granulation tissue (blue arrow) with infiltration by chronic inflammatory cells (black arrow) (×100). C) Group III presenting very mild dermal inflammation (black arrows). The epidermis is thickened and covered with excessive keratosis (blue arrow) with underlying mild collagenosis (red arrow) (×100). D) Group IV presenting epidermal healing with superficial dermal mild chronic inflammation (red arrows) and mild edema (blue arrows) (×100). E) Group V presenting absence of inflammation and the muscles are normal (black arrow). The epidermis is thickened and covered with excessive keratosis (red arrow) with underlying mild collagenosis (blue arrows) (×100). (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
3.3.3 Immunohistochemical studies
The COX-2 and TNF-α immunostaining of the paw tissues of the different studied groups is exhibited in Figs. 10 and 11.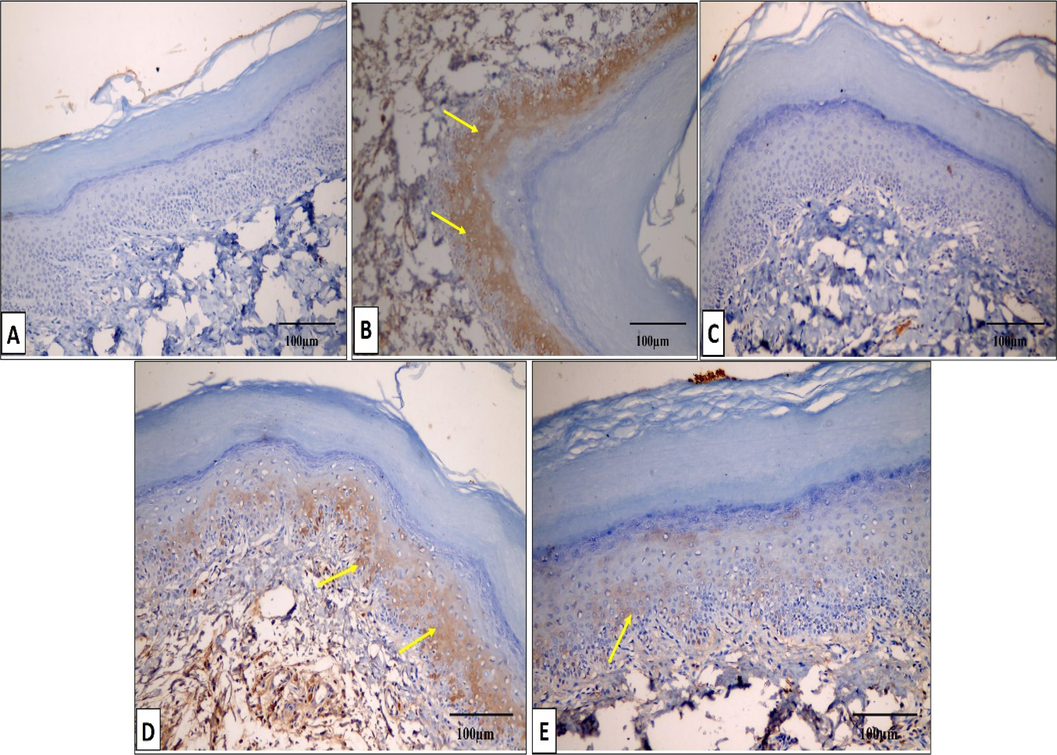
COX-2 immunostained paw tissue sections of: A) Group I presenting a negative COX-2 immunostaining (score 0). B) Group II presenting a strong positive COX-2 immunostaining (score 3). C) Group III presenting a negative COX-2 immunostaining (score 0). D) Group IV presenting a moderate positive COX-2 immunostaining (score 2). E) Group V presenting a mild positive COX-2 immunostaining (score 1). (×200, scale bar = 100 μm).
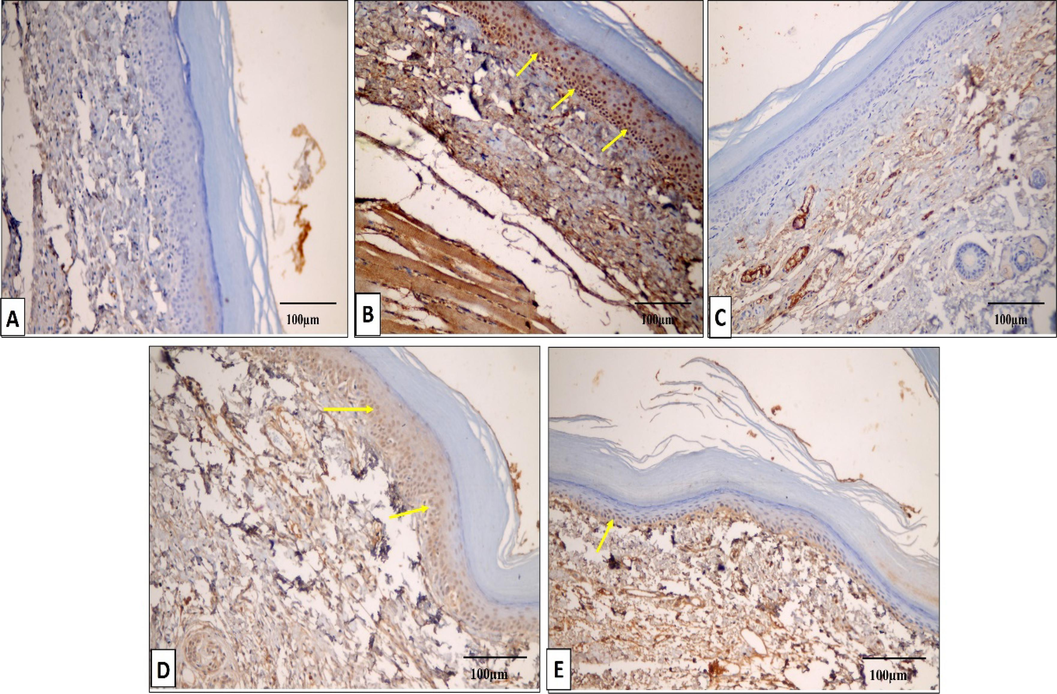
TNF-α immunostained paw tissue sections of: A) Group I presenting a negative TNF-α immunostaining (score 0). B) Group II presenting a strong positive TNF-α immunostaining (score 3). C) Group III presenting a negative TNF-α immunostaining (score 0). D) Group IV presenting a moderate positive TNF-α immunostaining (score 2). E) Group V presenting a mild positive TNF-α immunostaining (score 1). (×200, scale bar = 100 μm).
3.3.4 qRT-PCR
3.3.4.1 Gene expression of GM-CSF and MCP-1
The impact of EMEE on the relative gene expression of GM-CSF and MCP-1 was examined as revealed in Fig. 12. In comparison with group I, there was a significant overexpression of the GM-CSF and MCP-1 genes in group II by 380 % and 445 %, respectively. In addition, there was a significant downregulation in the gene expression of GM-CSF in groups IV and V by 58.13 % and 71.88 %, respectively, and the gene expression of MCP-1 in groups IV and V by 61.28 % and 73.03 %, respectively, in comparison with group II.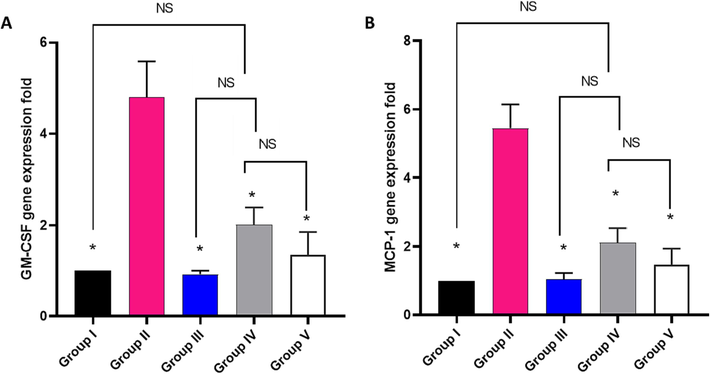
Impact of treatment with EMME on the gene expression of A) GM-CSF and B) MCP-1. The single asterisk represents a significant difference (p < 0.05) in comparison with group II. The abbreviation NS means non-significant difference (p > 0.05) in comparison with group IV.
3.3.4.2 Gene expression of IL-5, IL-10, and IL-12
The influence of EMEE on the relative gene expression of IL-5, IL-10, and IL-12 was studied as revealed in Fig. 13. Group II revealed a significant increase in the gene expression of IL-5 by 455 % and a marked decrease in the gene expression of IL-10 and IL-12 by 84.26 % and 83.7 %, respectively, in comparison with group I. Regarding IL-5, its expression significantly decreased in groups IV and V by 60.36 % and 75.86 %, respectively, relative to group II.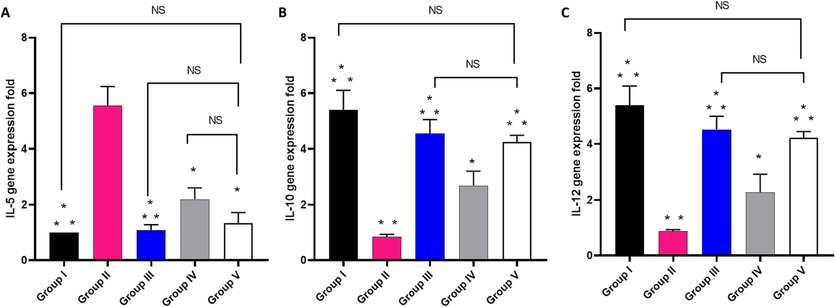
Impact of treatment with EMME on the gene expression of A) IL-5, B) IL-10, and C) IL-12. The single asterisk represents a significant difference (p < 0.05) in comparison with group II. The double asterisks represent a significant difference (p < 0.05) in comparison with group IV. NS means non-significant difference (p > 0.05) in comparison with group V.
Concerning IL-10 and IL-12, there was a significant increase in their expression in group V by 370.5 % and 354.5 %, respectively, relative to group II.
3.3.4.3 Gene expression of iNOS
The effect of EMEE on the relative gene expression of iNOS was elucidated and it is shown in Fig. 14. Group II revealed a significant increase in the gene expression of iNOS by 400 % in comparison with group I. Regarding group IV and V, they exhibited a significant downregulation of the gene expression of iNOS by 59.2 % and 73.4 %, respectively, in comparison with group II.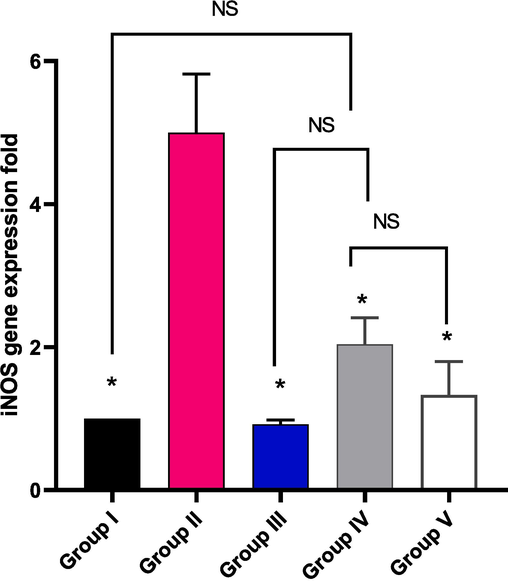
Impact of treatment with EMME on the gene expression of iNOS. The single asterisk represents a significant difference (p < 0.05) in comparison with group II. The abbreviation NS means non-significant difference (p > 0.05) in comparison with group IV.
4 Discussion
Inflammation usually leads to a progress of different pathophysiological proceedings that can provoke disease development (Chen et al., 2018, El-Banna et al., 2019). In the literature, there are many animal models that were used to elucidate the anti-inflammatory potential of various tested compounds. Carrageenan-induced paw edoema is regarded as one of the most used models to assess the anti-inflammatory activity of different natural and synthetic materials. Carrageenan is a chemical agent which is capable of stimulating the release of several inflammatory and pro-inflammatory mediators (Patil et al., 2019, Alherz et al., 2022).
Here, EMEE (200 mg/kg) treated group (group V) presented a significant decline (p < 0.05) in the paw edema weight in regard to groups II (positive control) and IV (treated with EMEE, 100 mg/kg). Interestingly, there wasn’t a significant difference (p > 0.05) between groups I (normal control), III (treated with celecoxib), and V. Edema is an important character for the local inflammation and it is formed owing to the fluid accumulation in the interstitial fluid of the tissues (Zamorano et al., 2021). Thus, it negatively affects tissue functions because the accumulated fluid hinders the diffusion of nutrients and oxygen to the cells of the affected tissue and compromises their metabolism (Wiig, 2011, Almukainzi et al., 2022).
During the inflammatory process, different cells like neutrophils and macrophages migrate to the site of the inflammation and they develop large amounts of inflammatory mediators such as TNF-α and interleukins. Such mediators induce a prolonged inflammatory reaction (Apostolova et al., 2020, Elekhnawy and Negm, 2022). Also, during inflammation, the stimulation of the COX-2 pathway is usually triggered and this leads to an increase in the release of prostaglandins like PGE2 which augment the inflammatory process (Guay et al., 2004, Attallah et al., 2022a). Herein, the carrageenan group (group II) revealed a strong positive COX-2 and TNF-α immuonostaining (score 3) and this is due to the inflammation which is triggered by carrageenan. Remarkably, this was mitigated by EMEE treatment as the low dose (100 mg/kg) resulted in a moderate positive COX-2 and TNF-α immunostaining (score 2) and the high dose (200 mg/kg) resulted in a mild positive COX-2 and TNF-α immunostaining (score 1). Decreasing the positive COX-2 and TNF-α immunostaining is an important indication as it means that both COX-2 and TNF-α are decreased which results in a decrease in the inflammatory reaction (Mercogliano et al., 2020, Alotaibi et al., 2021, Negm et al., 2021). Besides, the histological examination of the paw skin tissues of the different studied groups showed that treatment with EMEE (200 mg/kg) resulted in absence of inflammation and appearance of normal muscles are normal (black arrow).
EMEE (particularly at concentration of 200 mg/kg) significantly enhanced (p < 0.05) the levels of the anti-inflammatory cytokines (IL-12 and IL-10). It also significantly decreased (p < 0.05) the release of the pro-inflammatory mediators (IL-5, GM-CSF, iNOS, and MCP-1). These results imply that EMEE, at least in carrageenan-stimulated paw edema, acts to reduce inflammation by preventing the release of pro-inflammatory cytokines and mediators. Carrageenan solution into the paws produces an inflammatory reaction that is characterized by an infiltration of the inflammatory cells, an increase in exudate, and a marked production of pro-inflammatory mediators, such as prostaglandins, leukotrienes, and cytokines, as well as components of the oxidative stress response (Duarte et al., 2016, Attallah et al., 2022c).
It is commonly acknowledged that controlling inflammation involves a number of signalling channels and it is a difficult process. For instance, through controlling the expression of several pro-inflammatory mediators, the NF-kB signalling pathway contributes significantly to inflammation. The MAPK signalling route, in addition to the NF-kB signalling pathway, influences the release of several inflammatory mediators. There is a vast range of biological effects caused by the MAPK family, which comprises the enzymes ERK1/2, JNK, and p38 kinase. These enzymes are crucial in controlling the activity of several proteins, enzymes, and transcription factors (Thouverey and Caverzasio, 2015).
According to several research, inflammatory activation and oxidative stress-activated the MAPK signalling pathway, which led to the phosphorylation of ERK1/2, JNK, and p38 (Li et al., 2018). It is well known that the generation of the pro-inflammatory cytokines depends on the activation of MAPKs (Kang et al., 2016). In the current work, the phosphorylation of ERK1/2, JNK, and p38 by carrageenan-induced proteins promoted the release of TNF-α, which then induced the secretion of IL-6 and boosted the creation of COX-2. It is hypothesized that the MAPK signalling pathway contributes to the release of the inflammatory cytokines and the dysregulation of the production of COX-2 (Kang et al., 2016). The network pharmacology study determined the TNF gene to be the gene incorporated to most inflammatory disorders. The gene enrichment analysis of target genes identified by the compounds of EMEE determined the top signalling pathways; PI3K-Akt signalling pathway, MAPK signalling pathway, Ras signalling pathway, these results confirmed the hypothesis built in this study towards the signalling pathway contributed to the inflammation and proved that EMEE activity could be referred to a direct activation potential on MAPK signalling pathway and this anti-inflammatory potential attributed to TNF gene in addition to other identified 54 genes.
5 Conclusion
Our study suggested that EMEE had anti-inflammatory activity in a carrageenan-induced paw edoema model in rats. This was revealed via improving the histological findings as well as the mild positive TNF-α and COX-2 immunostaining of the EMEE-treated group (200 mg/kg). Our hypothesized mechanism is that EMEE inhibits the MAPK-mediated COX-2 signalling pathway, which dampens inflammation in carrageenan-induced paw edema. This was supported by the network pharmacology studies.
Funding
This research was funded by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2023R304), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
CRediT authorship contribution statement
Walaa A. Negm: Conceptualization, Methodology, Resources, Writing – original draft, Writing – review & editing. Engy Elekhnawy: Conceptualization, Investigation, Visualization, Methodology, Data curation, Writing – original draft. Fatma A. Mokhtar: Investigation, Methodology, Data curation, Writing – original draft. Reem Binsuwaidan: Methodology, Resources, Funding acquisition. Nashwah G.M. Attallah: Resources, Investigation, Data curation. Sally Abdallah Mostafa: Methodology, Resources, Writing – original draft. Ehssan Moglad: Methodology, Resources, Writing – review & editing. Sarah Ibrahim: Methodology, Investigation, Data curation. Omnia Momtaz Al-Fakhrany: Methodology, Investigation, Data curation, Writing – original draft. Duaa Eliwa: Methodology, Resources, Writing – original draft, Writing – review & editing.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- A Network Pharmacology Analysis of Cytotoxic Triterpenes Isolated from Euphorbia abyssinica Latex Supported by Drug-likeness and ADMET Studies. ACS Omega. 2022;7:17713-17722.
- [Google Scholar]
- Silver Nanoparticles Prepared Using Encephalartos laurentianus De Wild Leaf Extract Have Inhibitory Activity against Candida albicans Clinical Isolates. J. Fungi. 2022;8:1005.
- [Google Scholar]
- Co-delivery of gentiopicroside and thymoquinone using electrospun m-PEG/PVP nanofibers: In-vitro and In vivo studies for antibacterial wound dressing in diabetic rats. Int. J. Pharm.. 2022;625:122106
- [Google Scholar]
- Alnusaire, T.S., Sabouni, I.L., Khojah, H., Qasim, S., Al-Sanea, M.M., Siddique, S., Mokhtar, F.A., Ahmed, S.R., 2023. Integrating Chemical Profiling, In Vivo Study, and Network Pharmacology to Explore the Anti-inflammatory Effect of Pterocarpus dalbergioides Fruits and Its Correlation with the Major Phytoconstituents. ACS omega.
- Alotaibi, B.N.W., Elekhnawy, E., El-Masry, T.A., Elseady, W.S., Saleh, A., Alotaibi, K.N., El-Sherbeni, S.A., 2021. Antibacterial, Immunomodulatory, and Lung Protective Effects of Boswellia dalzielii Oleoresin Ethanol Extract in Pulmonary Diseases: In Vitro and In Vivo Studies. Antibiotics 10, 1444.
- Metabolic Profiling of Jasminum grandiflorum L. Flowers and Protective Role against Cisplatin-Induced Nephrotoxicity: Network Pharmacology and In Vivo Validation. Metabolites. 2022;12:792.
- [Google Scholar]
- Retama monosperma chemical profile, green synthesis of silver nanoparticles, and antimicrobial potential: a study supported by network pharmacology and molecular docking. RSC Adv.. 2023;13:26213-26228.
- [Google Scholar]
- Immunomodulatory and anti-inflammatory effects of fucoidan: A review. Polymers. 2020;12:2338.
- [Google Scholar]
- Anti-biofilm and antibacterial activities of Cycas media R. Br secondary metabolites: In silico, in vitro, and in vivo approaches. Antibiotics. 2022;11:993.
- [Google Scholar]
- Elucidation of the Metabolite Profile of Yucca gigantea and Assessment of its Cytotoxic, Antimicrobial, and Anti-Inflammatory Activities. Molecules. 2022;27:1329.
- [Google Scholar]
- In vivo and in vitro antimicrobial activity of biogenic silver nanoparticles against Staphylococcus aureus clinical isolates. Pharmaceuticals. 2022;15:194.
- [Google Scholar]
- Abelmoschus eculentus Seed Extract Exhibits In Vitro and In Vivo Anti-Alzheimer’s Potential Supported by Metabolomic and Computational Investigation. Plants. 2023;12:2382.
- [Google Scholar]
- Non-steroidal anti-inflammatory drugs (NSAIDs) and organ damage: A current perspective. Biochem. Pharmacol.. 2020;180:114147
- [Google Scholar]
- UniProtKB/Swiss-Prot, the manually annotated section of the UniProt KnowledgeBase: how to use the entry view. Plant Bioinform.: Methods Protocols 2016:23-54.
- [Google Scholar]
- Chen, L.D.H., Cui, H., Fang, J., Zuo, Z., Deng, J., Li, Y., Wang, X., Zhao, L., 2018. Inflammatory responses and inflammation-associated diseases in organs. Oncotarget 9, 7204.
- Duarte, D.B., Vasko, M.R., Fehrenbacher, J.C., 2016. Models of inflammation: carrageenan air pouch. Curr. Protocols Pharmacol. 72, 5.6.1-5.6. 9.
- Adaptation of Pseudomonas aeruginosa clinical isolates to benzalkonium chloride retards its growth and enhances biofilm production. Mol. Biol. Rep.. 2019;46:3437-3443.
- [Google Scholar]
- The potential application of probiotics for the prevention and treatment of COVID-19. Egypt. J. Med. Human Genet.. 2022;23:1-9.
- [Google Scholar]
- Evaluation of the impact of adaptation of Klebsiella pneumoniae clinical isolates to benzalkonium chloride on biofilm formation. Egypt. J. Med. Human Genet.. 2021;22:1-6.
- [Google Scholar]
- Network pharmacology and molecular docking study for biological pathway detection of cytotoxicity of the yellow jasmine flowers. BMC Complement. Med. Therapies. 2023;23:1-16.
- [Google Scholar]
- Vitis vinifera leaf extract liposomal Carbopol gel preparation's potential wound healing and antibacterial benefits: in vivo, phytochemical, and computational investigation. Food Funct.. 2023;14:7156-7175.
- [Google Scholar]
- Global medicinal uses of Euphorbia L. (Euphorbiaceae) J. Ethnopharmacol.. 2015;176:90-101.
- [Google Scholar]
- ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics. 2020;36:2628-2629.
- [Google Scholar]
- SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res.. 2014;42:W32-W38.
- [Google Scholar]
- Carrageenan-induced paw edema in rat elicits a predominant prostaglandin E2 (PGE2) response in the central nervous system associated with the induction of microsomal PGE2 synthase-1. J. Biol. Chem.. 2004;279:24866-24872.
- [Google Scholar]
- The role of interleukin-1 in general pathology. Inflammation Regeneration. 2019;39:1-16.
- [Google Scholar]
- Identification and mechanism of action of renoprotective constituents from peat moss Sphagnum palustre in cisplatin-induced nephrotoxicity. J. Funct. Foods. 2016;20:358-368.
- [Google Scholar]
- Cytoscape: software for visualization and analysis of biological networks. Data Min. Proteomics: Standards Appl. 2011:291-303.
- [Google Scholar]
- Immunosuppressive effect of geniposide on mitogen-activated protein kinase signalling pathway and their cross-talk in fibroblast-like synoviocytes of adjuvant arthritis rats. Molecules. 2018;23:91.
- [Google Scholar]
- Reactive oxygen species in autoimmune cells: function, differentiation, and metabolism. Front. Immunol.. 2021;12:635021
- [Google Scholar]
- Mercogliano, M.F.B.S., Elizalde, P.V., Schillaci, R., 2020. Tumor necrosis factor α blockade: an opportunity to tackle breast cancer. Front. Oncol. 10.
- Metabolic profiling and cytotoxic activities of ethanol extract of Dypsis leptocheilos aerial parts and its green synthesized silver nanoparticles supported by network pharmacology analysis. S. Afr. J. Bot.. 2023;161:648-665.
- [Google Scholar]
- Investigation of the Antibacterial Activity and Efflux Pump Inhibitory Effect of Cycas thouarsii R. Br. Extract against Klebsiella Pneumoniae Clinical Isolates. Pharmaceuticals. 2021;14:756.
- [Google Scholar]
- A review of the ethnomedical uses, phytochemistry and pharmacology of the Euphorbia genus. The Pharma Innov.. 2017;6:34.
- [Google Scholar]
- A novel community driven software for functional enrichment analysis of extracellular vesicles data. J. Extracellular Vesicles. 2017;6:1321455.
- [Google Scholar]
- Patil, K.R., M.U., Unger, B.S., Goyal, S.N., Belemkar, S., Surana, S.J., Ojha, S., Patil, C.R., 2019. Animal models of inflammation for screening of anti-inflammatory drugs: Implications for the discovery and development of phytopharmaceuticals. Int. J. Mol. Sci. 20, 4367.
- The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res.. 2020;48:D845-D855.
- [Google Scholar]
- Age and age-related diseases: role of inflammation triggers and cytokines. Front. Immunol.. 2018;586
- [Google Scholar]
- In vitro biological propensities and chemical profiling of Euphorbia milii Des Moul (Euphorbiaceae): A novel source for bioactive agents. Ind. Crop. Prod.. 2019;130:9-15.
- [Google Scholar]
- Antiviral and antibacterial potential of electrosprayed PVA/PLGA nanoparticles loaded with chlorogenic acid for the management of coronavirus and Pseudomonas aeruginosa lung infection. Artif. Cells Nanomed. Biotechnol.. 2023;51:255-267.
- [Google Scholar]
- Unravelling the antibacterial potential of biosynthesized selenium nanoparticles against Salmonella Typhimurium food pathogen: in vitro and in vivo investigation. Eur. Rev. Med. Pharmacol. Sci.. 2023;27:3631-3644.
- [Google Scholar]
- Euphorbia-derived natural products with potential for use in health maintenance. Biomolecules. 2019;9:337.
- [Google Scholar]
- Natural compounds with dual antimicrobial and anti-inflammatory effects. Phytochem. Rev.. 2020;19:1471-1502.
- [Google Scholar]
- A commensal symbiotic factor derived from Bacteroides fragilis promotes human CD39+ Foxp3+ T cells and Treg function. Gut Microbes. 2015;6:234-242.
- [Google Scholar]
- Focus on the p38 MAPK signaling pathway in bone development and maintenance. BoneKEy Reports. 2015;4
- [Google Scholar]
- Pathophysiology of tissue fluid accumulation in inflammation. J. Physiol.. 2011;589:2945-2953.
- [Google Scholar]
- Diterpenoids from the genus Euphorbia: Structure and biological activity (2013–2019) Phytochemistry. 2021;190:112846
- [Google Scholar]
- Diverse diterpenoids with α-glucosidase and β-glucuronidase inhibitory activities from Euphorbia milii. Phytochemistry. 2022;196:113106
- [Google Scholar]
- Tackling ischemic reperfusion injury with the aid of stem cells and tissue engineering. Front. Physiol.. 2021;12
- [Google Scholar]
- Phytochemical and pharmacological review of diterpenoids from the genus Euphorbia Linn (2012–2021) J. Ethnopharmacol. 2022115574
- [Google Scholar]
Appendix A
Supplementary material
Table S1. Sequences of the primers, Table S2. Top 20 biological KEGG pathways of genes related to inflammations among the studied data set, Figure S1. The total ion chromatogram of detected metabolites of E. milii ethanol extract in negative ion mode.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105568.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







