Translate this page into:
Quantitative profiling and mechanisms exploration of Epimedium total flavonoid capsules in neuroinflammation: An integrated study of pharmacokinetics, network pharmacology, and molecular pathways
⁎Corresponding authors at: State Key Laboratory on Technologies for Chinese Medicine Pharmaceutical Process Control and Intelligent Manufacture, Jiangsu Kanion Pharmaceutical Co. Ltd., Jiangsu, Lianyungang, 222001, China. lihaibo1985124@sina.com (Haibo Li), xwv8521@kanion.com (Wei Xiao),
⁎⁎Corresponding author. 1018yuyang@jnu.edu.cn (Yang Yu)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Abstract
Epimedium total flavonoid capsules (ETFCs), a traditional Chinese patent medicine derived from Epimedium brevicornu Maxim., have been used for centuries to treat primary osteoporosis and are associated with kidney yang deficiency symptoms. Based on the “kidney-brain axis” hypothesis, previous study demonstrated that ETFCs can improve the cognitive function of MCAO rats, and the mechanism may involve the inflammatory response. However, little is known about their therapeutic components and scientific connotations. This study systematically investigates the anti-neuroinflammatory material basis and the mechanisms of ETFCs, integrating UPLC-Q/TOF-MS and UPLC-TQ-MS for chemical profiling, quantitative analysis of key components in rats′ plasma and brain, along with network pharmacology and activity verification. As a result, 65 components were identified. Metabolite profiling showed a total of 130 xenobiotics, highlighting the extensive metabolic transformations. Pharmacokinetic studies showed that primary flavonoid glycosides were quickly absorbed, while secondary glycosides exhibited slower absorption and elimination. In brain tissue, both prototype glycosides and secondary glycosides reached their peak concentrations rapidly, with Tmax between 0.25–1 h, while the prototype glycosides were eliminated after 8 h and secondary glycosides exhibited a secondary peak at 6–8 h. A network pharmacology study revealed that prototypes in blood may play a therapeutic role through targets including AKR1B1, PDE5A, and PTGS2 (COX-2). The LPS-induced BV-2 cell model confirmed ETFCs′ constituents showed inhibitory activities on pro-inflammatory cytokine secretion, mRNA expressions of IL-6, TNF-α, COX-2, iNOS and could down-regulate the protein expressions of COX-2 and iNOS. This research lays the foundation for the further development and utilization of ETFCs.
Keywords
Epimedium total flavonoid capsules
Anti-neuroinflammation
Pharmacokinetics characteristics
Network pharmacology
Molecular mechanism
1 Introduction
Epimedium brevicornu Maxim., also known as Yin Yang Huo or Horny Goat Weed, is a Yang-enhancing and kidney-nourishing herb with a long history in traditional Chinese medicine (TCM) (Chen et al., 2015; Jiang et al., 2015; Ma et al., 2011). According to TCM theory, “the kidneys produce marrow, and the marrow nourishes the brain”, emphasizing the close relationship between kidneys and the brain (Feng et al., 2021). Therefore, the guiding principle of TCM in treating cognitive decline or neurological disorders is to tonify the kidneys, thus nourishing the marrow to enhance brain function.
Studies have shown that E. brevicornu is rich in isopentenyl flavanones, which exert a tonifying effect on the kidneys (Chen et al., 2015; Qian et al., 2024; Zhang et al., 2016). Numerous literature reports highlight the use of E. brevicornu in the treatment of neurological disorders, ischemic stroke, and other conditions, especially regarding its potential to enhance cognitive abilities by tonifying the kidneys (Lin et al., 2013; Wu et al., 2023; Xie et al., 2022; Xu et al., 2022). Epimedium total flavonoid capsule (ETFC), derived from the extraction of E. brevicornu, is a categorized Chinese patent medicine developed by Jiangsu Kangyuan Pharmaceutical Co., Ltd. and approved by the National Medical Products Administration. It is clinically used for primary osteoporosis associated with kidney yang deficiency syndrome, characterized by symptoms such as lower back pain, cold feet, frequent nocturia, and a pale tongue with a thin white coating. Extensive clinical applications have confirmed the safety and efficacy of ETFCs (Lu et al., 2013). In addition, a variety of bioactive compounds in Epimedium, especially flavonoids, have been the focus of research on chronic diseases such as tonifying the kidneys, strengthening bones, and improving cognitive ability (Jin et al., 2019; Li et al., 2015; Zhang et al., 2023).
It is well known that the therapeutic efficacy of orally administered TCM usually depends on the interaction between its active constituents and specific biological targets within the body. A comprehensive understanding of the serum pharmacokinetics of orally administered TCM is crucial for elucidating the absorption, distribution, metabolism, and excretion (ADME) of these active components (Li et al., 2022). In particular, considering the impact of the blood–brain barrier in neurological diseases, accounting for the temporal and spatial dynamics of components within target tissues will contribute to our understanding of the biological mechanisms underlying their efficacy. It is noteworthy that the characteristics of the mechanisms of action of TCM formulas are highly analogous to the holistic, systematic, and comprehensive nature of network pharmacology. Utilizing network pharmacology to explore the potential targets of TCM formulas, which are characterized by “multiple components,” is currently a mainstream research approach (Qin et al., 2024; Xiang et al., 2024; Zhao et al., 2023).
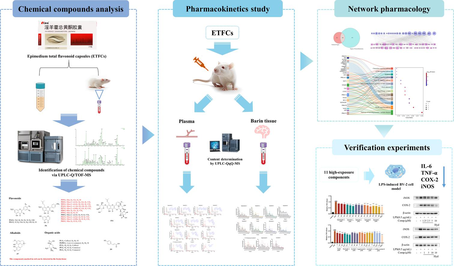
Based on the speculation that Epimedium tonifies the kidneys and may benefit the brain, we conducted a preliminary study on the improvement of post-stroke cognitive impairment (PSCI) by ETFCs (Yang et al., 2024). The results showed that ETFCs could improve cognitive function in middle cerebral artery occlusion (MCAO) stroke model rats, potentially through regulating the expression of p-IκBα and p-NF-κB proteins, thereby inhibiting neuroinflammation. However, the in vivo therapeutic components and scientific connotation remain unclear. To address these, our ongoing study aims to investigate the anti-neuroinflammatory material basis as well as the underlying mechanisms of ETFCs at both in vitro and in vivo levels. Hence, a five-step method was employed, involving UPLC qualitative and quantitative analysis techniques coupled with the network pharmacological study and in vitro experimental validation (Fig. 1.). Firstly, the composition analysis of ETFCs was elucidated using UPLC-Q-TOF-MS technology; Secondly, we described the complex multi-component metabolic characteristics of orally administered ETFCs in rats, while simultaneously characterizing the in vivo pharmacokinetic properties of its main constituents in both blood and brain tissues. This step has revealed highly exposed components within target tissues that may be associated with pharmacological effects, providing a more comprehensive interpretation of the chemical basis underlying the ETFCs' efficacy. Furthermore, we used network pharmacology combined with experimental validation to reveal the effects and mechanisms of ETFCs in improving neuroinflammation. By elucidating its complex composition, pharmacokinetics, and molecular mechanisms of action, our findings pave the way for the development of ETFCs-based therapeutics for the management of PSCI and related neuroinflammatory disorders.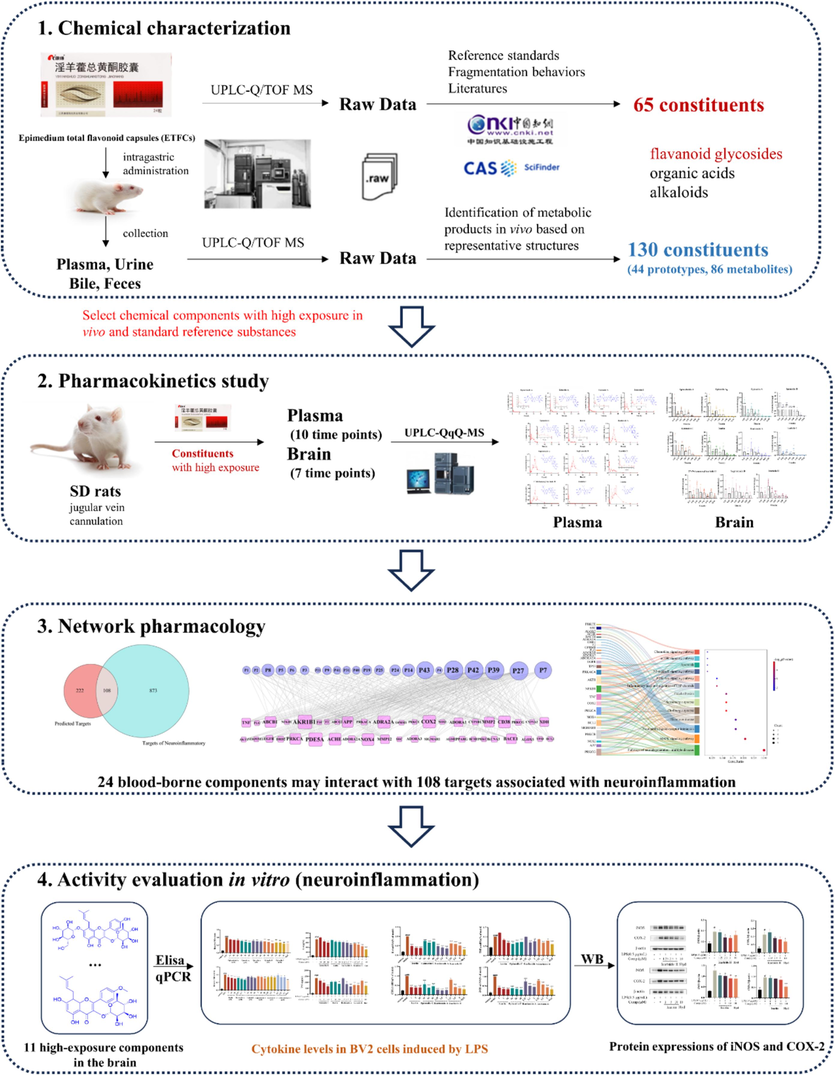
Research strategy.
2 Materials and methods
2.1 Chemicals and reagents
ETFCs were obtained from Kanion Pharmaceutical Co., Ltd ((Lot No. 210603, Jiangsu, China). Sagittatoside A, sagittatoside B, 2″-O-rhamnosyl icariside, ikarisoside A, epimedoside A, chlorogenic acid and hyperoside were purchased from Chengdu Purifa Technology Co., Ltd. (Chengdu, China). Icariin, epimedin A1, epimedin A, epimedin C, icariside I, icariside II and icaritin were purchased from Chengdu Herbpurify Co., Ltd. (Chengdu, China). Neochlorogenic acid, magnoflorine, and epimedin B were purchased from Shanghai Winherb Medical Technology Co., Ltd. (Shanghai, China). Cryptochlorogenic acid was purchased from Chengdu Push Bio-technology Co., Ltd (Chengdu, China). The purity of each compound was more than 98 % determined by HPLC analysis. Detailed information on the standards used in this study is shown in Table S1.
LC-MS-grade acetonitrile and methanol were purchased from Fisher Scientific (Fairlawn, NJ, USA). LC-MS-grade formic acid was obtained from Sigma-Aldrich (St. Louis, MO, USA). Information on other materials can be found in corresponding methods.
2.2 Animal and administration of ETFCs
Male Sprague Dawley (SD) rats, each weighing 250 ± 10 g, were obtained from the Medical Experimental Animal Center of Guangdong Province. They were kept under controlled conditions: a constant temperature of 23 °C, humidity of 55 %, a 12-hour light–dark cycle, and free access to food and water. Prior to the experiment, all rats underwent a one-week acclimation period. All procedures and management of all experimental animals strictly followed the “The Provision and General Recommendation for the Chinese Experimental Animals Administration Legislation guidance” approved by the Science and Technology Department of Jiangsu Province (Ethical Review No. ky2023030311 and ky2023010512.).
In the metabolic study, 10 male SD rats were randomly assigned to either the ETFCs group (n = 6) or the blank group (n = 4). The ETFCs group received an intragastric dose of ETFCs at 667 mg/kg/day for three days, while the blank group was given normal saline following the same schedule.
In the pharmacokinetic study, 7 male SD rats underwent jugular vein cannulation and recovered 12 h before administration. Then the rats received an oral dose of ETFCs at 1.33 g/kg/day.
In the brain tissue distribution study, after one week of acclimatization feeding, 48 male SD rats (250 ± 20 g) were also given ETFCs orally at the same dosage of 1.33 g/kg/day.
2.3 Biological Samples′ collection and pretreatment
2.3.1 For metabolic study
Plasma samples: Four rats were anesthetized with an intraperitoneal injection of 10 % chloral hydrate after the final dose. Blood from the hepatic portal vein was collected at 0.5, 1, 2, and 4 h. These plasma samples (500 µL each) were combined to create pooled plasma. Protein precipitation was performed by adding acetonitrile (three volumes) containing 0.1 % formic acid, followed by vortexing and centrifugation at 14,000 rpm for 10 min. The supernatant was then dried under nitrogen gas, and the residue was reconstituted in 200 μL of methanol.
Urine and feces samples: After ETFCs administration, urine and feces were collected over three days. Daily urine samples were pooled and centrifuged at 14,000 rpm for 10 min. The supernatant (10 mL) was applied to pre-activated SPE columns, washed with water, and eluted with methanol. The methanol eluent was dried under nitrogen at room temperature, and the residue was reconstituted in 200 µL methanol. Dried feces samples (1.0 g) were ground, soaked in 10 mL methanol for 24 h, and ultrasonically extracted for 60 min. The fecal extract was centrifuged at 14,000 rpm for 10 min, then the supernatant was evaporated to dryness and the residue re-dissolved in 5 mL water. This solution was processed on SPE columns similarly to the urine samples, with the final residue reconstituted in 500 µL methanol.
Bile samples: On the third-day post-administration, rats were anesthetized, and their bile ducts were cannulated using polyethylene tubing. A heating lamp was used to maintain body temperature. Bile (5 mL) was collected over 8 h and centrifuged at 14,000 rpm for 10 min. The supernatant was then processed using SPE columns following the same procedure as the urine samples. The final residue was reconstituted in 500 µL methanol.
Blank samples were processed identically. For analysis, 2 μL of plasma and urine samples and 1 μL of feces and bile samples were injected.
2.3.2 For quantitative study
Plasma: Blood samples were taken from the jugular vein at various time points (0.25, 0.5, 2, 4, 6, 8, 12, 24, 36, and 48 h) after ETFCs administration and placed in heparinized Eppendorf tubes. These were centrifuged at 14,000 rpm and 4 °C for 10 min to obtain plasma. Each 100 μL plasma sample was mixed with 50 μL methanol and 50 μL internal standard (isobavachin 10 ng/mL), followed by 300 μL acetonitrile (0.1 % formic acid) for protein precipitation. The mixture was vortexed for 2 min, centrifuged at 14,000 rpm for 10 min, then dried under nitrogen gas. The residue was reconstituted with 100 μL methanol, vortexed for 2 min, and centrifuged again at 14,000 rpm for 20 min. Finally, 2 μL of the supernatant was injected into UPLC-TQ/MS for analysis.
Brain tissue: After administration, the rats were anesthetized quickly and brain tissues were collected at 0.25, 0.5, 2, 4, 6, 8, and 12 h (n = 6 at each time point). The surface of the brain tissues was quickly washed with ice saline, dried with filter paper, and stored at −80 °C until analysis. The brain tissue was accurately weighed and homogenized with the four-fold volume of physiological saline to obtain a homogenate. 50 μL of methanol and 50 μL of IS were added to 100 μL of the homogenate. The subsequent steps were consistent with the procedures described for the treatment of plasma samples above.
2.4 UPLC-Q/TOF-MS
Chromatography analysis was performed using an Acquity UPLC I-Class system with a binary solvent system and an automatic sample manager. Separation occurred on a Waters BEH C18 column (2.1 mm × 100 mm, 1.7 μm) at 35 °C. The mobile phases were water (A) and acetonitrile (B), both with 0.1 % formic acid, delivered at 0.4 mL/min. The gradient program was: 0–2.5 min, 2–18 % B; 2.5–8.5 min, 18–31 % B; 8.5–11.5 min, 31–33 % B; 11.5–12.0 min, 33–50 % B; 12.0–13.5 min, 50–60 % B; 13.5–14.5 min, 60–100 % B; 14.5–15.0 min, 100 % B; 15–16 min, 100–2 % B; 16–17 min, 2 % B. The injection volume was 2 μL.
The UPLC system was coupled with a SYNAPTTM G2 HDMS quadrupole time-of-flight mass spectrometer (Waters, Manchester, U.K.), featuring an electrospray ionization (ESI) source. The settings were: capillary voltage of 3.0 kV (ESI+) or −2.0 kV (ESI−); sample cone voltage of 40 V (ESI+) or 30 V (ESI−); extraction cone voltage of 4 V; source temperature at 100 °C; desolvation temperature at 300 °C; cone gas flow at 50 L/h; and desolvation gas flow at 800 L/h. For mass accuracy in MSE mode, Sodium formate solution was used for calibration across 50–1500 Da, and Leucine-enkephalin (m/z 556.2771 in ESI+ mode; m/z 554.2615 in ESI− mode) served as an external reference at a constant flow rate of 5 μL/min.
2.5 UPLC-QqQ-MS
Components separation was achieved using an AcquityTM UPLC I-Class system (Waters Corp., Milford, MA, USA). Detailed information on chromatography analysis was given in Supplementary Material S1.2.
The UPLC system was paired with a Xevo TQ-XS mass spectrometer (Waters Corp., Milford, MA, USA) featuring an electrospray ionization (ESI) source. Multiple Reaction Monitoring (MRM) parameters for quantitative and semi-quantitative analytes in the pharmacokinetics and brain tissue study are optimized and shown in Table S2 and S3.
2.6 Method validation of pharmacokinetic research
In adherence to the 2018 FDA guidelines for bioanalytical method validation, a comprehensive assessment was conducted, encompassing specificity, linear response, sensitivity thresholds, as well as metrics for precision and accuracy. The extraction efficiency, matrix interference, and analyte stability were also rigorously evaluated. A detailed exposition of the validation procedures is delineated in Supplementary Material S1.3.
2.7 Network pharmacology
2.7.1 Target collection
The molecular structures of 23 bioavailable compounds in plasma, formatted in SMILES, were submitted to the SwissTargetPrediction platform (https://www.swisstargetprediction.ch/) for target prediction specific to 'Homo sapiens' (Gfeller et al., 2014). This initiative aimed to identify potential interactions between these compounds and pertinent protein targets. Concurrently, a targeted search within the CTD database (https://ctdbase.org/) was conducted, leveraging the keyword 'neuroinflammation' to extract genes associated with neuroinflammatory conditions. By mapping these gene names to their corresponding UniProt IDs, a comprehensive, non-redundant set of 330 neuroinflammatory-related target genes was compiled. Subsequently, the targets with a “Relevance score” greater than the mediator value were identified as neuroinflammation targets. The targets of 23 compounds for the treatment of neuroinflammation were obtained by overlaying the neuroinflammation targets with the predicted targets of these compounds. Ultimately, the compound-target network was built and the key target was extracted based on the mediator values of degree.
2.7.2 Enrichment analysis
All key targets were obtained from 2.7.1 and subsequently used for the enrichment analysis of Gene Ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. GO annotation and KEGG pathways analysis were conducted using the DAVID Bioinformatics Resources (https://david.ncifcrf.gov/), focusing on genes significantly associated with the study. The terms of PB, CC, MF and KEGG were obtained through a conditional screen that simultaneously satisfied 5 genes and a confidence level of 0.05. This analysis aimed to dissect the KEGG pathways linked to neuroinflammation, providing a visual representation of their interconnectedness and relevance.
2.7.3 Molecular docking
In order to verify the binding affinity between the active compound and targets, molecular docking was employed with the 5 highest-degree targets in the compound-target network to 23 bioavailable compounds. All molecular docking was performed using Autodock software. The target structures were extracted from the PDB database, which containing small molecule-target complexes, and their grid centers were determined based on the positions of the small molecules in the complexes. The grid size was 20 × 20 × 20. The 23 bioavailable compounds were prepared including hydrogenation and energy minimization.
2.8 In vitro anti-neuroinflammatory activity evaluation of potential bioactive components
2.8.1 Cell culture and treatment
The BV-2 murine microglial cell line was cultivated in DMEM (Gibco, USA) with 10 % fetal bovine serum (ExCell Bio, China) 100 IU/mL penicillin, and 100 μg/mL streptomycin. The cells were maintained in a controlled atmosphere of 37 °C and 5 % CO2, ensuring a humidified environment for optimal growth.
2.8.2 CCK8 assay
BV-2 cells were seeded into 96-well plates at 5 × 104 cells per mL (100 µL per well) overnight, and then the medium was discarded and 100 µL medium containing components were added to each well. Following a 24-hour exposure, 10 μL of the CCK8 reagent (GLPBIO, China) was introduced into each well for a subsequent 2-hour incubation period. Subsequently, the optical density was determined at a wavelength of 450 nm. Each trial was conducted three times to ensure reproducibility.
2.8.3 NO assay
Nitric oxide (NO) production, reflected by nitrite levels in the culture supernatant, was assessed using the Griess reaction. BV-2 cells were plated in a 96-well format at a concentration of 1.3 × 104 cells/mL for an initial 24-hour period, with 100 µL of medium per well. Subsequently, the growth medium was aspirated and replaced with 100 µL of fresh medium containing 0.5 μg/mL LPS (Sigma-Aldrich, USA) along with varying doses of the test compounds, followed by a 24-hour incubation. The assay was concluded by combining 50 µL of the supernatant with an equal volume of Griess reagent (Beyotime Biotechnology Co., Ltd, China), after which the optical density was recorded at 540 nm. The nitrite levels were then extrapolated from a sodium nitrite standard curve.
2.8.4 Elisa and quantitative PCR
Utilizing a protocol congruent with the procedures outlined for the “NO assay”, cells were subjected to treatment. Post-treatment, supernatants were harvested for the quantification of interleukin-6 (IL-6) and tumor necrosis factor-alpha (TNF-α) using enzyme-linked immunosorbent assay (ELISA) kits procured from 4A Biotech Co., Ltd, China. The assays were executed in strict accordance with the guidelines provided by the manufacturer.
BV-2 cells' total RNA was harvested utilizing Servicebio's extraction kit, following the manufacturer's recommended protocol. Subsequently, 1 μg RNA was converted to cDNA by using SweScript All-in-One SuperMix and gDNA Remover according to the reaction system in the instructions (Servicebio, China). Finally, the intended DNA bands were amplified with 10 μL SYBR Green qPCR Master Mix (Servicebio, China), 0.4 μL primers, and 2 μL cDNA. The sequences of PCR primers used are listed in Table S15.
2.8.5 Western blot
After LPS induction and drug treatment for 24 h, BV-2 cells were collected and lysed with RIPA buffer (Thermo Fisher, USA). The supernatant was obtained by high-speed centrifugation, followed by total protein quantification analysis using the BCA protein assay kit (Beyotime Biotechnology Co., Ltd, China). Protein samples, each weighing 40 μg, were resolved on a 10 % SDS-PAGE gel and subsequently transferred to a PVDF membrane. Post-transfer, the membranes were saturated with a 5 % non-fat milk solution for 2 h at ambient temperature prior to an overnight incubation with primary antibodies at 4 °C. Following three rounds of washing with TBST, the membranes were exposed to secondary antibodies for 2 h at room temperature. The immunoreactive bands were detected using an ECL detection kit from Absin and their intensities were measured employing the Image J software.
3 Results
3.1 Identification of EFTC-related prototypes in rats′ bio-samples
The base peak intensity (BPI) chromatograms of ETFCs in positive and negative ion modes are shown in Fig. 2. The results showed that 65 chemical components were detected in ETFCs (Fig. S1, Table S4), including 54 flavonoids (41 isopentenyl flavanones), 8 organic acids, 1 alkaloid, and 2 other types. Among them, 16 chromatographic peaks were accurately identified by comparison with the reference standards. Based on the analysis of chemical composition, it can be observed that isopentenyl flavanones are the main ingredients of ETFCs. The detailed analysis of the compounds can be found in the Supplementary Material S2.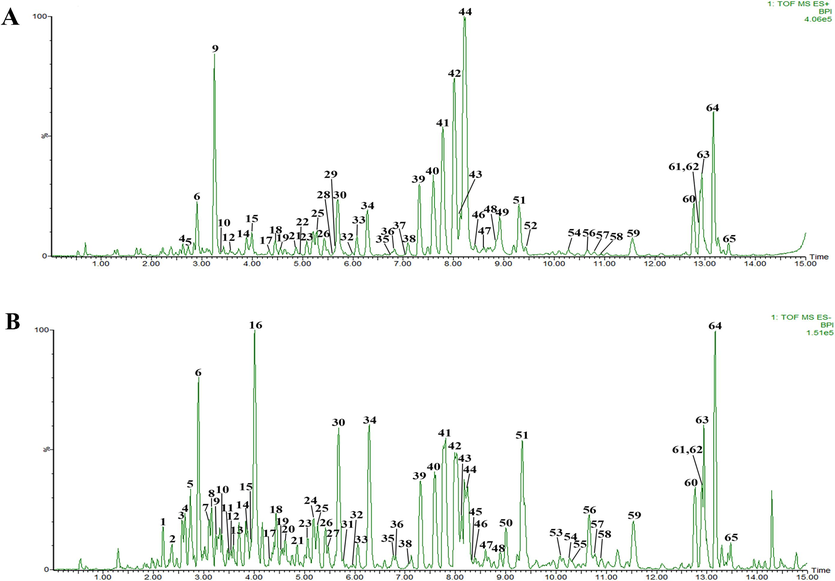
The base peak intensity (BPI) chromatograms of ETFCs. (A) (+) ESI-MS chromatogram of ETFCs. (B) (−) ESI-MS chromatogram of ETFCs.
Based on the chemical profile of ETFCs, characteristic mass patterns of ingredients were used for the identification of prototypes. Utilizing retention times and mass spectrometry data, 44 distinct prototypes were identified following oral ingestion of ETFCs. Of these, Detection across biological matrices revealed the presence of 24 prototypes in plasma, 25 in urine, 30 in bile, and 33 in fecal matter, as detailed in Fig. 3A and Table 1. Notably, the plasma-exposed prototypes were predominantly isopentenyl flavanone glycosides, suggesting their candidacy as bioactive constituents, as depicted in Fig. 4. Note: P, U, B and F represents plasma, urine, bile, and feces; N/A: not detected.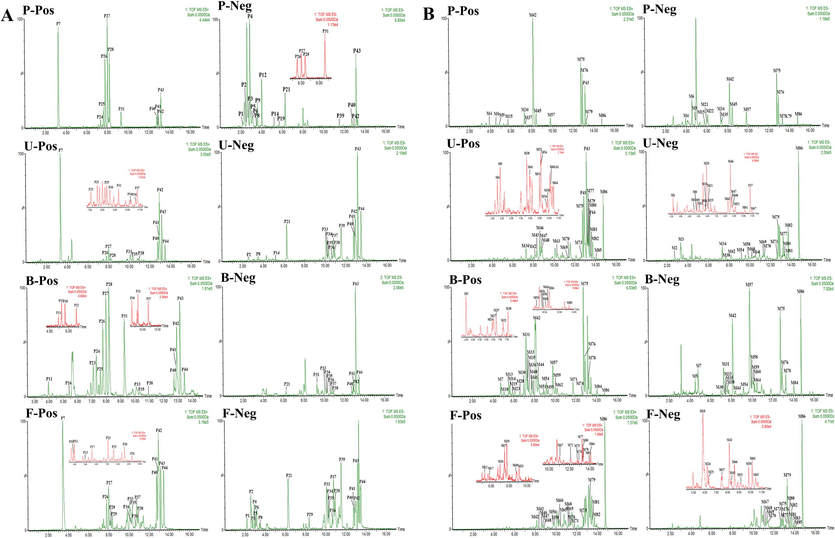
Extracted ion chromatograms (EICs) of xenobiotics after oral administration of ETFCs. (A) EICs of ETFCs related prototypes and (B) EICs of ETFCs related metabolites in rats. (P, U, B, and F represented rat plasma, urine, bile and fecal samples; Pos and Neg meant positive and negative ion modes).
No
tR
Elemental composition
Selected ion
Measured mass
Calculated mass
Mass error
MS/MS or MSE fragmentation
Identification
Source
ESI+
ESI-
P1
2.18
C16H18O9
[M−H]-
353.0876
353.0873
0.8
N/A
191.0525, 179.0354, 135.0457
neochlorogenic acid
P F
P2
2.56
C16H18O8
[M−H]-
337.0923
337.0923
0.9
N/A
191.0556, 163.0404
trans-5-O-p-coumaroylquinic acid
P U F
P3
2.73
C16H18O9
[M−H]-
353.0870
353.0873
−0.8
N/A
191.0493, 179.0345
chlorogenic acid
P F
P4
2.89
C9H6O4
[M−H]-
177.0191
177.0188
0.3
N/A
133.0298
5,7-dihydroxychromone
P F
P5
3.09
C16H18O8
[M−H]-
337.0914
337.0923
−0.9
N/A
191.0562, 163.0434
4-O-p-coumaroylquinic acid
P F
P6
3.15
C16H18O8
[M−H]-
337.0922
337.0923
−0.3
N/A
191.0544, 163.0391
3-O-p-coumaroylquinic acid
P F
P7
3.26
C20H23NO4
[M + H]+
342.1708
342.1705
0.9
297.1135, 282.0866, 265.0872, 237.0906, 191.0857
N/A
magnoflorine
P U F
P8
3.52
C16H18O8
[M−H]-
337.0928
337.0923
1.5
N/A
191.0558, 163.0390
cis-5-O-p-coumaroylquinic acid
P U F
P9
3.59
C9H8O3
[M−H]-
163.0402
163.0395
4.3
N/A
119.0501
p-hydroxycinnamic acid
P
P10
3.90
C21H20O12
[M + H]+
465.1041
465.1033
1.7
303.0511, 287.0555, 177.0915
N/A
hyperoside
F
P11
3.97
C27H30O14
[M + H]+
579.1715
579.1714
0.2
N/A
431.0986, 285.0422
kaempferol-3,7-di-O-rhamnoside
F B
P12
4.03
C18H24O10
[M−H]-
399.1284
399.1291
−1.8
N/A
237.0762, 219.0658, 193.0860
unknow
P
P13
4.93
C33H42O17
[M + H]+
711.2486
711.2500
−2.1
565.1931, 403.1412, 385.1279
N/A
unknow
F B
P14
5.19
C20H18O10
[M−H]-
417.0820
417.0822
−0.5
N/A
285.0396
kaempferol-3-O-xylopyranoside
P U
P15
5.26
C38H48O20
[M + H]+
825.2800
825.2817
−2.1
663.2275, 517.1714, 355.2614
N/A
diphylloside A
B
P16
5.43
C37H46O19
[M + H]+
795.2681
795.2712
−3.9
663.2304, 517.1724, 355.1189
N/A
epimedoside E
B
P17
5.60
C39H52O20
[M + H]+
841.3121
841.3130
−1.1
695.2548, 549.1592, 387.1438, 369.1323
N/A
icariside D
F
P18
5.62
C33H42O16
[M + H]+
695.2550
695.2551
−0.1
549.1975, 387.1433, 369.1323, 313.0713
N/A
icariside B
B
P19
5.68
C32H38O15
[M−H]-
661.2132
661.2132
0.0
N/A
514.1442, 499.1745, 353.0998
epimedoside A
P
P20
5.76
C38H48O19
[M + H]+
809.2850
809.2868
−2.2
663.2325, 517.1689, 355.185
N/A
diphylloside B
B
P21
6.29
C24H34O9
[M−H]-
465.2129
465.2125
0.5
N/A
303.1598, 285.1509
taxifolin-7-O-glucoside
P U F B
P22
6.82
C39H48O20
[M + H]+
837.2816
837.2817
−0.1
705.2366, 367.1174, 355.1172
N/A
4′'-O-acetyl-3-O-xylopyranosyl epimedoside A
B
P23
7.09
C28H32O14
[M + H]+
593.1872
593.187
0.3
447.1274, 285.0766
N/A
acacetin 7-O-rutinoside
U F B
P24
7.31
C39H50O20
[M + H]+
839.2973
839.2974
−0.1
677.2449, 531.1893, 369.1347, 313.0722
N/A
epimedin A1
P B
P25
7.60
C39H50O20
[M + H]+
839.2953
839.2974
−2.5
677.2421, 531.1862, 369.1336, 313.0714
N/A
epimedin A
P U F B
P26
7.77
C38H48O19
[M + H]+
809.2847
809.2868
−2.6
677.2443, 531.1856, 369.1323, 313.0710
645.2172, 512.2684, 366.1107, 351.0872
epimedin B
P U F B
P27
8.01
C39H50O19
[M + H]+
823.3006
823.3025
−2.3
677.2428, 531.1854, 369.1337, 313.0706
659.2327, 366.1097, 351.0867
epimedin C
P U F B
P28
8.21
C33H40O15
[M + H]+
677.2437
677.2445
−1.2
531.1866, 369.1333, 313.0709
513.1795, 367.1162, 352.0911, 297.0391
icariin
P U F B
P29
8.35
C33H42O15
[M−H]-
677.2462
677.2445
2.5
N/A
384.1214, 367.1151, 341.1018
wanepimedoside A
U F
P30
8.60
C27H32O11
[M + H]+
533.2021
533.2023
−0.4
387.1446, 369.1335, 313.0715
N/A
4′-methoxynoricaritin-3-O-rhamnoside
U F B
P31
9.28
C39H48O19
[M + H]+
821.288
821.2868
1.5
677.2415, 531.1870, 369.1317, 313.0717
529.1692, 367.1181
icaritin-3-O-(2-hydroxy-2-carboxyl-5-methyltetrahydrofuran-1-yl) rhamnopyranoside-7-O-glucopyranoside
P U F B
P32
9.41
C40H50O20
[M + H]+
851.2985
851.2974
−1.9
719.2525, 531.1849, 369.1320
N/A
sempervirenoside B
B
P33
10.09
C32H38O15
[M−H]-
661.2135
661.2132
0.5
355.1183, 299.0562
353.1014, 297.0405
ikarisoside B or its isomer
U F B
P34
10.28
C27H30O11
[M−H]-
529.1704
529.1710
−1.5
385.1288, 369.2430
383.1128, 297.0397
caohuoside C or its isomer
U F B
P35
10.32
C32H38O15
[M−H]-
661.2136
661.2132
0.6
355.1178, 299.0551
353.1019
ikarisoside B or its isomer
U F B
P36
10.64
C27H30O11
[M−H]-
529.1697
529.1710
−2.5
N/A
383.1099, 297.0391
caohuoside C or its isomer
U F B
P37
10.78
C31H36O14
[M−H]-
631.2022
631.2027
−0.8
355.1185, 299.0555
353.1003
ikarisoside F
U F B
P38
10.92
C32H38O14
[M−H]-
645.2188
645.2183
0.8
501.1762, 355.1181, 299.0557
352.0953
2′'-O-rhamnosyl ikarisoside A
U F B
P39
11.54
C26H28O10
[M−H]-
499.1612
499.1604
1.6
N/A
353.1023, 309.0399, 281.0459
ikarisoside A
P U F
P40
12.76
C33H40O15
[M + H]+
677.2441
677.2445
−0.6
369.1337, 313.0712
531.2291, 367.1180, 352.0931, 298.0481
sagittatoside A
P U F B
P41
12.90
C32H38O14
[M + H]+
647.2332
647.2340
−1.2
369.1344, 313.0719
366.1101, 351.0886, 323.0907
sagittatoside B
P U F B
P42
12.93
C33H40O14
[M + H]+
661.2495
661.2496
−0.2
369.1340, 313.0716
366.1104, 351.0870, 323.0919
2″-O-rhamnosyl icariside Ⅱ
P U F B
P43
13.17
C27H30O10
[M + H]+
515.1910
515.1917
−1.4
369.1333, 313.0710
366.1107, 351.0865, 323.0920
icariside Ⅱ
P U F B
P44
13.46
C33H38O14
[M−H]-
657.2186
657.2183
0.5
515.1936, 369.1349, 313.0717
513.1763, 367.1185, 352.0951
3′''-carbonyl-2′'-β-L-quinovosyl icariside II
U F B
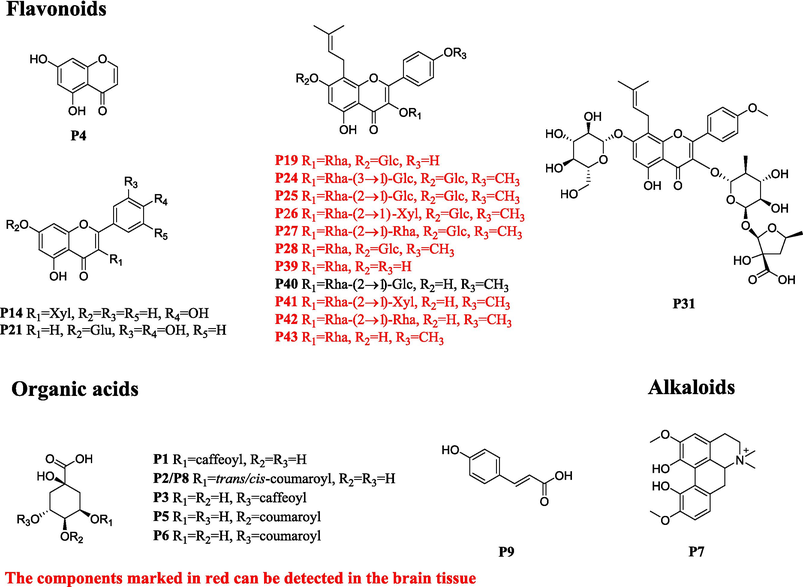
Chemical structures of the prototypes absorbed in plasma after oral administration of ETFCs.
3.2 Identification of EFTC-related metabolites in rats′ bio-samples
The analysis strategy of “homologous metabolites identification based on representative structures” was employed to quickly identify related metabolites in rats following oral administration of ETFCs (Geng et al., 2014). Three aglycones, kaempferol, quercetin, and icariin, were defined as representative structures and imported into Metabolynx XS software. Using MDF technology, potential metabolites in rats′ biological samples were screened. As a result, a total of 86 metabolites were characterized, including 16 metabolites in plasma, 41 in urine, 43 in bile, and 37 in feces (Fig. 3B, Table 2). There were few metabolites found in plasma, primarily glucuronidation products of secondary glycosides and aglycones, and their responses were higher than those of the prototypes of isopentenyl flavanones glycosides. The detailed analysis of the compounds can be found in the Supplementary Material S3. Note: P, U, B and F represents plasma, urine, bile, and feces; N/A: not detected.
No
tR
Elemental composition
Selected
ionMeasured
massCalculated
massMass
errorMS/MS or MSE fragmentation
Identification
Source
ESI+
ESI-
M1
2.16
C10H12O4
[M−H]-
195.0664
195.0657
3.6
N/A
150.0321
methyl-hydrocaffeic acid
U
M2
2.76
C17H20O9
[M−H]-
367.1033
367.1029
1.1
N/A
191.0347
(trans or cis) 4-O-methylcaffeoylquinic acid
U
M3
3.34
C17H20O9
[M−H]-
367.1024
367.1029
−1.4
N/A
191.0562, 173.0461
(trans or cis) 4-O-methylcaffeoylquinic acid
U
M4
3.92
C21H18O13
[M + H]+
479.0836
479.0826
2.1
303.0507
301.0339
quercetin + GluA
P
M5
4.48
C21H18O12
[M + H]+
463.0873
463.0877
−0.9
287.0551
285.0401
kaempferol + GluA
B
M6
4.55
C21H18O12
[M + H]+
463.0874
463.0877
−0.6
287.0557
285.0401
kaempferol + GluA
P U
M7
4.72
C32H34O18
[M + H]+
707.1824
707.1823
0.1
531.1509, 355.1189, 299.0563
529.1342, 353.1029
icaritin − CH2 + 2GluA
B
M8
4.83
C22H20O13
[M−H]-
491.0815
491.0826
−2.2
N/A
315.0500, 300.0281
quercetin + CH2 + GluA
U
M9
4.87
C21H18O12
[M + H]+
463.0886
463.0877
1.9
287.0561
285.0403
kaempferol + GluA
P U
M10
5.04
C33H38O17
[M + H]+
707.2167
707.2187
−6.9
561.1601, 543.1605, 385.1623, 367.1179
N/A
icariside II + GluA + O
B
M11
5.08
C22H20O13
[M−H]-
491.0825
491.0826
−0.2
N/A
315.0502
quercetin + CH2 + GluA
U
M12
5.27
C39H52O21
[M + H]+
857.3047
857.3079
−3.7
711.2467, 565.1907, 403.1392, 385.1279, 313.0721
N/A
epimedin C + O + H2O
F
M13
5.49
C32H36O16
[M + H]+
677.2085
677.2082
0.4
531.1508, 355.1188, 299.0565
N/A
icariside II − CH2 + GluA
B
M14
5.62
C38H48O19
[M + H]+
809.2866
809.2868
−0.2
663.2283, 517.1714, 355.1187
N/A
epimedin C − CH3
B
M15
5.74
C26H28O12
[M + H]+
533.1655
533.1659
−0.8
357.1341
355.1165, 327.1225,193.0857
icaritin − H2 − CH2 + GluA
P
M16
5.79
C22H20O13
[M−H]-
491.0824
491.0826
−0.4
N/A
315.0503, 300.0269
quercetin + CH2 + GluA
U
M17
5.79
C39H50O20
[M + H]+
839.2935
839.2974
−4.6
693.2343, 547.1782, 385.1283, 367.1186
N/A
epimedin C + O
F
M18
5.89
C15H10O9S
[M−H]-
364.9975
364.9967
2.2
N/A
285.0400
kaempferol + sul
U F
M19
5.89
C33H38O17
[M + H]+
707.2172
707.2187
−2.1
561.1619, 385.1288
N/A
icariside II + GluA + O
B
M20
5.92
C15H10O10S
[M−H]-
380.9918
380.9916
0.8
N/A
301.0352, 151.0038
quercetin + sul
U
M21
5.95
C15H10O9S
[M−H]-
364.9968
364.9967
0.3
N/A
285.0394
kaempferol + sul
P U
M22
6.06
C15H10O9S
[M−H]-
364.997
364.9967
0.8
N/A
285.0410
kaempferol + sul
P
M23
6.08
C32H34O18
[M + H]+
707.1814
707.1823
−1.3
531.1501, 355.1182, 299.0557
N/A
icaritin − CH2 + 2GluA
B
M24
6.2
C16H12O10S
[M−H]-
395.0087
395.0073
3.5
N/A
315.0508, 300.0273
quercetin + CH2 + sul
F
M25
6.34
C16H12O10S
[M−H]-
395.0072
395.0073
−0.3
N/A
315.0506, 300.0273
quercetin + CH2 + sul
U F
M26
6.43
C26H28O13
[M + H]+
549.1614
549.1608
1.1
373.1277, 355.1180
N/A
icaritin − CH2 + GluA + H2O
B
M27
6.50
C26H26O13
[M + H]+
547.145
547.1452
−0.4
371.1123, 353.2486
N/A
icaritin − CH2 + GluA + O
B
M28
6.59
C33H38O17
[M + H]+
707.217
707.2187
−2.4
561.1602, 385.1299, 367.1187
N/A
icariside II + GluA + O
B
M29
6.72
C26H26O13
[M + H]+
547.1451
547.1452
−0.2
371.1137, 353.2484
N/A
icaritin − CH2 + GluA + O
B
M30
6.98
C39H48O21
[M + H]+
853.2761
853.2766
−0.6
545.1674, 369.1338, 313.0722
675.2269, 513.2712, 367.1186
sagittatoside A + GluA
B F
M31
7.21
C33H36O18
[M + H]+
721.1982
721.198
0.3
545.1682, 369.1342, 313.0722
N/A
icaritin + 2GluA
B
M32
7.23
C39H46O22
[M + H]+
867.2551
867.2559
−0.9
545.1666, 369.1349, 313.0718
N/A
icariside II + 2GluA
B
M33
7.30
C39H48O21
[M + H]+
853.2759
853.2766
−0.8
545.1666, 369.1342, 313.0715
N/A
sagittatoside A + GluA
B
M34
7.32
C33H36O18
[M + H]+
721.1982
721.1980
0.3
545.1682, 369.1342, 313.0722
543.1509, 367.1184
icaritin + 2GluA
P U
M35
7.36
C33H38O17
[M + H]+
707.2181
707.2187
−0.8
531.1863, 369.1348, 313.0718
N/A
icaritin + glu + GluA
B P
M36
7.45
C33H38O17
[M + H]+
707.2173
707.2187
−2.0
545.1650, 369.1341, 313.0719
N/A
Icaritin + glu + GluA
B
M37
7.5
C32H40O16
[M + H]+
681.2375
681.2395
−2.9
549.1976, 403.1393, 385.1288, 369.1333
547.2902, 400.1168, 367.1184
sagittatoside B + O + H2O
F
M38
7.64
C27H30O13
[M + H]+
563.1761
563.1765
−0.7
387.1434, 369.1335, 313.0715
385.1271, 367.1181
icaritin + H2O + GluA
B U
M39
7.76
C33H42O16
[M + H]+
695.2537
695.2551
−2.0
549.1976, 531.1846, 387.1111, 369.1333
N/A
icariin + H2O
F
M40
7.72
C38H46O20
[M + H]+
823.2659
823.2661
−0.2
691.2255, 545.1655, 369.1334
N/A
sagittatoside B + GluA
B
M41
7.96
C39H48O20
[M + H]+
837.2803
837.2817
−1.7
691.2224, 545.1657, 369.1335, 313.0713
N/A
2″-O-rhamnosyl icariside ⅡI + GluA
U B
M42
8.16
C33H38O16
[M + H]+
691.2222
691.2238
−2.3
545.1655, 369.1341, 313.0776
513.1759, 366.1098
icariside II + GluA
P U F B
M43
8.29
C15H10O5
[M−H]-
269.0463
269.045
4.8
N/A
N/A
kaempferol − O
U F
M44
8.39
C26H26O12
[M + H]+
531.1497
531.1503
−1.1
355.1177, 299.0551
353.10332, 298.0484
icaritin − CH2 + GluA
B
M45
8.44
C26H26O12
[M + H]+
531.1503
531.1503
0.0
355.1175, 299.0559
353.10332, 298.0484
icaritin − CH2 + GluA
P
M46
8.59
C15H10O6
[M−H]-
285.0402
285.0399
1.1
N/A
N/A
qercetin − O
U F
M47
8.73
C16H12O6
[M−H]-
299.0562
299.0556
2
286.0473
285.1492, 284.0329
kaempferol + CH2
U F
M48
8.81
C16H12O6
[M−H]-
299.056
299.0556
1.3
286.0472
285.1499, 284.0334
kaempferol + CH2
U F
M49
8.81
C38H46O20
[M + H]+
823.2682
823.2661
2.6
677.2120, 531.1875, 355.2682
N/A
2″-O-rhamnosyl icariside Ⅱ − CH2 + GluA
F
M50
8.91
C27H30O11
[M + H]+
531.1863
531.1866
−0.6
385.1292, 367.1186
N/A
icariside II + O
B
M51
8.94
C27H28O13
[M + H]+
561.1616
561.1608
1.4
545.1667, 385.1297, 367.1195
N/A
icaritin + GluA + O
B U
M52
9.12
C16H12O7
[M−H]-
315.0515
315.0505
3.2
302.0436
300.0286
quercetin + CH2
U
M53
9.12
C32H38O16
[M + H]+
679.2238
679.2238
0.0
661.2437, 531.1865, 481.2814, 371.1136, 315.0499
N/A
icariin − CH3 + O
F
M54
9.15
C27H28O13
[M + H]+
561.1612
561.1608
0.7
385.1293, 367.1187, 313.0719
N/A
icaritin + O + GluA
B U
M55
9.32
C27H28O13
[M + H]+
561.1601
561.1608
−1.2
385.2181, 369.1327
N/A
icaritin + O + GluA
B
M56
9.74
C32H38O15
[M + H]+
663.2292
663.2298
0.5
385.1296, 367.1189, 313.0716
N/A
sagittatoside B + O
F B U
M57
9.77
C26H26O12
[M + H]+
531.1520
531.1503
3.2
355.1179, 299.0555
353.1013
icaritin − CH2 + GluA
P B
M58
9.85
C33H40O15
[M + H]+
677.2438
677.2445
−1.0
659.2379, 531.1879, 385.1279, 367.1191
N/A
2″-O-rhamnosyl icariside Ⅱ + O
B F U
M59
10.04
C27H28O13
[M + H]+
561.1605
561.1608
−0.5
385,1288, 367.1194, 355.2643, 313.0716
383.1132, 352.0946
icaritin + GluA + O
B
M60
10.16
C33H40O15
[M + H]+
677.2433
677.2445
−1.8
659.2323, 531.1873, 385.1290, 367.1190
N/A
2″-O-rhamnosyl icariside Ⅱ + O
B F U
M61
10.18
C17H12O8
[M−H]-
343.0461
343.0454
2
N/A
300.1655
quercetin + acetylation
U
M62
10.18
C33H36O18
[M + H]+
721.1965
721.198
−2.1
545.1653, 369.1333, 313.0713
N/A
icaritin + 2GluA
B
M63
10.26
C33H36O18
[M + H]+
721.1968
721.198
−1.7
545.1675, 369.1335
N/A
icaritin + 2GluA
U
M64
10.43
C27H30O13
[M + H]+
563.1761
563.1765
−0.7
545.1624, 387.1442, 369.1337, 313.0715
385.1283, 331.1911, 311.0556
icaritin + GluA + H2O
B U
M65
10.45
C33H40O16
[M + H]+
693.2390
693.2395
−0.7
531.1875, 385.1286, 367.1190
N/A
icariin + O
F
M66
10.66
C33H36O16
[M + H]+
689.2075
689.2082
−1.0
543.1508, 513.1772, 367.1180
N/A
icarisdie II + GluA − H2
B
M67
11.00
C31H36O14
[M + H]+
633.2181
633.2183
−0.3
355.1181
353.1031
sagittatoside B − CH2
U F
M68
11.17
C32H38O15
[M + H]+
663.2282
663.2289
−1.1
531.1855, 385.1286
383.1115, 367.0804
sagittatoside B + O
F
M69
11.32
C32H38O14
[M + H]+
647.2318
647.234
−3.4
501.1751, 355.1180, 299.0552
352.0952
2″-O-rhamnosyl icariside Ⅱ − CH2
U F
M70
11.42
C33H40O15
[M + H]+
677.2435
677.2445
−1.5
531.1852, 385.1278,
N/A
2″-O-rhamnosyl icariside Ⅱ + O
U F
M71
11.73
C33H38O16
[M + H]+
691.2229
691.2238
−1.3
545.1658, 369.1340, 313.0717
N/A
icarisdie II + GluA
B F
M72
12.13
C27H30O11
[M + H]+
531.1868
531.1866
0.4
385.129
N/A
icariside II + O
F
M73
12.65
C33H38O15
[M + H]+
675.2285
675.2289
−0.6
543.1872, 355.1189, 299.0569
N/A
desmethylicaritin 3-O-rha(Ac)-xyl
U F
M74
12.73
C33H38O16
[M + H]+
691.2220
691.2238
−2.6
545.1658, 369.1341, 313.0714
N/A
icarisdie II + GluA
B
M75
12.77
C27H28O12
[M + H]+
545.1663
545.1659
0.7
369.1336, 313.0708
367.1179, 352.0948
icaritin + GluA
P U F B
M76
12.91
C27H28O12
[M + H]+
545.1666
545.1659
1.3
369.1344, 13.0715
367.1183, 352.0942
icaritin + GluA
P F B
M77
13.24
C21H22O7
[M + H]+
387.1443
387.1444
−0.3
369.1343, 313.0716
N/A
icaritin + H2O
F U
M78
13.26
C27H28O12
[M + H]+
545.1660
545.1659
0.2
369.1338, 313.0719
367.1190, 352.0947
icaritin + GluA
P F B
M79
13.30
C20H18O6
[M + H]+
355.1180
355.1182
−0.6
299.0555
283.2636
icaritin − CH2
P U F
M80
13.32
C34H40O15
[M + H]+
689.2429
689.2445
−2.3
369.1330, 299.0556, 313.0719
515.3051, 453.2318, 353.1035
anhydroicaritin 3-O-rha(Ac)-xyl
F U
M81
13.54
C21H20O7
[M + H]+
385.1280
385.1287
−1.8
367.1176, 313.0705
365.1118
icaritin + O
U F
M82
13.73
C21H20O7
[M + H]+
385.1287
385.1287
0.0
367.1188, 313.0721
365.1101, 337.2049
icaritin + O
U F
M83
13.93
C30H32O12
[M + H]+
585.1979
585.1972
1.2
369.1336, 313.0714
367.1177, 352.0936
anhydroicaritin-3-O-xyl(2Ac)
F
M84
14.05
C21H18O6
[M + H]+
367.1190
367.1182
2.2
313.1561
350.0764
icartin − H2
B
M85
14.06
C21H18O7
[M + H]+
383.1119
383.1131
−3.1
365.1014, 313.0708
N/A
icartin − H2 + O
U F
M86
14.77
C21H20O6
[M + H]+
369.1342
369.1338
1.1
313.0710
N/A
icaritin
P U F B
3.3 Method validation of pharmacokinetic research
Based on the metabolic study of ETFCs in rats, 11 isopentenyl flavanone glycosides, including epimedoside A, epimedin A1, epimedin A, epimedin B, epimedin C, icariin, ikarisoside A, sagittatoside A, sagittatoside B, 2″-O-rhamnosyl icariside and icariside, which were detected in rat plasma, showing high exposure levels with available reference substances. Furthermore, compounds such as icariside I and icaritin, identifiable in urine samples, are documented for their bioactivity (Bi et al., 2022; Zhang et al., 2023). Therefore, the above 13 compounds were selected as the quantitative components of plasma pharmacokinetics, with 11 being absorbable into the brain tissue. The method validation was carried out according to FDA bioanalytical method validation guidelines with the following results:
Plasma-specificity testing confirmed the absence of interference from endogenous substances at the analytes' and internal standard's (IS) retention times, as illustrated in Fig. S3. The plasma's quantitative constituents demonstrated a robust linear correlation (R2 > 0.99) across the specified linear range detailed in Table S5. The LLOQs for plasma analytes spanned a range from 0.03 to 0.25 ng/mL, ensuring precision and accuracy aligned with the stipulated analytical criteria, as noted in Table S6. As displayed in Table S7, the intra-day and inter-day precision (RSD%) of three concentrations in plasma ranged between 1.71 % and 9.93 %. Concurrently, the RE% for accuracy was confined within a range from −4.25 % to 5.02 %. The matrix effects of the 13 quantitative components and IS in plasma at three concentrations were between 85.49 % and 106.94 % with RSDs within 8.79 %. Consistently, the recovery rates for the aforementioned analytes ranged between 80.66 % and 100.44 %, with icariside I showing a slightly lower range of 62.79 % to 69.80 %. The RSDs for these recoveries were contained within an 11.74 % margin, as summarized in Table S8. The stability test data of the 13 analytes under four different conditions are shown in Table S9. Their RSDs ranged from −4.01 % to 14.08 %, and accuracy was less than 14.60 %. Moreover, this method ensured no residual interference from the 13 compounds in plasma (Fig. S4).
Above all, the validated method can be applied to the pharmacokinetic analysis of the 13 components in rat plasma. Similarly, full method validation in rat brain tissue demonstrated good specificity, linearity, accuracy, precision, matrix effect, extraction recovery, and stability for 11 components. The results are summarized in Fig. S5-S6 and Table S10–S14.
3.4 Pharmacokinetic studies
3.4.1 Plasma
The optimized approach was successfully applied to simultaneously measure 13 key components in rat plasma after oral administration of ETFCs. The mean plasma concentration–time profile and corresponding pharmacokinetic parameters are displayed in Fig. 5 and Table 3, respectively. According to the metabolic profile, the plasma metabolites of isopentenyl flavone glucosides primarily existed in the form of glucuronidation. The dynamic changes in the metabolites together with the prototypes contributed to describing the overall dynamic characteristics of ETFCs in vivo. Therefore, the pharmacokinetic characteristics of 8 semi-quantitative components were also monitored by using peak area-time change curves (Fig. S7).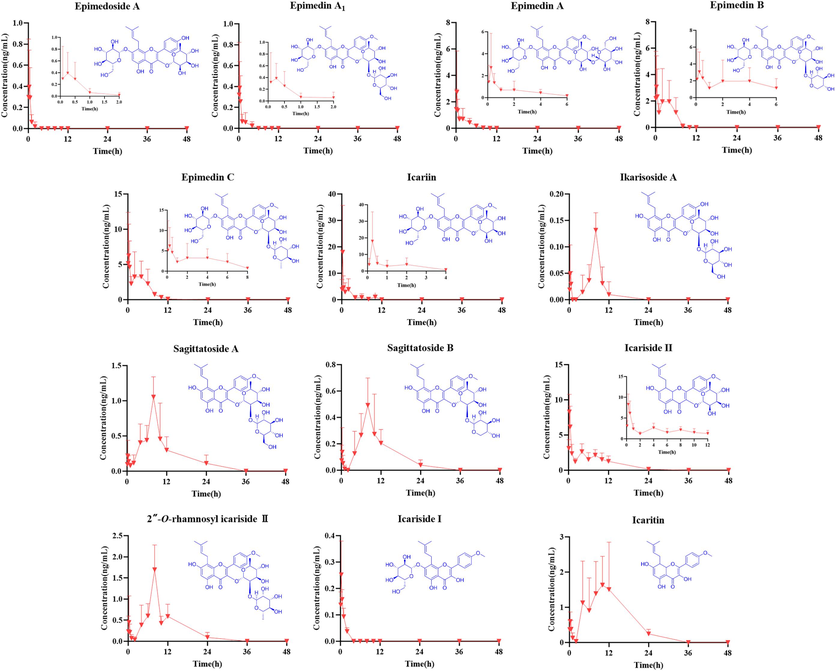
Mean plasma concentration–time profiles of 13 components after oral administration of ETFCs (mean ± SD, n = 6).
Name
Tmax (h)
t1/2 (h)
Cmax (ng/mL)
AUC0-t (ng*h*mL−1)
AUC0-∞ (ng*h*mL−1)
MRT0-t (h)
MRT0-∞ (h)
epimedoside A
1.22 ± 0.96
N/A
0.53 ± 0.18
0.41 ± 0.16
N/A
1.76 ± 0.83
N/A
epimedin A1
0.26 ± 0.13
N/A
0.44 ± 0.38
0.37 ± 0.43
N/A
0.99 ± 0.89
N/A
epimedin A
0.26 ± 0.05
2.14 ± 0.36
2.83 ± 1.27
3.96 ± 0.96
4.41 ± 0.96
2.11 ± 0.38
2.92 ± 0.39
epimedin B
1.84 ± 1.03
1.01 ± 0.24
4.00 ± 1.10
11.17 ± 3.30
13.00 ± 4.90
2.97 ± 0.53
2.99 ± 0.73
epimedin C
1.80 ± 1.04
2.65 ± 1.03
8.00 ± 2.38
23.48 ± 4.60
24.68 ± 5.50
3.69 ± 0.35
4.45 ± 0.66
icariin
0.22 ± 0.02
16.11 ± 5.24
18.22 ± 7.16
22.98 ± 7.16
62.32 ± 21.59
2.68 ± 0.60
19.51 ± 8.17
ikarisoside A
7.67 ± 0.80
N/A
0.24 ± 0.08
0.66 ± 0.16
N/A
7.06 ± 0.27
N/A
sagittatoside A
7.67 ± 0.80
5.60 ± 0.65
1.14 ± 0.16
8.83 ± 1.23
9.14 ± 1.28
10.18 ± 0.96
10.94 ± 0.94
sagittatoside B
8.33 ± 0.33
13.75 ± 7.96
0.67 ± 0.14
4.52 ± 0.71
5.81 ± 1.33
10.04 ± 0.50
21.90 ± 10.67
icariside I
0.29 ± 0.04
0.80 ± 0.09
0.25 ± 0.05
0.22 ± 0.02
0.27 ± 0.02
0.70 ± 0.03
1.16 ± 0.14
icariside II
0.29 ± 0.04
12.72 ± 5.16
8.38 ± 0.98
28.63 ± 2.36
59.04 ± 22.26
6.24 ± 0.53
18.57 ± 7.64
icaritin
8.67 ± 1.12
5.64 ± 0.44
2.24 ± 0.43
22.79 ± 5.41
24.67 ± 5.34
10.73 ± 0.74
10.93 ± 0.69
2″-O-rhamnosyl icariside Ⅱ
7.01 ± 1.42
7.54 ± 3.51
1.85 ± 0.34
12.31 ± 1.63
14.14 ± 2.29
9.53 ± 0.53
14.61 ± 4.82
The pharmacokinetic profiles of the 13 ingredients revealed that the mean plasma concentration–time curves for 6 prototype glycosides (epimedoside A, epimedin A1, epimedin A1, epimedin B, epimedin C, and icariin) exhibited a consistent trend. These glycosides were rapidly absorbed (Tmax < 1h) and quickly eliminated in vivo, with a short mean residence time (MRT). Additionally, the prototype glycosides exhibited a small maximum plasma concentration (Cmax). Their poor absorption in plasma is likely due to the difficulty in transporting through intestinal epithelial cells and initial hydrolysis in the intestines, converting them into secondary glycosides (Chen et al., 2011; Zhou et al., 2013).
Compared with prototype glycosides, the secondary glycosides and aglycones (sagittatoside A, sagittatoside B, 2″-O-rhamnosyl icariside, icariside, and icaritin) displayed distinct pharmacokinetic characteristics. Their pharmacokinetic profiles exhibited two peaks, and the MRT values were obviously prolonged compared to prototype glycosides. Among them, icariside II had a Tmax value of 0.29 ± 0.10 h, differing from other secondary glycosides, as monoglycosides can pass through the intestinal epithelium more quickly than oligoglycosides and aglycones (Fokialakis et al., 2019).
The results of pharmacokinetic parameters showed that icariin, icariside II, and epimedin C had higher exposure levels than others in plasma, with their AUC0-t (ng*h/mL) values at 28.63 ± 5.79, 22.98 ± 17.55, and 23.48 ± 11.27, respectively. Although the initial glycoside forms and their subsequent derivatives had modest areas under the AUC and Cmax values, the glucuronide conjugates exhibited greater bioavailability and a more prolonged elimination phase.
3.4.2 Brain tissue
According to the pharmacokinetics of TCM, only those prototypes or related metabolites that reach effective concentrations and exposure times in blood and target tissues may be the potential bioactive components (Wang, 2002). Considering the influence of the blood–brain barrier in neurological diseases, studying the dynamic changes of components in brain tissue can help identify the effective components of EFTCs. Consequently, a method for the simultaneous determination of 11 prototypes absorbed in brain tissue was established and applied to study the concentration–time changes after ETFCs administration. As depicted in Fig. 6 and Table 4, icariin showed the maximum AUC value, followed by icariside II and epimedin C, with values of 35.36 ± 5.56, 33.29 ± 6.32, and 28.53 ± 4.51 ng*h/mL, respectively. The quantitative ingredients reached maximum concentration rapidly with Tmax values between 0.25–1 h, indicating that isopentenyl flavanones can rapidly distribute through the blood–brain barrier to reach the target tissue. Among them, the prototype glycosides were basically eliminated after 8h, but secondary glycosides had a secondary peak at 8 h, likely due to the transformation of prototype glycosides.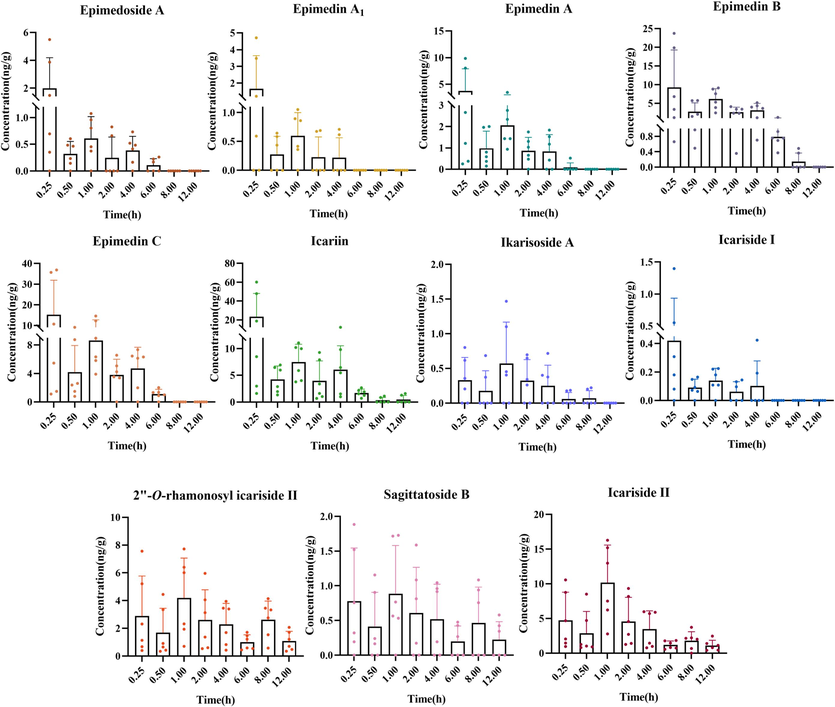
Mean brain tissue concentration–time profiles of 11 components after oral administration of ETFCs (mean ± SD, n = 6).
Name
Tmax (h)
t1/2 (h)
Cmax (ng/g)
AUC0-t (ng*h*g−1)
AUC0-∞ (ng*h*g−1)
MRT0-t (h)
MRT0-∞ (h)
epimedoside A
1.13 ± 0.59
3.63 ± 1.06
3.63 ± 1.06
2.68 ± 0.55
5.04 ± 0.73
1.85 ± 0.34
4.52 ± 1.00
epimedin A1
1.13 ± 0.59
N/A
1.88 ± 0.72
1.70 ± 0.46
N/A
1.21 ± 0.30
N/A
epimedin A
1.25 ± 0.57
N/A
4.29 ± 1.51
5.02 ± 0.98
N/A
1.59 ± 0.36
N/A
epimedin B
1.25 ± 0.57
3.18 ± 1.11
10.68 ± 3.62
18.35 ± 2.92
20.69 ± 3.20
2.45 ± 0.27
4.27 ± 1.51
epimedin C
1.25 ± 0.57
2.30 ± 0.50
17.49 ± 6.04
28.53 ± 4.51
40.91 ± 1.80
2.07 ± 0.24
2.81 ± 1.60
icariin
1.13 ± 0.60
94.95 ± 77.71
25.50 ± 9.35
35.36 ± 5.56
286.79 ± 216.25
3.03 ± 0.57
94.95 ± 77.20
ikarisoside A
1.05 ± 0.28
N/A
0.88 ± 0.18
2.30 ± 0.66
N/A
2.28 ± 0.42
N/A
sagittatoside B
0.96 ± 0.12
10.43 ± 1.17
1.20 ± 0.28
5.75 ± 1.98
16.78 ± 0.99
3.21 ± 0.83
13.71 ± 1.69
icariside I
1.13 ± 0.59
N/A
0.49 ± 0.19
0.41 ± 0.15
N/A
1.15 ± 0.39
N/A
icariside II
1.04 ± 0.23
7.54 ± 1.71
9.53 ± 1.79
33.39 ± 6.32
44.71 ± 7.23
4.27 ± 0.22
10.07 ± 2.48
2″-O-rhamnosyl icariside Ⅱ
2.13 ± 1.20
17.53 ± 9.35
4.85 ± 0.96
24.47 ± 4.59
64.96 ± 15.20
5.50 ± 0.33
24.15 ± 12.32
In addition, we compared the contents of quantitative ingredients in ETFC extract (Xu et al., 2018) with their exposure in plasma and brain tissue. The results revealed that the abundance of icariside II in ETFC extract was much lower than icariin and epimedin C, but their exposure in blood and brain tissue was comparable. The exposure characteristics of prototypes in rat plasma and brain tissue were generally similar. For example, icariin, icariside II, and epimedin C exhibited significant exposure in both plasma and brain tissue, suggesting that they were crucial for the pharmacological effects of ETFCs. The above findings provide a basis for selecting representative components for subsequent in vitro activity evaluations.
3.5 Network pharmacology
A target prediction using the SwissTargetPrediction database revealed that 24 compounds acted on 330 targets. A further 981 neuroinflammation-related protein targets were extracted from the GenCard database. A total of 108 target duplications were identified in the context of neuroinflammation-related targets and compound targets of action (Fig. 7A), indicating that the 24 compounds acted on 108 neuroinflammation-related targets. Subsequently, a compound-target interaction network was constructed, comprising 132 nodes and 385 edges. In this network, 22 compounds and 44 targets exhibited degree values equal to or greater than the mediator value (4) (Fig. 7B and Table S16). Among these, compounds P28, P39, and P42, which had the highest degree values, may represent promising therapeutic agents for neuroinflammation. Similarly, AKR1B1, PDE5A PTGS2 (COX-2), ADRA2A, and ACHE, which also had the highest degree values, may represent promising key targets for the therapeutic effects of these compounds on neuroinflammation and were used for molecular docking (Table S17). A KEGG pathway analysis of 44 key targets revealed that these targets were primarily focused on neuroinflammation disease-related pathways, including MAPK signaling pathway, neuroactive ligand-receptor interaction, and NF-κB signaling pathway pathways (Fig. 7C) (Shabab et al., 2017; Vezzani et al., 2019). To gain further insight into the potential of these compounds for treating neuroinflammation, we conducted a more detailed examination of their impact on inflammatory factors.
(A) The Venn diagram illustrating the common gene targets of ETFCs and neuroinflammatory diseases. (B) The constructed compound-target interaction network. (C) Enriched KEGG pathways and their associated targets (via DAVID).
The results of molecular docking indicated that the binding affinities between the 5 highest-degree targets in the compound-target network and the 23 bioavailable compounds were better than −5.8 kcal/mol (Fig. S8), which suggested the presence of a robust interaction between the small molecule and the target (Li et al., 2020a). Among these targets, COX-2 played a primary role in neuroinflammation and thus became a focus target. In consideration of the docking scoring and the accessibility of the small molecules, only compounds P27, P28, P39, P42 and P43 were subjected to testing for their inhibitory activity against COX-2.
3.6 Anti-neuroinflammatory activity evaluation of the potential active components in ETFCs
Microglial cells play a critical role in regulating neuronal activity, promoting learning, and shaping social behavior. When activated, microglia release various inflammatory mediators that can lead to neuroinflammation (Guruswamy and ElAli, 2017; Woodburn et al., 2021). Given their crucial regulatory role in the central nervous system, inhibiting their activation is key to alleviating neuroinflammation and achieving neuroprotection. The LPS-induced BV-2 cell model is often employed as an in vitro model for anti-neuroinflammatory studies. Based on the quantitative results of plasma and brain tissue, combined with the results of compounds-targets interaction network analysis, 11 ingredients were selected for in vitro anti-neuroinflammatory activity evaluation. First, the maximum non-toxic concentration of each compound was determined (Fig. S9). The results shown in Fig. 8A demonstrated that all 11 compounds could significantly inhibit NO production, exhibiting significant anti-neuroinflammatory activity.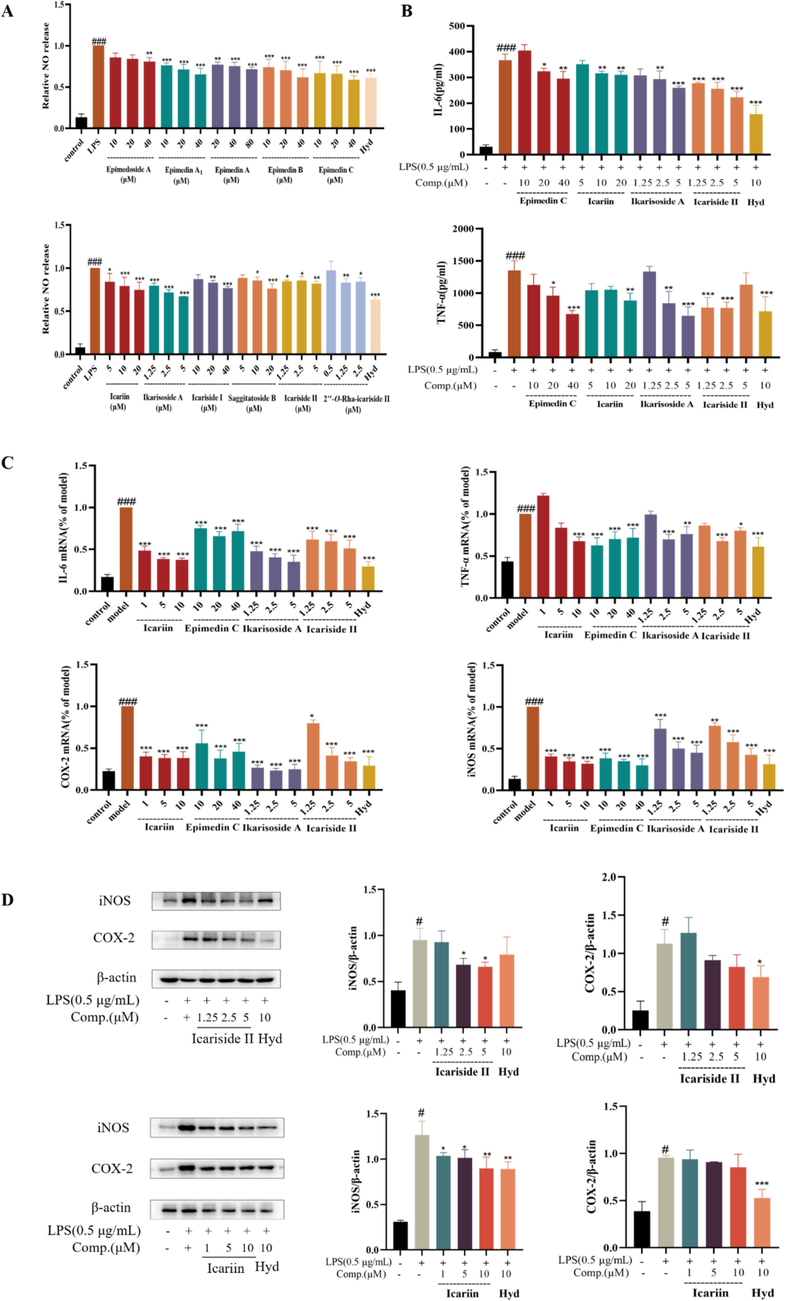
(A) The production of NO in the supernatant. (B) The concentration of IL-6 and TNF-α in the supernatant. (C) Relative mRNA expressions of IL-6, COX-2, TNF-α, and iNOS. (D) The protein expressions of iNOS and COX-2. Data are mean ± SD (n = 3). *P < 0.05, **P < 0.01 vs. cells treated with LPS.
Additionally, based on the exposure levels of the 11 compounds in plasma and brain tissue, as well as their structural characteristics, we selected icariin, icariside II, epimedin C, and ikarisoside A as representative compounds to further verify their anti-neuroinflammatory effects. The results from Elisa and qPCR experiments indicated that these four compounds could inhibit the secretion of the pro-inflammatory cytokines IL-6 and TNF-α, as well as the mRNA expressions of COX-2, iNOS, IL-6, and TNF-α (Fig. 8B-C). Specifically, icariside II and icariin dose-dependently reduced the protein expression levels of iNOS and COX-2 (Fig. 8D). These findings further support the role of these active ingredients in contributing to the efficacy of ETFCs.
4 Discussion
Stroke is a leading cause of mortality and long-term disability worldwide, with devastating consequences on both physical and cognitive functions (Kuriakose and Xiao, 2020; Li et al., 2020b). Among the complications that arise following a stroke, cognitive impairment stands out as a significant challenge, affecting patients' quality of life and posing a burden on healthcare systems (Kalaria et al., 2016; Mijajlović et al., 2017; Van Der Flier et al., 2018). Despite advancements in acute stroke management, there remains a critical need for effective therapeutic strategies targeting post-stroke cognitive impairment (PSCI) (Van Der Flier et al., 2018; Wu et al., 2019). ETFC is a Chinese patent medicine developed based on the TCM theory of “the kidney governing the bones”. Considering the kidney-brain axis hypothesis, we are attempting to explore its indications for treating post-stroke cognitive impairment. In our previous study, an MCAO model was established to investigate cognitive impairments of ETFCs (Yang et al., 2024). The results suggested that ETFCs can effectively improve cognitive impairment and may act through the neuroinflammatory pathway. Therefore, to clarify the in vivo pharmacodynamic basis and mechanisms of ETFCs, this study focuses on neuroinflammation and analyzes the pharmacokinetics of ETFCs, as well as the quantification of exogenous components in target tissues. Through the integration of network pharmacology, potential pharmacodynamically active components were identified, followed by experimental validations against neuroinflammation. Our findings provide scientific support for the mechanisms underlying the amelioration of PSCI by ETFCs through neuroinflammatory pathways.
In the treatment of nervous system diseases, the therapeutic effect of orally administrated TCM depends on whether its components can pass through the blood–brain barrier to reach the target organs. Some TCM components have sufficient bioavailability and brain tissue affinity, while many others struggle to reach the brain due to their chemical properties or molecular size, limiting their application in treating neurological diseases. The chemical composition of ETFCs was characterized by a complex array of flavone compounds, with poor absorption properties (Chen et al., 2008; Szabó et al., 2022). In view of the cognition-enhancing efficacy of ETFCs observed at the animal level, we further studied the blood circulation components of ETFCs and their plasma pharmacokinetics by using mass spectrometry. In particular, to address the question of whether the components reach the brain, a method was established for quantifying the content of components in brain tissue. Additionally, we described the temporal variation characteristics of the brain components at multiple time points. Thus, we have demonstrated that 11 key components of ETFCs can indeed enter brain tissue, thereby providing experimental support for its efficacy in improving cognitive function after stroke.
In network pharmacology study, we believe that it not only helps to reveal the multi-component and multi-target mechanisms of TCM but also provides significant support for the clinical application and pharmacological substance basis research (Shao and Zhang, 2013; Van Hasselt and Iyengar, 2019). The chemical composition of ETFCs is complex, comprising 65 identified components within the formula and over 130 exogenous substances detected in biological samples such as plasma. It is often difficult to evaluate and verify one by one by using traditional experimental methods, and it is even more challenging to comprehensively understand its mechanism of action. Therefore, in this study, the network pharmacology technique was used to reveal the interaction network of multiple components and targets of ETFCs at the overall level and to identify its key active ingredients. A target prediction and network analysis identified 24 compounds acting on 108 neuroinflammation-related targets, highlighting key compounds (P28, P39, P42) and targets (AKR1B1, PDE5A, and PTGS2) involved in crucial neuroinflammation pathways. Based on previous animal experiments, the highly exposed components in brain tissue, as well as the structural representativeness of the components, we conducted experimental validation of 11 potential bioactive components in neuroinflammation pathways. The results showed that all 11 compounds could significantly inhibit the release of NO. Icariin, icariside II, epimedin C, and ikarisoside A were demonstrated to significantly reduce the secretion of inflammatory factors IL-6 and TNF-α, and inhibit their expression at the mRNA level, while also reducing the mRNA expressions of COX-2 and iNOS. Notably, icariin and icariside II, dose-dependently reduced iNOS protein expression. These results suggest that the structural diversity of isoprenyl flavonoids in E. brevicornu, undergo complex and diverse metabolic reactions in vivo, transforming into corresponding secondary glycosides and metabolites to exert their effects. This provides important support for the new clinical application and pharmacological substance basis of ETFCs. This study acknowledges its limitations: it has thus far only evaluated the neuroinflammatory activity of 11 potential active ingredients in vitro. Future research should consider assessing a broader range of inflammatory markers and conducting further validation in animal models with larger sample sizes to enhance the generalizability of the findings. Moreover, the current investigation focused on the activity of individual compounds, neglecting the “multi-components” nature inherent in Traditional Chinese Medicine formulas. Subsequent studies should aim to explore the combined in vivo and in vitro neuroinflammatory effects and mechanisms of these compound mixtures.
5 Conclusion
This study investigated the pharmacodynamic components of ETFCs against neuroinflammation through an integrated four-step strategy involving both in vitro and in vivo analyses. First, the metabolic profile of ETFCs was systematically characterized using UPLC-Q/TOF-MS, resulting in the detection of a total of 130 xenobiotics (44 prototypes and 86 metabolites) in rat biological samples after oral administration of ETFCs. Second, two UPLC-TQ-MS/MS analytical methods were developed to describe the pharmacokinetic characteristics of the major absorbed components in rat plasma and brain tissue, respectively. Based on the quantitative data of plasma and brain tissue, combined with the network pharmacology analysis, 11 components with high exposure levels were screened out, potentially contributing to the anti-neuroinflammatory effect. Finally, the bioactivity evaluations provided the interpterion of the ETFCs’s anti neuroinflammatory activities and mechanism. Conclusively, these findings provide a valuable reference for further research.
CRediT authorship contribution statement
Xiaochun Zeng: Writing – original draft, Methodology, Investigation, Formal analysis, Data curation, Conceptualization. Junran Shao: Writing – original draft, Methodology, Investigation, Formal analysis, Data curation, Conceptualization. Dabo Pan: Software, Formal analysis, Data curation. Siying Zeng: Investigation, Formal analysis, Data curation. Zhenzhong Wang: Supervision, Conceptualization. Xinsheng Yao: Software, Conceptualization. Haibo Li: Writing – review & editing, Supervision, Methodology, Investigation, Funding acquisition, Data curation, Conceptualization. Wei Xiao: Writing – review & editing, Supervision, Methodology, Investigation, Funding acquisition, Data curation, Conceptualization. Yang Yu: Writing – review & editing, Methodology, Investigation, Funding acquisition, Data curation, Conceptualization.
Acknowledgements
This work is supported by Basic Research Program Natural Science Fund -Frontier Leading Technology Basic Research Special Project (No. SBK2023050003). University Science and Technology Innovation Team of Department of Education of Guizhou Province (QJJ [2023]099).
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Anti-inflammatory and immunoregulatory effects of icariin and icaritin. Biomed. Pharmacother.. 2022;151:113180
- [CrossRef] [Google Scholar]
- Chemical constituents, quality control, and bioactivity of Epimedii Folium (Yinyanghuo) Am. J. Chin. Med.. 2015;43(05):783-834.
- [CrossRef] [Google Scholar]
- Intestinal absorption mechanisms of prenylated flavonoids present in the heat-processed Epimedium koreanum Nakai (Yin Yanghuo) Pharm. Res.. 2008;25(9):2190-2199.
- [CrossRef] [Google Scholar]
- Role of intestinal hydrolase in the absorption of prenylated flavonoids present in Yinyanghuo. Molecules. 2011;16(2):1336-1348.
- [CrossRef] [Google Scholar]
- Research progress on the regulation of kidney-tonifying Chinese herbs on brain based on theory of interaction between kidney and brain. Chin. J. Tradit. Chin. Med.. 2021;36(04):2179-2183.
- [Google Scholar]
- Biological evaluation of isoflavonoids from Genista halacsyi using estrogen-target cells: activities of glucosides compared to aglycones. PLoS One. 2019;14(1):e0210247.
- [Google Scholar]
- Metabolites profile of Xian-Ling-Gu-Bao capsule, a traditional Chinese medicine prescription, in rats by ultra performance liquid chromatography coupled with quadrupole time-of-flight tandem mass spectrometry analysis. J. Pharm. Biomed. Anal.. 2014;96:90-103.
- [CrossRef] [Google Scholar]
- SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res.. 2014;42(W1):W32-W38.
- [CrossRef] [Google Scholar]
- Complex roles of microglial cells in ischemic stroke pathobiology: new insights and future directions. Int. J. Mol. Sci.. 2017;18(3):496.
- [CrossRef] [Google Scholar]
- Phytochemistry and ethnopharmacology of Epimedium L. species. Chin. Herb. Med.. 2015;7(3):204-222.
- [CrossRef] [Google Scholar]
- An outline for the pharmacological effect of icariin in the nervous system. Eur. J. Pharmacol.. 2019;842:20-32.
- [CrossRef] [Google Scholar]
- Stroke injury, cognitive impairment and vascular dementia. Biochim. Biophys. Acta Mol. Basis Dis.. 2016;1862(5):915-925.
- [CrossRef] [Google Scholar]
- Pathophysiology and treatment of stroke: present status and future perspectives. Int. J. Mol. Sci.. 2020;21(20):7609.
- [CrossRef] [Google Scholar]
- Pharmacological effects and pharmacokinetic properties of icariin, the major bioactive component in Herba Epimedii. Life Sci.. 2015;126:57-68.
- [CrossRef] [Google Scholar]
- A systems pharmacology approach for identifying the multiple mechanisms of action for the Rougui‐Fuzi herb pair in the treatment of cardiocerebral vascular diseases. Evid. Based Complement. Alternat. Med.. 2020;2020(1):5196302.
- [CrossRef] [Google Scholar]
- Multi-compound and drug-combination pharmacokinetic research on Chinese herbal medicines. Acta Pharmacol. Sin.. 2022;43(12):3080-3095.
- [CrossRef] [Google Scholar]
- Trends in stroke incidence in high-income countries in the 21st century. Stroke.. 2020;51(5):1372-1380.
- [CrossRef] [Google Scholar]
- Clinical curative effect in depression patients with kidney deficiency and liver stagnation treated with kidney-tonifying and depression resolving method. Chin. Arch. Tradit. Chin. Med.. 2013;31(10):2143-2145.
- [Google Scholar]
- Treatment of primary osteoporosis with epimedium total flavone capsule: a multicenter clinical observation on 360 cases. Chin. J. Osteoporos.. 2013;19(3):279-282.
- [CrossRef] [Google Scholar]
- The genus Epimedium: an ethnopharmacological and phytochemical review. J. Ethnopharmacol.. 2011;134(3):519-541.
- [CrossRef] [Google Scholar]
- Post-stroke dementia – a comprehensive review. BMC Med.. 2017;15(1):1-12.
- [CrossRef] [Google Scholar]
- A systematic review of traditional uses, phytochemistry, pharmacology and toxicity of Epimedium koreanum Nakai. J. Ethnopharmacol.. 2024;318:116957
- [CrossRef] [Google Scholar]
- Determination of the pharmacodynamic substances and mechanism of Shiwuwei Saierdou Pills against cholestatic hepatitis through chemical profile identification and network pharmacology analysis. Arab. J. Chem.. 2024;17(2):105504
- [CrossRef] [Google Scholar]
- Neuroinflammation pathways: a general review. Int. J. Neurosci.. 2017;127(7):624-633.
- [CrossRef] [Google Scholar]
- Traditional Chinese medicine network pharmacology: theory, methodology and application. Chin. J. Nat. Med.. 2013;11(2):110-120.
- [CrossRef] [Google Scholar]
- Bioavailability improvement strategies for icariin and its derivates: a Review. Int. J. Mol. Sci.. 2022;23(14):7519.
- [CrossRef] [Google Scholar]
- Systems pharmacology: defining the interactions of drug combinations. Annu. Rev. Pharmacol. Toxicol.. 2019;59:21-40.
- [CrossRef] [Google Scholar]
- Neuroinflammatory pathways as treatment targets and biomarkers in epilepsy. Nat. Rev. Neurosci.. 2019;15(8):459-472.
- [CrossRef] [Google Scholar]
- Studies on serum pharmacochemistry of traditional Chinese medicine. World Sci. Technol.. 2002;4(2):1-5.
- [Google Scholar]
- The semantics of microglia activation: neuroinflammation, homeostasis, and stress. J. Neuroinflammation. 2021;18(1):1-16.
- [CrossRef] [Google Scholar]
- Epimedium aqueous extract ameliorates cerebral ischemia/reperfusion injury through inhibiting ROS/NLRP3-mediated pyroptosis. Antioxidants. 2023;12(5):999.
- [CrossRef] [Google Scholar]
- Stroke in China: advances and challenges in epidemiology, prevention, and management. Lancet Neurol.. 2019;18(4):394-405.
- [CrossRef] [Google Scholar]
- Revealing the potential bioactive components and mechanism of Qianhua Gout Capsules in the treatment of gouty arthritis through network pharmacology, molecular docking and pharmacodynamic study strategies. Heliyon. 2024;10(10):e30983.
- [Google Scholar]
- Molecular mechanism of Epimedium in the treatment of vascular dementia based on network pharmacology and molecular docking. Front. Aging Neurosci.. 2022;14:940166
- [CrossRef] [Google Scholar]
- Determination of flavonoids constituents in epimedium total flavone capsule and research on its anti-osteoporosis activity. Chin. J. Chin. Mater. Med.. 2018;43(15):3140-3144.
- [CrossRef] [Google Scholar]
- Molecular mechanism of Epimedium extract against ischemic stroke based on network pharmacology and experimental validation. Oxid. Med. Cell Longev.. 2022;2022:3858314.
- [CrossRef] [Google Scholar]
- Effects of epimedium total flavone capsules on post-stroke cognitive impairment in rats. Chin. J. Chin. Mater. Med.. 2024;49(8):2262-2272.
- [CrossRef] [Google Scholar]
- Flavonoids of herba Epimedii stimulate osteogenic differentiation and suppress adipogenic differentiation of primary mesenchymal stem cells via estrogen receptor pathway. Pharm. Biol.. 2016;54(6):954-963.
- [CrossRef] [Google Scholar]
- Advancements in the biotransformation and biosynthesis of the primary active flavonoids derived from Epimedium. Molecules. 2023;28(20):7173.
- [CrossRef] [Google Scholar]
- Network pharmacology, a promising approach to reveal the pharmacology mechanism of Chinese medicine formula. J. Ethnopharmacol. 2023;309:116306
- [CrossRef] [Google Scholar]
- A comparative study on the metabolism of Epimedium koreanum Nakai-prenylated flavonoids in rats by an intestinal enzyme (Lactase Phlorizin Hydrolase) and intestinal flora. Molecules. 2013;19(1):177-203.
- [CrossRef] [Google Scholar]
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2024.106084.
Appendix A
Supplementary data
The following are the Supplementary data to this article:Supplementary Data 1
Supplementary Data 1







