Translate this page into:
Sesquiterpenoids of Dendrobium nobile Lindl. aqueous extract for inhibition alcoholic liver injury through RTK/ELF4regulated inflammation in mouse hepatic macrophages
⁎Corresponding authors. tandp@zmu.edu.cn (Daopeng Tan), yqhe2016@zmu.edu.cn (Yuqi He)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Sesquiterpenoids of Dendrobium nobile Lindl. aqueous extract for inhibition alcoholic liver injury through RTK/ELF4-regulated inflammation in hepatic macrophages.
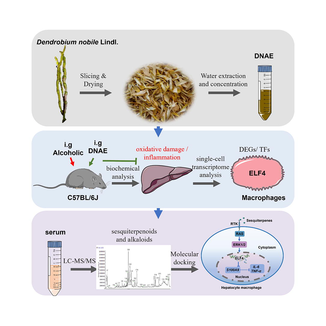
Abstract
DNAE as source of anti-inflammation molecules for alcoholic liver injury. Cell types, DEGs, and TFs of liver immune cells in mice were profiled by scRNA-Seq. 20 sesquiterpenoids and 12 alkaloids of D.nobile in vivo were identified by LC-MS/MS. Interactions between sesquiterpenoids and RTK/ERK1/2/ELF4 signaling were calculated by molecular docking.
Abstract
Dendrobium nobile Lindl. (D.nobile) contributes to an hepatoprotective effect. Unfortunately, the active components and molecular mechanism of D.nobile in alcoholic liver injury is still elusive. Here, we investigated the function of main active ingredients in D.nobile aqueous extract (DNAE) in vivo on regulating the expression of E74-like factor 4 (ELF4) in liver macrophages to against alcoholic liver injury through reducing inflammatory response. The differential expressed genes (DEGs) and transcription factors (TFs) in mouse liver immune cells were determined by single-cell transcriptome analysis, and ELF4 expression in macrophages was determined by RT-PCR and Western blot. The main metabolites of DNAE in vivo were determined by LC-MS/MS analysis. The molecular docking was used to analyze the interactions between metabolites of DNAE and ELF4 signaling. The results showed that DNAE administration showed a reduction of oxidative damage and inflammatory response when mice were treatment with a daily alcoholic intragastric for 14 days. What is more, alcohol primarily affected the expression of macrophages genes among liver immune cells and the DEGs were mainly regulated by transcription factor ELF4, which was relevant to inflammation reaction in macrophages. Meanwhile, DNAE upregulated ELF4 expression through the RTK/ERK1/2 axis. ELF4 inhibits macrophage inflammatory responses via transactivating its upstream target genes S100A9. Furthermore, the main metabolites of DNAE were sesquiterpenoids and alkaloids in vivo, and sesquiterpenoids are more strongly correlated with the cell membrane cascade response RTK/ERK1/2 signaling pathway, which could regulate the expression of ELF4. This study explores the essentiality of sesquiterpenoids to regulate ELF4 in macrophages and indicates that sesquiterpenoids could be as an anti-inflammation drug or a daily functional healthcare product to alcoholic liver disease.
Keywords
Macrophages
Alcoholic liver disease
D.nobile
Inflammatory response
1 Introduction
Dendrobium nobile Lindl. (D.nobile), also known as Jin Chai Shi Hu in Chinese, is a well-known edible plant material and used as immunity enhancer for thousands of years (Li et al., 2019; Liu et al., 2022a, 2022b). D.nobile is a species of orchid in the family Orchidaceae, distributed throughout Asia, including Thailand, Laos, Vietnam, and China (Li et al., 2022a, 2022b). Accumulating evidence suggests that aqueous and ethanol extracts of D.nobile exhibit regulatory effects on hepatic lipid homeostasis, liver inflammation and liver fibrosis through ameliorated oxidative stress, inflammation and metabolism disruption (Xu et al., 2017a, 2017b; Huang et al., 2022). And it is widely known alcoholic liver injury, which ranges from asymptomatic hepatic steatosis to alcoholic steatohepatitis, consequently leading to liver fibrosis and cirrhosis, is intimately associated with inflammatory response (Bajaj, 2019; Dukic et al., 2023). The active ingredients of D.nobile including alkaloids, sesquiterpenes, polysaccharides, and phenanthrenes compounds, which have anti-oxidation, anti-inflammation, anti-aging properties (Nie et al., 2020; Lei et al., 2022). D.nobile contributes to an hepatoprotective effect, thus, we hypothesize that D.nobile has a protective effect against alcoholic liver injury. However, the complex metabolic process limits the clarification of the anti-inflammatory mechanism of D.nobile in vivo. Liquid chromatography-mass spectrometry/mass spectrometry (LC-MS/MS) is a powerful approach for identifying of biologically active compounds. Based on analysis of metabolic profiling data measured by LC-MS/MS technology, the critical bioactive metabolites of D.nobile has been characterized, which including 11 alkaloids, 10 sesquiterpenes, and 13 other metabolites (Tan et al., 2020; Liu et al., 2022a, 2022b; Daopeng et al., 2023; Tan et al., 2023), thus, a comprehensive understanding of those ingredients in vivo and the hepatoprotective effects in alcohol-induced liver injury is needed for further investigation.
Inflammatory response plays a crucial role in the development of alcoholic liver disease. Acetaldehyde, the key alcohol toxic metabolite in the liver, is involved in the generation of reactive oxygen species (ROS) (Yan and Zhao, 2020), and eventually leading to hepatotoxic effect through oxidative stress and an inflammatory cascade (Contreras-Zentella et al., 2022). In addition, as alcohol intake, liver macrophages including the resident macrophages (Kupffer cells, KCs) and monocyte-derived macrophages are activated by exogenous and endogenous antigens (such as lipopolysaccharide and ROS) and final resulted in the activation of inflammation response. It has been proven that macrophages perform a central role in the pathogenesis of acute and chronic inflammatory liver damage (Slevin et al., 2020; Li et al., 2022a, 2022b), and thus, the effect of macrophages in the treatment of alcoholic liver injury needs further investigated.
Alcoholic liver injury develops via a complex process, which involving parenchymal and nonparenchymal cells, as well as recruitment of other cell types to the liver in response to damage and inflammation. Single-cell RNA sequencing (scRNA-seq) represents a powerful approach for the unsupervised characterization of molecular variation in heterogeneous biological systems, is changing the recognitions of the diseases pathogenesis (Chen et al., 2019). Compared to transcriptome sequencing, scRNA-seq could distinguish the gene expression of different cell populations and recognized the expression in cell type specific manner (Johansen and Quon, 2019). It is widely known that alcohol targets numerous signaling pathways in hepatocytes and immune cells (Bukong et al., 2016), and our previous studies also demonstrated hepatocytes and KCs respond more aberrantly differential expressed genes (DEGs) to alcohol than other cell types based on the results of scRNA-seq (Cao et al., 2023). What is more, the aberrantly activated transcription factors (TFs) of Kupffer cells are involved in the process of antigen presentation and inflammation response regulation (Cao et al., 2023). Therefore, comprehensive studies on function of macrophages are required to assess the detailed mechanisms and therapeutic regulation of macrophages-related-inflammation response in the alcoholic liver damage pathogenesis.
In this study, we combined single-cell RNA sequencing (scRNA-seq) and LC-MS/MS technology to reveal the active ingredient of D.nobile in vivo and their anti-inflammation activity on macrophages of mouse with alcoholic liver injury, with the hope to valorize it as a prominent edible plant material for the prevention and co-therapy of inflammation-mediated alcoholic liver pathologies.
2 Materials and methods
2.1 Preparation of D.nobile
The fresh stems of D.nobile were collected from Chishui planting area in Guizhou province and identified of Dendrobium in the Orchidaceae family by Professor Wu from the Department of pharmacognosy, Zunyi Medical University. Specifically, 108.0 g D.nobile dry samples and 3.2 L distilled water (m/v = 1:30) were mixed and soaked overnight in a 5 L round-bottom flask. On the second day, the mixture was boiled reflux extraction for 1 h, and then the extract was filtered and collected. The filter residue was mixed with distilled water and then extracted again using the same method. Combined the twice extracts filtrate and concentrated to 120 mL by the rotary evaporator with reduced pressure. Finally, the D.nobile aqueous extract (DNAE) with a concentration of 0.9 g/mL was stored in a refrigerator at −80 °C. As used, DNAE was removed from the refrigerator to an ice box for thawing, and then diluted to the target concentration with distilled water.
2.2 Animal experiments
Male C57BL/6J mice (22.0 ± 2.0 g) of 8-week-old were purchased from SPF(Beijing) Biotechnology Co., Ltd. Mice were allowed to acclimatize to laboratory conditions (SPF) for at least one week before the experiments. All mice were housed on a 12 h light/dark cycle at a room temperature of 25 °C and had free access to water and food. To study the hepatoprotective effect of D.nobile, mice were randomly divided into 6 groups (n = 15), which including normal group (NG), alcoholic group (AG), low-dose DNAE (0.36 g/kg) treatment group (DNAE-L), middle-dose DNAE (1.8 g/kg) treatment group (DNAE-M), high-dose DNAE (9.0 g/kg) treatment group (DNAE-H), and Silymarin (Sil, 0.05 g/kg) positive treatment group. Sil (Cat. No. R10F11Y108464) was obtained from Shanghai yuanye Bio-Technology Co., Ltd. According to grouping, mice were given distilled water, DNAE, and Sil by gavage for 14 consecutive days, respectively. And on the 14th day, mice were gavage with 53 % ethanol (10 mL/kg) once a day after drug administration 2 h for 14 consecutive days, except the normal group.
After the last administration of alcohol for 12 h, mice were anesthetized with 5 % ethyl carbamate (urethane, 0.25 g/kg). Then, blood and tissue samples were collected (n = 6). Blood was incubated at room temperature for 30 min and then centrifuged at 4500 rpm for 10 min at 4℃ to obtain serum, and then serum was store at −80℃ for further investigated. Liver tissue (approximate 100 mg) was fixed in 10 % formalin for hematoxylin and eosin (H&E) staining, other tissues were stored at −80 °C for further research. To analyze the main components of D.nobile in vivo, DNAE (9.0 g/kg) was administered by gavage on mice. And then blood was collected at 2 h after gavage. Serum was collected by centrifuged and used for the LC-MS/MS analysis. Acetonitrile (Cat. No. A120771), Methanol (Cat. No. M116125) and other organic chemicals used for LC/MS were LC-MS-grade and purchased from Shanghai Aladdin Biochemical Technology Co., Ltd.
All animal experiments were performed by the ethics policy of the Laboratory Animal Welfare & Ethics Committee of Zunyi Medical University (ZMU23-2303–255).
2.3 Single-cell suspensions prepared and isolation, identification the liver macrophages
After the last administration of alcohol for 12 h, mice were anesthetized with 5 % ethyl carbamate (urethane, 0.25 g/kg) and single-cell suspensions from mouse liver (n = 9) were collected. Particularly, hepatocytes and hepatic non-parenchymal cells (NPCs) were isolated by differential centrifugation (DC) after the two-step liver perfusion (Naito et al., 2020; Poggel et al., 2022). Collagenase from Clostridium histolyticum (Cat. No. C5138), DNase I (Cat. No. AMPD1), HEPES (Cat. No. H4034), EGTA (Cat. No. 324626), 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA, Cat. No.287810) were obtained from Sigma Aldrich. For single-cell RNA sequencing, hepatocytes were added to NPCs to give a final hepatocytes concentration of approximately 10 % of the total cell number. The viability of the mixed cells should be higher than 85 %, and the cell concentration was adjusted to 1 × 105 cells/mL in PBS. Chromium Next GEM Single Cell 3′ Reagent Kits v3 was used to construct the single-cell RNA-seq libraries. Illumina HiSeq sequencing platform was selected to sequence data and subsequent data analysis. The Seurat R package was used for quality control, dimensionality reduction, and clustering analyses. (Cao et al., 2023).
For macrophages isolation and identification, NPCs were suspended in Dulbecco's Modified Eagle Medium (DMEM, C11965500BT, Gibco) with 10 % fetal bovine serum (10099141, Gibco) and centrifuged at 350 g for 5 min at 4 °C after lysis of erythrocytes. Then, cell precipitates were collected, resuspended with cell culture medium, and cultured at 37 °C in a humidified incubator containing 5 % CO2 for 3 ∼ 4 h. Remove the cell supernatant softly and collected macrophages for further identification and analysis. Flow cytometry (Navios, Beckman) was used to detect the F4/80 green fluorescence signal which labeled on macrophages (Rabbit monoclonal anti-F4/80 antibody (Cat. No. ab6640) was obtained from Abcam at a dilution of 1:50) and the ROS level of macrophages (2′,7′-dichlorodihydrofluorescein diacetate, DCFH-DA, Cat. No.287810 were obtained from Sigma Aldrich). Cells not immediately used were stored frozen at −80 °C.
2.4 Serum biochemical indicators assay
Total Cholesterol (Alam et al., 2020) Assay Kit (A111-1–1), Triglyceride (TG) Assay Kit (A110-2–1), Low-density Lipoprotein Cholesterol (LDL-C) Assay Kit (A113-1–1), High-density Lipoprotein Cholesterol (HDL-C) Assay Kit (A112-1–1), Aspartate Aminotransferase (Mendrick et al., 2018) Assay Kit (C010-2–1), Alanine Aminotransferase (ALT) Assay Kit (C009-2–1), Gamma‐glutamyl Transferase (GGT) Assay Kit (C017-2–1), Total Superoxide Dismutase (T-SOD) Assay Kit (A001-1–2), Catalase (CAT) Assay Kit (A007-2–1), and Reduced glutathione (GSH) Assay Kit (A006-1–1) were purchased from Nanjing Jiancheng Bioengineering Institute. The content of AST, ALT, TC, TG, LDL-C, HDL-C, SOD, CAT, GSH in serum were measured with the appropriate kits according to the manufacturing instructions.
2.5 Histopathology
According to the Manufacturer's instructions for Staining Assay Kit (G1120, Solarbio Life Sciences), the paraffin sections of liver samples were used for H&E staining. Histomorphology of tissues was observed on an inverted microscope (Olympus, BX43).
2.6 Reverse transcription quantitative PCR (RT-PCR)
Total RNA of macrophages was extracted using TRIzol reagent (15596926CN, Invitrogen TM). Then, purified RNA (2 μg) was reversely transcribed into cDNA using Hifair II 1st Strand cDNA Synthesis SuperMix (11120ES60, Yeasen Biotech Co., Ltd). And then, 2 × SYBR Green qPCR Master Mix (B21202, Bimake) was used to amplify cDNA in a PCR machine (CFX96, Bio-Rad). Primers were supplied by Sangon Biotech Co., Ltd and the sequences were listed in Supplementary Table 1. The mRNA levels were determined by the 2-ΔΔCT method and β-actin as a control.
2.7 Western blotting analysis
Cells were collected and lysed in RIPA lysis buffer (P0013K, Beyotime Institute of Biotechnology) in the presence of a protease inhibitor cocktail (B14001, Selleck). Cellular lysates were centrifuged at 12,000 g at 4 °C for 10 min and the proteins in the supernatant was quantified using the BCA Assay Kit (BCA02, Dingguo Biotechnology Co., Ltd). The proteins (20 μg) were separated by SDS-PAGE and visualized by western blotting using specific antibodies (Rabbit monoclonal anti-ELF4 antibody (Cat. No. ab96075) was obtained from Abcam at a dilution of 1:1000. Aabbit polyclonal anti-β-actin antibody (Cat. No. D110001) was supplied from Sangon Biotech at a dilution of 1:4000). Finally, the BeyoECL Star analysis regent (P0018S, Beyotime Biotechnology) was used to visualize the protein signals under a Biomolecular imager (LAS-4000MINI, BIO-RAD). Fiji software was used for the semi-quantitative analysis.
2.8 Analyze the main metabolites of D.nobile in vivo by LC-MS/MS
Liquid chromatography triple quadrupole tandem mass spectrometer (API 4000™, AB SCIEX) was used for chromatographic separation and quantitative determination. The chromatography separation was performed on an ACQUITY UPLC HSS T3 column (2.1 mm × 100 mm, 1.8 μm). The mobile phases consisted of 0.1 % formic acid water (A) and acetonitrile (B) and the elution gradient program was shown in Supplementary Table 2. The injection volume was 2 μL, the flow rate was 0.3 mL/min, and the temperature of column was 30℃. The mass spectrometry was performed on an electrospray ionization source (ESI) with a positive and negative ion multiple reaction detection mode (scan range of 100 ∼ 1000 m/z). ESI-MS/MS operating parameters were set as follows: ion spray voltage of 3.0 kV (+) / 2.5 kV (-), ion transfer tube temperature of 320 °C, sheath gas flow rate of 35 arb, auxiliary gas flow rate of 10 arb, purge gas flow rate of 0 arb, probe heating temperature of 350 °C. And the mass spectrometry data were imported into Agilent Mass Hunter Profinder 10.0 software for peak matching. Peak alignment, ion fusion and junctional convolution processing, and the fragmented peaks with false positives were excluded based on peak area, retention time and molecular weight. The corresponding molecular formulae were obtained by fitting and calculating with Qualitative Analysis B.07.00 software and matched with the local database for preliminary structure inference. The chemical composition was further identified based on the secondary mass spectrometry data with the information provided by references, Scifinder database, etc.
2.9 Molecular docking
Resolved protein structures of potential targets could be searched in the UniProt database with following filtering conditions: resolution of less than 3 Å, wild-type protein structures, and protein structure with eutectic ligands, then the protein structure was download in a “pdb” format. Opened the “pdb” file through Chimera 1.16 software and the protein structure was selected and saved. Then the AutoDock Tools 1.5.7 software was used to process proteins as follows: remove eutectic ligands and water, add polar hydrogen, calculate the Gasteiger charge and energy optimization, and then the protein was saved as a “pdbqt” file. Furthermore, selected the original protein in the “pdb” format by PyMOL software and searched the center coordinates (x, y, z) of the docked binding site using the GetBox plug-in. The small molecule metabolites of DNAE in vivo were imported into Chem3D Pro 20.0 software and then energy minimization was performed by the MM2 algorithm. Finally, PyMOL and Discovery Studio were used to analyze interactions and visualize the results from the molecular docking analyses to obtain 2D and 3D interactions images (Zhou et al., 2022).
2.10 Single-cell RNA sequencing (scRNA-seq)
Single-cell suspensions were loaded onto microfluidic devices and scRNA-seq libraries were constructed according to the GEM protocol using the Chromium Next GEM Single Cell 3ʹ Reagent Kits v3. After quality checks, individual libraries were diluted to 4 nM and pooled for sequencing data and subsequent data on an Illumina HiSeq sequencing platform. After data normalizing, highly variable genes were identified and used for the following PCA (principal component analysis). FindIntegration Anchors and Integrate Data be used to integrate samples and performed downstream analysis. Subsequently, clustering with 25 principal components and resolution 0.8 was performed by graph-based clustering and visualized using t-Distributed Stochastic Neighbor Embedding (t-SNE) or Uniform manifold approximation and projection (UMAP) with Seurat functions RunTSNE and RunUMAP. Cells were filtered by gene counts between 300 and 6000 and UMI counts between 500 and 40,000. Cells with over 25 % mitochondrial content and 1 % hemoglobin (Oskouei et al., 2023) content were removed.
2.11 Statistical analysis
Data were generated from at least three independent experiments and expressed as mean ± SD. Adjusted p values were calculated with SPSS 17.0 software and differences were considered statistically significant at p < 0.05. Significant differences using a one-way analysis of variance (ANOVA) with Tukey multiple comparison test correction or Unpaired t-test, and the level of confidence is assumed to be 0.05(Son et al., 2017). Figures were assembled by using Prism GraphPad Prism, version 7 (GraphPad Software, Inc., La Jolla, CA, USA).
3 Results and discussion
3.1 Protective effect of D.nobile on alcoholic liver injury
D.nobile, an edible plant material, has been considered as vital allies in the promotion of a healthy lifestyle by preventing chronic diseases through against oxidation damage and inflammation (Pinto et al., 2023). Alcoholic liver injury is caused by acute or chronic excessive alcohol consumption (Rungratanawanich et al., 2021). Many studies have confirmed excessive alcohol consumption in mice for 1 day could increase serum levels of ALT and AST (Hou et al., 2019), for 3 to 7 days induce hepatic steatosis (Fleming and McGee, 1984), for 14 days cause liver cell edema and necrosis (Cao et al., 2021). To explore the hepaprotective effect of D.nobile, mice were given DNAE by gavage for 14 days and liver function enzymes including ALT, AST, GGT, serum lipid content including TC, TG, LDL-C and HDL-C, and hepatic cell morphology were detective after 14 days of alcohol pretreatment (Fig. 1a). Alcohol administration resulted in liver injury in mice, as indicated by augmented liver weight to body weight ratio (Supplementary Fig. 1) and increased serum ALT, AST, and GGT levels. Notably, a prominent does-dependent decreased in ALT, AST and GGT level were detected in alcoholic liver injury mice after DNAE treatment (Fig. 1b), which revealed a substantially hepatoprotective effect of DNAE. Hepatic steatosis severity could be characterized by excessive accumulation of TG (Zhou et al., 2020a, 2020b). Besides, TG and LDL-C levels were significantly up-regulated in mice after alcohol pretreatment, while DNAE could down-regulate serum TG and LDL-C levels. However, there were no significant influence on TC and HDL-C neither alcohol stimulation nor DNAE treatment (Fig. 1c). The result revealed that DNAE substantially downregulated hepatic steatosis. And as shown in Fig. 1d, lipid accumulation, inflammatory infiltration, and hepatocytes death were observed in the liver of mice after alcohol stimulation, while mice liver morphology appeared normal with the DNAE treatment. Histopathological studies on the liver sections were consistent with the liver function tests. These results suggest that consumption of D.nobile during alcohol consumption may improve liver health compared with drinking alcohol alone.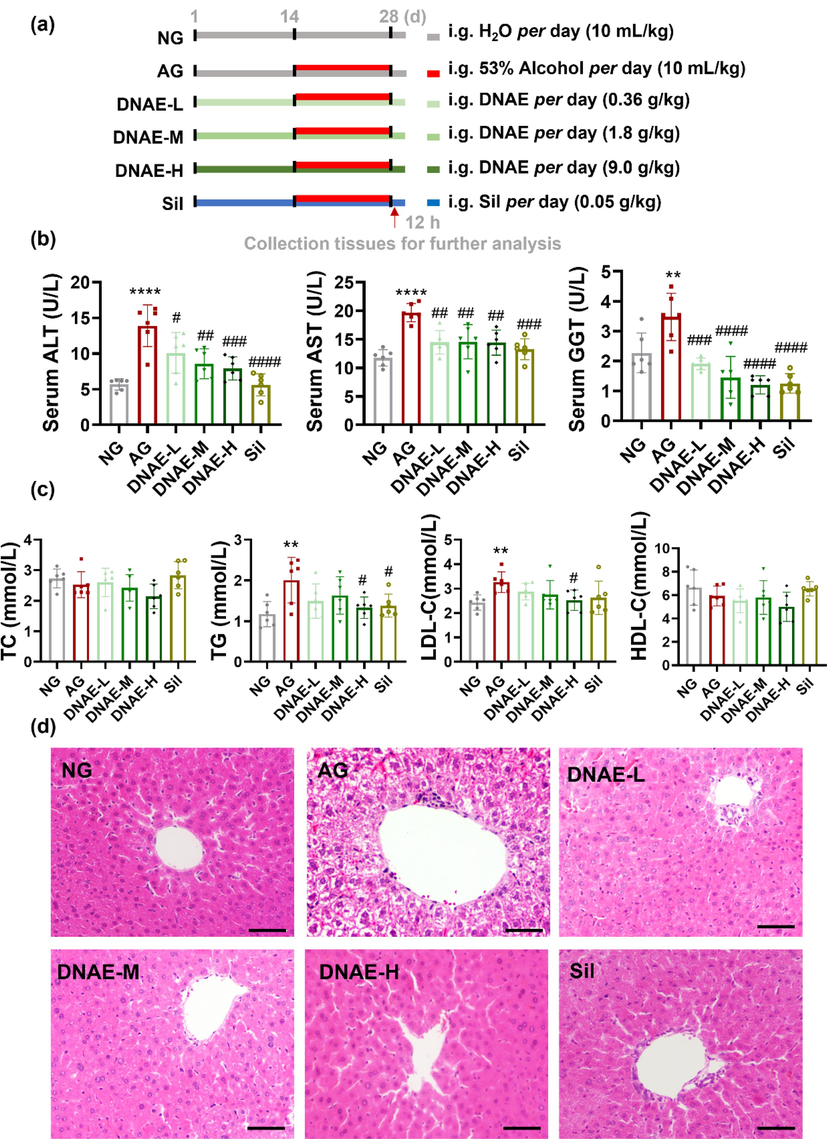
Protective effect of D.nobile on alcoholic liver injury. (a) Overview of the experimental design routinely used in this study. (b) Serum ALT, AST, and GGT levels. (c) serum TC, TG, LDL-C, and HDL-C levels. (d) H&E staining pictures, scar bar = 50 μm. All data are means ± SD (n = 6). * vs. NG. (*p < 0.05, **p < 0.01, ****p < 0.0001), # vs. AG. (#p < 0.05, ##p < 0.01, ### p < 0.001, #### p < 0.0001).
3.2 Changes of gene expression in immune cells of mice with alcoholic liver injury
The pathogenesis of alcoholic liver disease (Alam et al., 2020) includes acetaldehyde-mediated toxicity, hepatic steatosis, oxidative stress, and inflammation (Seitz et al., 2018). In which, inflammation is a consequence of excessive alcohol consumption causing direct and indirect damage to the liver. Clinical and experimental studies of ALD have identified many inflammatory mediators play important roles in the pathogenesis of ALD, and targeting inflammation has already been a clinical treatment of ALD (Xu et al., 2017a, 2017b). Heterogeneity and molecular complexity are characteristic features of ALD. Our recent study highlighting the importance of hepatocyte and hepatic non-parenchymal cells (NPCs) in the development of alcoholic liver injury through the single-cell RNA-sequence (Cao et al., 2023). To investigate the key role of inflammation in alcoholic liver injury, we further performed the changing in TFs of immune cells from alcoholic stimulated liver by single-cell RNA sequence analysis. After standard scRNA-seq filtering excluded cells (Xiong et al., 2019), total 21,397 cells were analyzed by Seurat R package and tSNE plot showed that the liver cells were evenly distributed and no significant intergroup batch effect was observed between the normal group (NG) and alcoholic group (AG) (Fig. 2a). Using uniform manifold approximation and projection (UMAP), we identified 7 clusters of liver immune cells, which were B lymphocytes (B), T lymphocytes (T), Granulocyte (Gran), Dendritic cells (DCs), macrophages (Mac), Natural Killer cells (NK), and Plasma cells (Fig. 2b). Each cell type showed a distinct gene expression profile (Fig. 2c). We further analyzed the differential expressed genes (DEGs) in the liver immune cells. As shown in Fig. 2d, there were 476 DEGs in macrophages, 239 DEGs in Natural Killer cells, 239 DEGs in B lymphocyte, 214 DEGs in T lymphocytes, 237 DEGs in Dendritic cells, 198 DEGs in Granulocyte, and 125 DEGs in Plasma cells. These results indicated that differences of gene expression in immune cells during the progression of alcoholic liver injury in mice and alcohol primarily affected the expression of macrophages genes. Transcription factors (TFs) are key cellular proteins which could be bound to specific regulatory DNA motifs and modulate the expression of target gene and protein synthesis (Bogaert et al., 2023). TFs has been shown to contribute to the development of many different diseases, including immune-related disorders and malignancies (Lee and Young, 2013). Furthermore, our study aimed to investigate immune-related inflammation in alcoholic liver injury, and the change of TFs activity also be analyzed by single-cell regulatory network inference and clustering analysis (SCENIC) in macrophages. As shown in Fig. 2e, there were 8 TFs significantly activated after alcohol stimulated, in which Batf3, Spi1, Spic and Elf4 were decreased while Zmiz1, Z1b1, Gata4 and Sox18 were increased. Co-analyzing the DEGs with TFs target genes, we found that these DEGs were mainly regulated by Spi1, Spic and Elf4 (Fig. 2f). Spi1 is a transcriptional activator involved in macrophages or B lymphocytes differentiation and activation (Gao et al., 2019). Spic is involved in iron homeostasis-associated macrophage development (Alam et al., 2020). ELF4 both sustained the expression of anti-inflammatory genes and limited the upregulation of inflammation amplifiers, thereby restraining the inflammatory response (Tyler et al., 2021). These results suggest that ELF4 might be more relevant to regulate the inflammatory response process during alcoholic liver injury.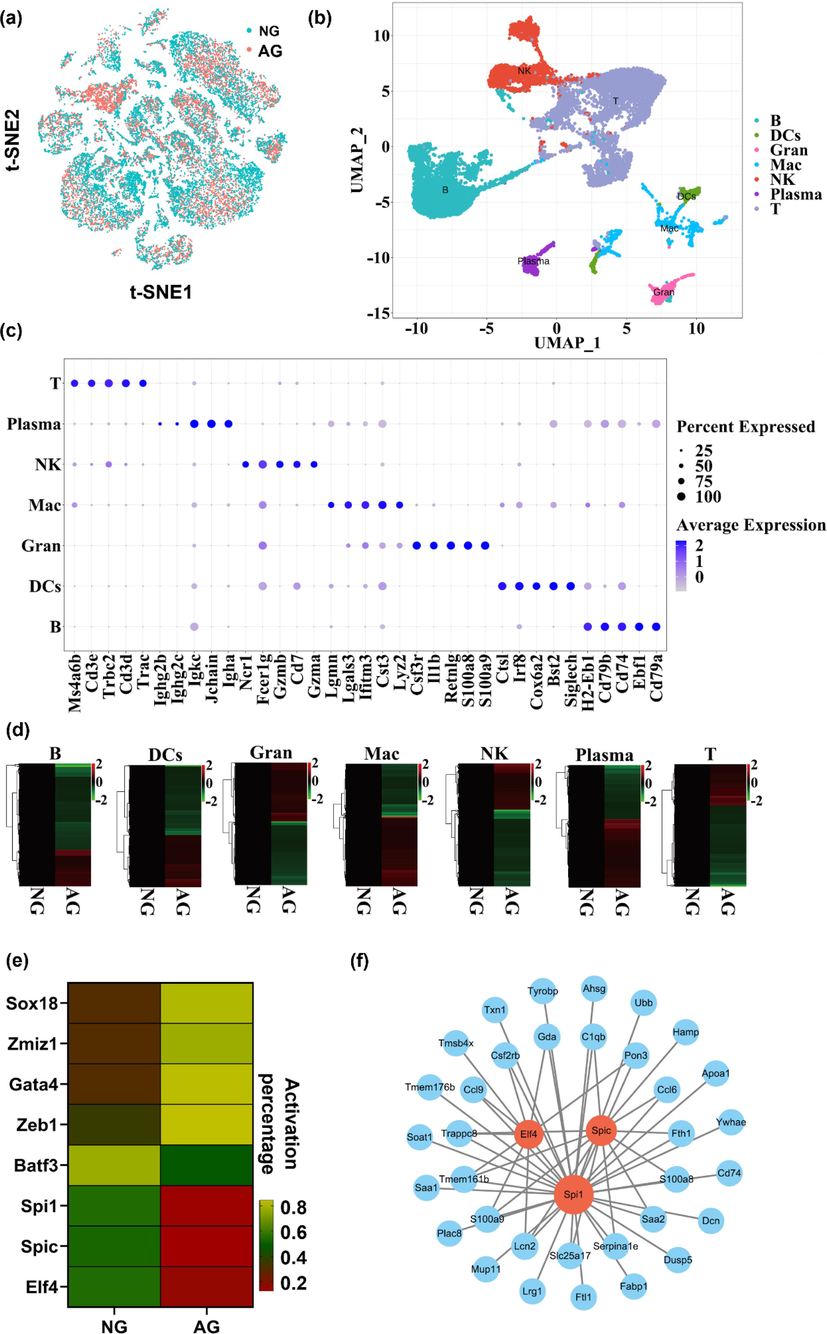
Changes of gene expression in immune cells of mice with alcoholic liver injury. (a) t-SNE plot of the BG and AG cells after integration, each dot represents a cell, and the colors indicate different groups. (b) UMAP dimensional reduction of the immune cells. The immune cells including lymphocytes (B), T lymphocytes (T), Granulocyte (Gran), Dendritic cells (DCs), macrophages (Mac), Natural Killer cells (NK), and Plasma cells in the liver of mice. Population identities were determined based on marker gene expression and 7 distinct immune cells populations were identified. Each point represents a single-cell and the color scheme depicts a cluster. (c) Expression of representative enriched genes for each immune cell type. Gene expression plots are shown in log-scale UMI. Colors correspond to expression level. (d) Heatmap of DEGs (P < 0.01 vs. NG) in 7 different immune cells populations with or without alcohol stimulation. Color scale represents expression levels, red colors indicate high expression level (right upper panel). (e) Heatmap of the percentage of cells with active regulators in different groups of macrophages. Color scale represents activation percentage, green colors indicate high percentage (right lower panel). (f) Intersection network diagram of macrophage DEGs and TFs target genes.
3.3 DNAE reduces oxidative damage and the inflammatory response in macrophages of mice with alcoholic liver injury
Alcoholic liver injury via a complex process involving multicellular in response to oxidative damage and inflammation (Nagy et al., 2016). The generation of ROS during alcohol metabolism leading to oxidative damage and further promotes inflammation. The antioxidant superoxide dismutase (SOD) and catalase (CAT) could nullify the pro-inflammatory effect of ROS (Calvani et al., 2022). And in our study, SOD and CAT level in serum of mice were significantly decreased after alcohol stimulated, while DNAE with low does could increase the level of those antioxidant enzymes (Fig. 3a). The results suggested DNAE had certain antioxidant capacity. However, the ability of D.nobile to regulate oxidative damage and inflammation needs further investigation. The important role of macrophages in alcoholic liver injury has been demonstrated by single-cell RNA sequence analysis. F4/80 has been used widely as a marker for mouse macrophages (Michalet et al., 2005), macrophages labeled with green fluorescent F4/80 protein were detected by flow cytometry in this study. The results showed that almost all macrophages derived from mice were labeled with green fluorescence (Fig. 3b), indicated that the extracted macrophages were of high purity for subsequent analysis. Then, ROS production was measured by flow cytometry and results showed that ROS were significantly elevated in the mouse liver macrophages after alcohol administrated, meanwhile, DNAE reduced the generation of ROS (Fig. 3c). The antioxidant function of D.nobile protects against hepatic injury in mice (Li et al., 2019; Zhou et al., 2020a, 2020b; Lei et al., 2022), and our study further reveals that the DNAE could inhibit oxidative stress by reducing ROS in mouse liver macrophages, which may be associated with the inhibition of inflammation responses. ELF4 belongs to the E74-like factor subfamily of the E26 transformation-specific transcription factor family and is involved in regulating immune responses and development of immune-related cells (Tyler et al., 2021). Our recent work has shown that ELF4 in KCs were sensitive to alcohol induced liver injury in mouse. RT-PCR results also showed that the expression of ELF4 in mouse liver macrophages decreased with alcohol stimulation, which was consistent with the single-cell transcriptome results, meanwhile, DNAE could regulate the expression of TFs ELF4, and the results of western blot were consistent with RT-PCR results (Fig. 3d). To further explore the inflammatory regulation role of ELF4 in macrophages, the expression level of the target gene S100A9 was also examined. As shown in Fig. 3e, the expression of S100A9 was upregulated after alcohol stimulation, and DNAE significantly inhibited the expression of S100A9. ELF4-mutant macrophages had hyperinflammatory responses to a range of innate stimuli and the deficiency of ELF4 augments immune cells inflammatory response (Liu et al., 2022a, 2022b). It is reported that ELF4 limited the upregulation of inflammation amplifiers, including S100A9, S100A8, Lcn2, Trem1 and neutrophil chemoattractant (Tyler et al., 2021). S100A9 with high level proteins had been detective during inflammatory conditions and could stimulated the production of proinflammatory cytokines (Sunahori et al., 2006), such as tumor necrosis factor alpha (TNF-α) and interleukin 6 (IL-6). And in our study, the expressions of proinflammatory cytokines IL-6 and TNF-α in mouse liver macrophages were significantly upregulated after alcohol stimulation, which is consistent with the results reported in previous studies (Ambade et al., 2019). Notably, DNAE significantly inhibited the upregulation of those inflammatory factors (Fig. 3e). These results further confirmed that DNAE could regulate the inflammatory response through ELF4 and thus protect against alcoholic liver injury.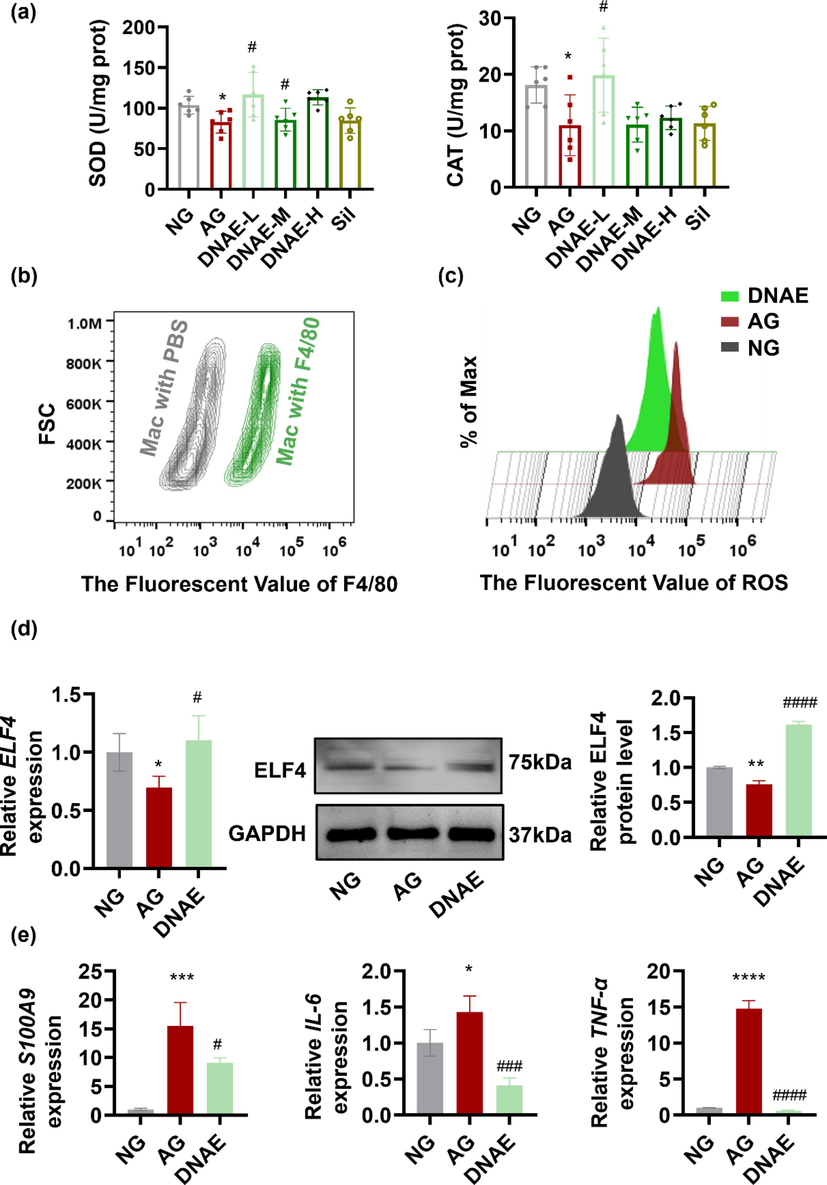
DNAE reduces oxidative damage and the inflammatory response in macrophages of mice with alcoholic liver injury. (a) SOD and CAT level in serum (n = 6). (b) Macrophages (Mac) labeled with green fluorescent protein F4/80 were determined by Flow cytometry. (c) Macrophages were obtained from mice and stained with the fluorescent probe DCFH-DA for 30 min, the FITC fluorescence signal was recorded for intracellular ROS in macrophages by Flow cytometry (n = 3). Data were analyzed using the FlowJo v10 software. (d) The expression level of ELF4 mRNA and protein in macrophages. Fiji was used for the semi-quantitative analysis. (e) The expression level of S100A9, IL-6, and TNF-α mRNA. All data are means ± SD. * vs. NG. (*p < 0.05, **p < 0.01), # vs. AG. (#p < 0.05, ### p < 0.001, #### p < 0.0001).
3.4 The main metabolites of D.nobile in vivo
To further investigate the hepatoprotective effects of DNAE, the main metabolites of DNAE in vivo were investigated. Based on the established LC-MS/MS method, the prepared serum extracts of mice that administrated DNAE by gavage were analyzed. The Compound Discoverer 3.2 software were used to identify and characterize the metabolites. A total of 32 compounds including 20 sesquiterpenoids and 12 alkaloids were separated and identified by the negative and positive ion modes of LC-MS/MS. Detailed information on the identifications, including compound type, name, formula, and fragments were performed in Supplementary Table 3 and the representative extract ion chromatogram (EIC) of DNAE is shown in Supplementary Fig. 2. The structures of these compounds were identified by matching the MS/MS data with a reference substance or through public databases such as MassBank (https://www.massbank.jp/), SciFinder (https://scifinder-n.cas.org), or PubChem (https://pubchem.ncbi.nlm.nih.gov/search/). The identified sesquiterpenoids were shown in Fig. 4. The results showed that sesquiterpenes and alkaloids were the main metabolites of DNAE in vivo, which is in accordance with the previous reports that the mainly active compounds in the stems of D. nobile were alkaloids and sesquiterpenes. Sesquiterpenes not only showed relatively high content but also were present in all regions of the D.nobile stem (Liu et al., 2022a, 2022b). Moreover, sesquiterpenes acted as vital pharmacological activity, which exhibited multiple bioactivities such as anti-inflammatory and immunomodulatory (Gong et al., 2021).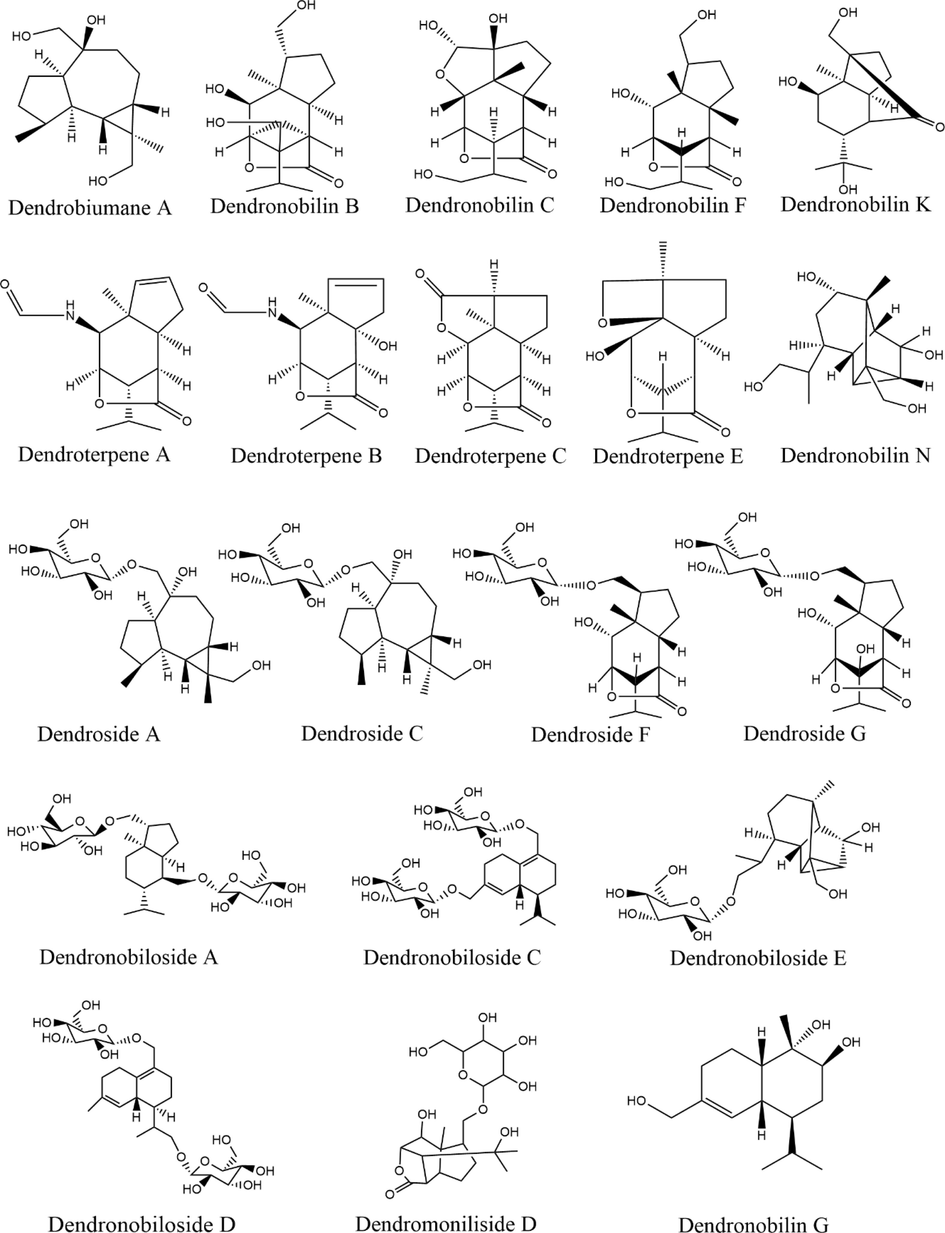
The structures of identified sesquiterpenoids metabolites of D.nobile in vivo.
3.5 Sesquiterpenes mainly activated the RTK signaling links ELF4 with inflammation
Recent transcriptomic analyses revealed that ELF4 modulation directly affected receptor tyrosine kinase (RTK) signaling (Kosti et al., 2023). ELF4 as a transcriptional activator, or through interactions with its partner proteins, could carry out the function which involve in immune response and signaling (Tyler et al., 2021; Liu et al., 2022a, 2022b). Nineteen RTK subfamilies are single-span transmembrane receptors in which divergent extracellular modules are coupled to intracellular kinase domains. The diversity of extracellular domains structures allows for coupling of many unique signaling inputs to intracellular tyrosine phosphorylation to initiate receptor signaling (Trenker and Jura, 2020). Many issues remain to be learned, however, about the complex signaling networks downstream from RTK and how alterations in these networks are translated into cellular responses. Using molecular docking technology, we could visualize the structures of active RTK signaling units, predict the active ingredients and potential targets of DNAE for the treatment of alcoholic liver injury. The detail results of molecular docking were shown in Supplementary Table 4. The binding energy values of the RTK signaling units targets with twenty-five of the 32 metabolites of DNAE acting on them were all less than −7 kcal/mol, which indicated these metabolites have good binding affinity to the key RTK signaling units. According to the theory of receptor-ligand docking, binding affinity is inversely correlated with docking energy, and a more negative docking energy suggests stronger binding affinity between the protein receptor and the ligand (Ye et al., 2021). And the heat map of the correlation between the active metabolites and signaling networks downstream from RTK showed that sesquiterpenes compounds had higher binding affinity to RTK signaling networks compared to alkaloids compounds (Fig. 5a). Particularly, four pairs of metabolites and proteins with the best binding affinity (binding energy values were all less than −8 kcal/mol) were RTK and Dendronobiloside C, RAS and N-trans-feruloyl tyramine, ERK and Dendroside A, MAPK and Dendronobiloside C, respectively. The docking data of RTK and Dendronobiloside C were imported into Ligplot software to analyze the interactions of compound and protein, and Pymol software was used to visualize, the 3D interaction was shown in Fig. 5b and the 2D interaction was shown in Supplementary Fig. 3. There were hydrophobic forces between Dendronobiloside C and Ser147, Ala101, Arg104, Thr92, Glu28, Arg66, Arg68; and hydrogen bound with Cys102 (2.88 Å), Lys100 (2.97 Å), Val103 (3.06 Å, 3.09 Å, 2.55 Å), Lys93(2.95 Å), Phe91(3.09 Å), Glu97 (3.11 Å), Phe67 (2.73 Å, 2.70 Å), Asp69 (2.91 Å), Cys146 (3.01 Å). Those results indicated that Dendronobiloside C had good binding for RTK and the main forces involved were van der Waals forces and hydrophobic forces. Based on the results of molecular ducking, the core active ingredients in DNAE for the treatment of alcoholic liver injury were Dendronobiloside C, Dendroside A and N-trans-feruloyl tyramine targeted on RTK signaling. To further verified the therapeutic mechanism of DNAE is dependent on RTK signaling, RT-PCR was performed with the regulated kinase ERK1/2. The results showed that DNAE have ability to reverse the decrease in ERK1 and ERK2 mRNA expression of mice macrophages due to alcohol administration (Fig. 5c). Extracellular signal-regulated kinases (ERK1/2) could integrate multiple intracellular signaling modules to regulate of various stimulated cellular processes (Arosh and Banu, 2019). Upstream, ERK1/2 could be activated by a small G protein Ras-Raf family members followed by RTK (Unni et al., 2018). Downstream, ERK1/2 regulates several signaling molecules that include protein kinases, protein phosphatases, receptors, TFs, and other proteins. The ERK signaling also involved in ELF4 expression. The sequence on the ELF4 promoter could presume binding sites of transcription factors correlated with the ERK pathway. Furthermore, The ERK1/2/SP1 axis was essential for ELF4 expression (Chen et al., 2023). Taken together, we can conclude that sesquiterpenes-RTK axis upregulates ELF4 expression by activating the ERK1/2 signaling pathway. ELF4 upregulation suppresses inflammatory responses via transactivating S100A9 (Fig. 5d).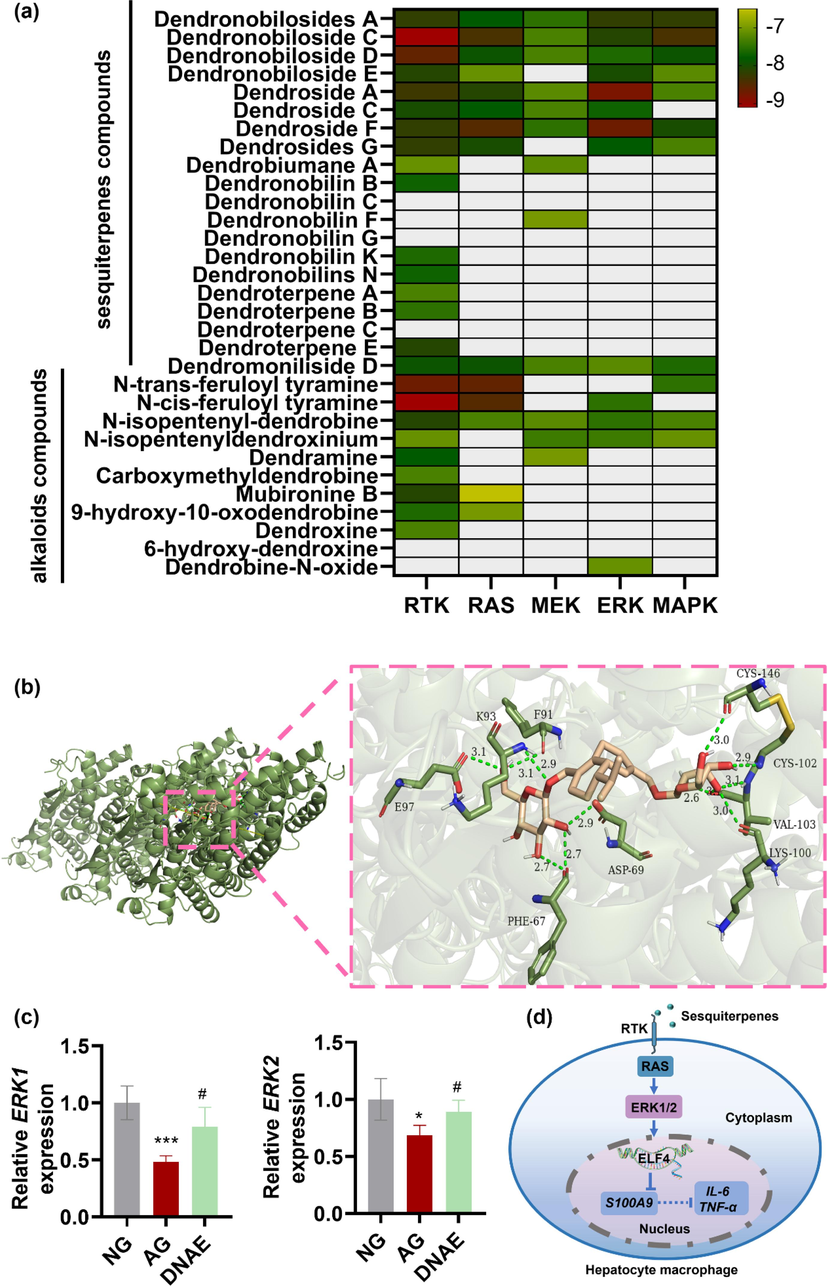
Sesquiterpenes mainly activated the RTK signaling links ELF4 with inflammation. (a) Heat map of the molecular docking results correlation between the active ingredients of DNAE in blood and signaling networks downstream from RTK. (b) In 3D interactions, the colorful ring structures represent the Dendronobiloside C, the green chain structures represent the amino acids forming hydrogen bonds with Dendronobiloside C and the green dashed lines represent the hydrogen bonds formed between the Dendronobiloside C and the protein. (c) ERK1and ERK2 mRNA expression level (n = 4). All data are means ± SD. * vs. NG. (*p < 0.05, ***p < 0.001), # vs. AG. (#p < 0.05). (d) An illustration depicting the signaling pathway of the sesquiterpenes-RTK-ERK1/2-ELF4 in mouse hepatocyte macrophage.
4 Conclusion
This study provides that fourteen days of continuous alcohol intervention could result in alterations of DEGs in mouse immune cells, especially macrophages, and the expression of transcription factor ELF4 in macrophages was associated with alcohol-induced liver inflammatory responses. Furthermore, DNEA administration can reverse alcohol-induced inflammation damage to liver macrophages, by increasing ELF4 expression, and inhibiting S100A9, IL-6, TNF-α expression through the RTK/ERK1/2 signaling pathway. The active metabolites of DNAE in vivo were sesquiterpenes and alkaloids, in which, sesquiterpenes showed the higher binding affinity to RTK signaling networks. And mechanisms underlying the interactions among the sesquiterpenes, oxidative stress, and inflammatory responses are complex and need more in-depth studies in future. Overall, the anti-inflammation and antioxidant capacity of D. nobile make it possible to be a daily functional healthcare product for alcoholic liver injury and inflammatory-associated chronic diseases.
Author contributions
Di Wu performed the experiments. Chengcheng Feng, Ligang Cao and Ju Ye assisted in animal experiments, collecting tissue samples and macrophages, scRNA-seq, bioinformatic analysis, RT-PCR, Western blot, H&E staining. Xingdong Wu Daopeng assisted in active metabolites analysis. Tan, Lin Qin and Yuqi He assisted in conceiving experiments and analyzing data. Di Wu, Daopeng Tan and Yuqi He designed the studies and wrote the paper.
Acknowledgments
The research was supported by grants from the National Nature Science Foundation of China (82160812, 82360741), Guizhou Engineering Research Center of Industrial Key-technology for Dendrobium Nobile (QJJ [2022]048, QJJ [2022]006), the Department of Science and Technology of Guizhou Province (QKHZC [2021]420, QKHZC [2023]261).
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Counter regulation of spic by NF-kappa B and STAT signaling controls inflammation and iron metabolism in macrophages. Cell Rep... 2020;31:107825
- [CrossRef] [Google Scholar]
- Pharmacological inhibition of CCR2/5 signaling prevents and reverses alcohol-induced liver damage, steatosis, and inflammation in mice. Hepatology. 2019;69:1105-1121.
- [CrossRef] [Google Scholar]
- Dual inhibition of ERK1/2 and AKT pathways is required to suppress the growth and survival of endometriotic cells and lesions. Mol. Cell Endocrinol.. 2019;484:78-92.
- [CrossRef] [Google Scholar]
- Alcohol, liver disease and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol.. 2019;16:235-246.
- [CrossRef] [Google Scholar]
- Editorial: the role of transcription factors in inborn errors of immunity. Front. Immunol.. 2023;14:1189312.
- [CrossRef] [Google Scholar]
- Inhibition of spleen tyrosine kinase activation ameliorates inflammation, cell death, and steatosis in alcoholic liver disease. Hepatology. 2016;64:1057-1071.
- [CrossRef] [Google Scholar]
- Two Distinct Superoxidase Dismutases (SOD) secreted by the helminth parasite fasciola hepatica play roles in defence against metabolic and host immune cell-derived Reactive Oxygen Species (ROS) during growth and development. Antioxidants (basel).. 2022;11
- [CrossRef] [Google Scholar]
- Single-Cell RNA transcriptome profiling of liver cells of short-term alcoholic liver injury in mice. Int. J. Mol. Sci.. 2023;24
- [CrossRef] [Google Scholar]
- The effects of Gentiana dahurica Fisch on alcoholic liver disease revealed by RNA sequencing. J. Ethnopharmacol.. 2021;279:113422
- [CrossRef] [Google Scholar]
- FGF19-mediated ELF4 overexpression promotes colorectal cancer metastasis through transactivating FGFR4 and SRC. Theranostics. 2023;13:1401-1418.
- [CrossRef] [Google Scholar]
- Revolutionizing immunology with single-cell RNA sequencing. Cell. Mol. Immunol.. 2019;16:242-249.
- [CrossRef] [Google Scholar]
- Ethanol metabolism in the liver, the induction of oxidant stress, and the antioxidant defense system. Antioxidants (basel). 2022;11
- [CrossRef] [Google Scholar]
- Sesquiterpene glycosides from Dendrobium nobile and their chemotaxonomic significance. Biochem. Syst. Ecol. 2023
- [Google Scholar]
- Alcohol, inflammation, and microbiota in alcoholic liver disease. Int. J. Mol. Sci.. 2023;24
- [CrossRef] [Google Scholar]
- Inflammatory pathways in alcoholic steatohepatitis. J. Hepatol.. 2019;70:249-259.
- [CrossRef] [Google Scholar]
- Recent advances and new insights in biosynthesis of dendrobine and sesquiterpenes. Appl. Microbiol. Biotechnol.. 2021;105:6597-6606.
- [CrossRef] [Google Scholar]
- Characterization of natural melanin from Auricularia auricula and its hepatoprotective effect on acute alcohol liver injury in mice. Food Funct.. 2019;10:1017-1027.
- [CrossRef] [Google Scholar]
- Transcriptome analysis of protection by Dendrobium Nobile Alkaloids (DNLA) against chronic alcoholic liver injury in mice. Biomedicines.. 2022;10
- [CrossRef] [Google Scholar]
- scAlign: a tool for alignment, integration, and rare cell identification from scRNA-seq data. Genome Biol.. 2019;20:166.
- [CrossRef] [Google Scholar]
- ELF4 is a critical component of a miRNA-transcription factor network and is a bridge regulator of glioblastoma receptor signaling and lipid dynamics. Neuro Oncol.. 2023;25:459-470.
- [CrossRef] [Google Scholar]
- Transcriptional regulation and its misregulation in disease. Cell. 2013;152:1237-1251.
- [CrossRef] [Google Scholar]
- Antioxidant and anti-inflammatory activity of constituents isolated from Dendrobium nobile (Lindl.) Front. Chem.. 2022;10:988459
- [CrossRef] [Google Scholar]
- Heterogeneity and function of Kupffer cells in liver injury. Front. Immunol.. 2022;13:940867
- [CrossRef] [Google Scholar]
- Dendrobium nobile Lindl. Polysaccharides protect fibroblasts against UVA-induced photoaging via JNK/c-Jun/MMPs pathway. J. Ethnopharmacol.. 2022;298:115590
- [CrossRef] [Google Scholar]
- Induction of Nrf2 pathway by Dendrobium nobile Lindl. alkaloids protects against carbon tetrachloride induced acute liver injury. Biomed. Pharmacother.. 2019;117:109073
- [CrossRef] [Google Scholar]
- Metabolic profiling, in-situ spatial distribution, and biosynthetic pathway of functional metabolites in Dendrobium nobile stem revealed by combining UPLC-QTOF-MS with MALDI-TOF-MSI. Front. Plant Sci.. 2022;13:1125872.
- [CrossRef] [Google Scholar]
- Intestinal ELF4 deletion exacerbates alcoholic liver disease by disrupting gut homeostasis. Int. J. Mol. Sci.. 2022;23:4825
- [CrossRef] [Google Scholar]
- Metabolic syndrome and associated diseases: from the bench to the clinic. Toxicol. Sci.: An Off. J. Soc. Toxicol.. 2018;162:36-42.
- [CrossRef] [Google Scholar]
- Quantum dots for live cells, in vivo imaging, and diagnostics. Science. 2005;307:538-544.
- [CrossRef] [Google Scholar]
- Linking pathogenic mechanisms of alcoholic liver disease with clinical phenotypes. Gastroenterology. 2016;150:1756-1768.
- [CrossRef] [Google Scholar]
- Isolation of tissue-resident vascular endothelial stem cells from mouse liver. Nat. Protoc.. 2020;15:1066-1081.
- [CrossRef] [Google Scholar]
- Anti-aging properties of Dendrobium nobile Lindl.: From molecular mechanisms to potential treatments. J. Ethnopharmacol.. 2020;257:112839
- [CrossRef] [Google Scholar]
- The effects of Dendrobium species on the metabolic syndrome: a review. Iran. J. Basic Med. Sci.. 2023;26:738-752.
- [CrossRef] [Google Scholar]
- Metabolomic insights into phenolics-rich chestnut shells extract as a nutraceutical ingredient-A comprehensive evaluation of its impacts on oxidative stress biomarkers by an in-vivo study. Food Res. Int.. 2023;170:112963
- [CrossRef] [Google Scholar]
- Isolation of hepatocytes from liver tissue by a novel. Semi-Automated Perfusion Technol. Biomed.. 2022;10
- [CrossRef] [Google Scholar]
- Advanced glycation end products (AGEs) and other adducts in aging-related diseases and alcohol-mediated tissue injury. Exp. Mol. Med.. 2021;53:168-188.
- [CrossRef] [Google Scholar]
- Alcoholic liver disease (vol 4, 16, 2018) Nat. Rev. Dis. Primers. 2018;4
- [CrossRef] [Google Scholar]
- Kupffer cells: inflammation pathways and cell-cell interactions in alcohol-associated liver disease. Am. J. Pathol.. 2020;190:2185-2193.
- [CrossRef] [Google Scholar]
- Thiomer solidification of an ASR bottom ash: optimization based on compressive strength and the characterization of heavy metal leaching. J. Clean. Prod.. 2017;166:12-20.
- [CrossRef] [Google Scholar]
- The S100A8/A9 heterodimer amplifies proinflammatory cytokine production by macrophages via activation of nuclear factor kappa B and p38 mitogen-activated protein kinase in rheumatoid arthritis. Arthritis Res. Ther.. 2006;8:R69.
- [CrossRef] [Google Scholar]
- UDP-glycosyltransferases play a crucial role in the accumulation of alkaloids and sesquiterpene glycosides in Dendrobium nobile. Arab. J. Chem.. 2023;16
- [CrossRef] [Google Scholar]
- Simultaneous quantitative determination of polyphenolic compounds in Blumea balsamifera (Ai-Na-Xiang, Sembung) by high-performance liquid chromatography with photodiode array detector. Int. J. Anal. Chem.. 2020;2020:9731327.
- [CrossRef] [Google Scholar]
- Receptor tyrosine kinase activation: from the ligand perspective. Curr. Opin. Cell Biol.. 2020;63:174-185.
- [CrossRef] [Google Scholar]
- Human autoinflammatory disease reveals ELF4 as a transcriptional regulator of inflammation. Nat. Immunol.. 2021;22:1118-+.
- [CrossRef] [Google Scholar]
- Hyperactivation of ERK by multiple mechanisms is toxic to RTK-RAS mutation-driven lung adenocarcinoma cells. Elife. 2018;7
- [CrossRef] [Google Scholar]
- Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol. Cell. 2019;75(644–660):e645.
- [CrossRef] [Google Scholar]
- Dendrobium nobile Lindl. alkaloids regulate metabolism gene expression in livers of mice. J. Pharm. Pharmacol.. 2017;69:1409-1417.
- [CrossRef] [Google Scholar]
- Targeting inflammation for the treatment of alcoholic liver disease. Pharmacol. Ther.. 2017;180:77-89.
- [CrossRef] [Google Scholar]
- Acetaldehyde induces phosphorylation of dynamin-related protein 1 and mitochondrial dysfunction via elevating intracellular ROS and Ca(2+) levels. Redox Biol.. 2020;28:101381
- [CrossRef] [Google Scholar]
- Exploring the molecular mechanism of action of Yinchen Wuling powder for the treatment of hyperlipidemia, using network pharmacology, molecular docking, and molecular dynamics simulation. Biomed Res. Int.. 2021;2021:9965906
- [CrossRef] [Google Scholar]
- Assessment of pulmonary infectious disease treatment with Mongolian medicine formulae based on data mining, network pharmacology and molecular docking. Chin Herb Med.. 2022;14:432-448.
- [CrossRef] [Google Scholar]
- Dendrobium nobile Lindl. alkaloids-mediated protection against CCl4-induced liver mitochondrial oxidative damage is dependent on the activation of Nrf2 signaling pathway. Biomed. Pharmacother.. 2020;129
- [CrossRef] [Google Scholar]
- Gadofullerene inhibits the degradation of apolipoprotein B100 and boosts triglyceride transport for reversing hepatic steatosis. Sci. Adv.. 2020;6
- [CrossRef] [Google Scholar]
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.arabjc.2023.105501.
Appendix A
Supplementary material
The following are the Supplementary data to this article:Supplementary data 1
Supplementary data 1







